Plant Widely-Targeted Metabolomics Service
Plant Widely-Targeted Metabolomics Service
What Is Plant Widely-Targeted Metabolomics
Plant metabolomics is a powerful mass spectrometry-based approach for revealing the biochemical foundation of plant traits, physiology, and adaptation. Plants produce a vast and highly diverse repertoire of primary and secondary metabolites that govern growth, development, signaling, defense, environmental responses, and the formation of nutritional, sensory, and medicinal characteristics. By capturing these small-molecule changes at scale through LC-MS/MS-based profiling, plant metabolomics enables deeper insight into phenotype, pathway activity, and biological function, making it a valuable tool for functional genomics, agricultural metabolomics, crop improvement, quality evaluation, stress biology, and natural product research.
MetwareBio's Plant Widely-Targeted Metabolomics Service is a high-performance LC-MS/MS metabolite profiling solution specifically designed for plant samples, delivering comprehensive coverage, accurate quantification, and robust reproducibility in a single workflow. The service integrates untargeted and targeted strategies by combining high-resolution MS-based DDA for broad metabolite identification with high-sensitivity MS-based MRM for large-scale quantitative analysis, thereby uniting discovery power with quantitative reliability. Supported by a standardized workflow, rigorous end-to-end quality control, and a proprietary database of more than 61,000 plant metabolites, the platform typically detects over 2,500 metabolites across primary and secondary metabolism, including flavonoids, phenolic acids, alkaloids, terpenoids, lignans, organic acids, amino acids, nucleotides, and lipids. This service is ideally suited for metabolic fingerprinting, differential metabolite profiling, metabolic pathway analysis, and comparative studies across genotypes, tissues, developmental stages, and environmental or treatment conditions.
For readers evaluating platform choice or planning a plant metabolomics project, see our guide to widely-targeted metabolomics workflow, advantages, and applications, as well as our practical plant metabolomics guide.
Why Choose MetwareBio for Plant Metabolomics?
Extensive Plant Metabolite Database—61,000+ Compounds
Broad Coverage with Accurate MRM-Based Quantification
Comprehensive Quality Control for Reproducible Results
Professional Bioinformatics Analysis and Expert Data Interpretation
Publication-Ready Outputs Backed by Proven Project Experience
Multi-Omics Integration Capability
Plant Metabolite Database and Coverage
MetwareBio’s Plant Widely-Targeted Metabolomics Service is powered by a proprietary plant metabolite database containing more than 61,000 plant-associated metabolites associated with plant growth, development, environmental adaptation, nutritional quality, and secondary metabolism. This broad database coverage supports comprehensive plant metabolite profiling and enhances profiling depth, annotation confidence, and pathway interpretation in comparative plant metabolomics studies.
MetwareBio’s in-house metabolite database for Plant Widely-Targeted Metabolomics Service
| Types | Num. | Representative compounds |
| Flavonoids | 7800+ | Rutin, Phloretin, Phelligrin A, Hesperetin, Licorice glycoside A, Pelargonidin-3-O-glucoside, Ginkgetin, Formononetin |
| Phenolic acids | 2500+ | Chlorogenic acid, Momordicoside A, Oleuropein, Salvianolic acid A, Tatariside A, Veratric acid, Salidroside, Parishin B |
| Alkaloids | 8500+ | α-Solanine, Verticine, Nuciferine, Stachydrine, Matrine, Camptothecin, Arecoline, DIMBOA, Avenanthramide A, Lycorenine |
| Terpenoids | 12000+ | Artemisinine, Genipin, Paclitaxel, Wilforlide A, Protopanaxdiol, Saikosaponin A, Cucurbitacin B, Crocin I, Cyclocarioside I |
| Quinones | 1000+ | Emodin, Obtusin, Lapachone, Shikonin, Tectograndone, Morindaparvin A, Aloesin, 5-Hydroxydigitolutein, Trijuganone A |
| Steroid | 1600+ | Asparagoside C, Polyphyllin I, Timosaponin A-III, Gracillin, Sarsasapogenin, Tigogenin, Digitonin, Oleandrin |
| Tannins | 400+ | Ellagic acid, Gemin D, Casuariin, Punicalin, Chebulagic acid, 1,3,6-Tri-O-galloylglucose, Chebulanin, Tellimagrandin I |
| Lignans | 1100+ | Honokiol, Syringaresinol, Arctigenin, Pinoresinol, Schisanhenol, Sesamin, Chestnutlignansoide, Trachelogenin, Fargesin |
| Glucosinolates | 200+ | Sulforaphane, Gluconasturtiin, Sinalbin, Glucocheirolin, Glucoraphanin, 4-Hydroxy-3-indolylmethyl glucosinolate, Sinigrin |
| Coumarins | 1500+ | Umbelliprenin, Psoralen, Glycycoumarin, Xanthotoxol, Scopolin, Bengenin, Bergapten, Decursinol, Dihydrocoumarin |
| Organic acids | 2100+ | Succinic acid, Malic acid, Citric acid, Quinic acid, Abscisic acid, Tartaric acid, Shikimic acid, Aconitic acid, Salicylic acid |
| Vitamins | 50+ | Vitamin C, Vitamin B2, Vitamin A1, Vitamin U, Ginkgotoxin, Nicotinic acid, Nicotinamide, Retinol, Vitamin D3, Tocotrienol |
| Amino acids and derivatives | 800+ | Tryptophan, Theanine, Beauvericin, Dencichin, Heterophyllin, Saccharopine, Alliin, Dopa, S-Adenosylmethionine, γ-Glu-Cys |
| Nucleotides and derivatives | 300+ | Adenine, Cytosine, Thymine, Inosine, Eritadenine, Xanthosine, Cordycepin, Sepiapterin, Adenosine 5'-monophosphate |
| Saccharides and Alcohols | 450+ | Glucose, Fructose, Sucrose, Fucose, Xylitol, Rhamnose, Maltose, Raffinose, Allitol, Mannitol |
| Lipids | 900+ | Linolenic acid, 4-Hydroxysphinganine, Lauric acid, Myristic acid, Palmitic acid, Arachidonic acid, Stearic acid |
| Others | 20000+ | Aflatoxin B1, Secoxyloganin, Kavain, Terreic acid, Mansonone E, Litchiol A, Myricanol, Safranal, Bruceine A, Gambogic acid |
| Total | 61150+ |
Plant Widely-Targeted Metabolomics Workflow
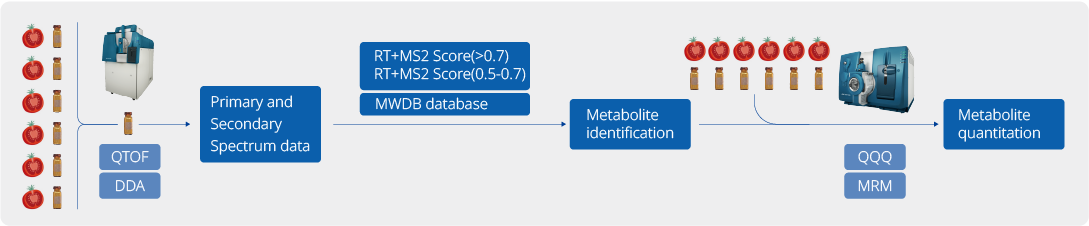
Our Plant Widely-Targeted Metabolomics Service is built on a standardized workflow that covers study design, sample processing, UPLC-MS/MS analysis, metabolite identification, relative quantification, quality control, statistical modeling, and pathway interpretation. Samples are prepared using matrix-appropriate extraction methods and analyzed on a UPLC-MS/MS platform. Metabolites are identified by DDA-supported spectral matching against a proprietary database and quantified in MRM mode for robust and reproducible comparison. Integrated downstream analysis includes quality evaluation, multivariate statistics, differential metabolite analysis, and pathway annotation, enabling reliable insights for plant biology, cultivar comparison, and mechanism studies.
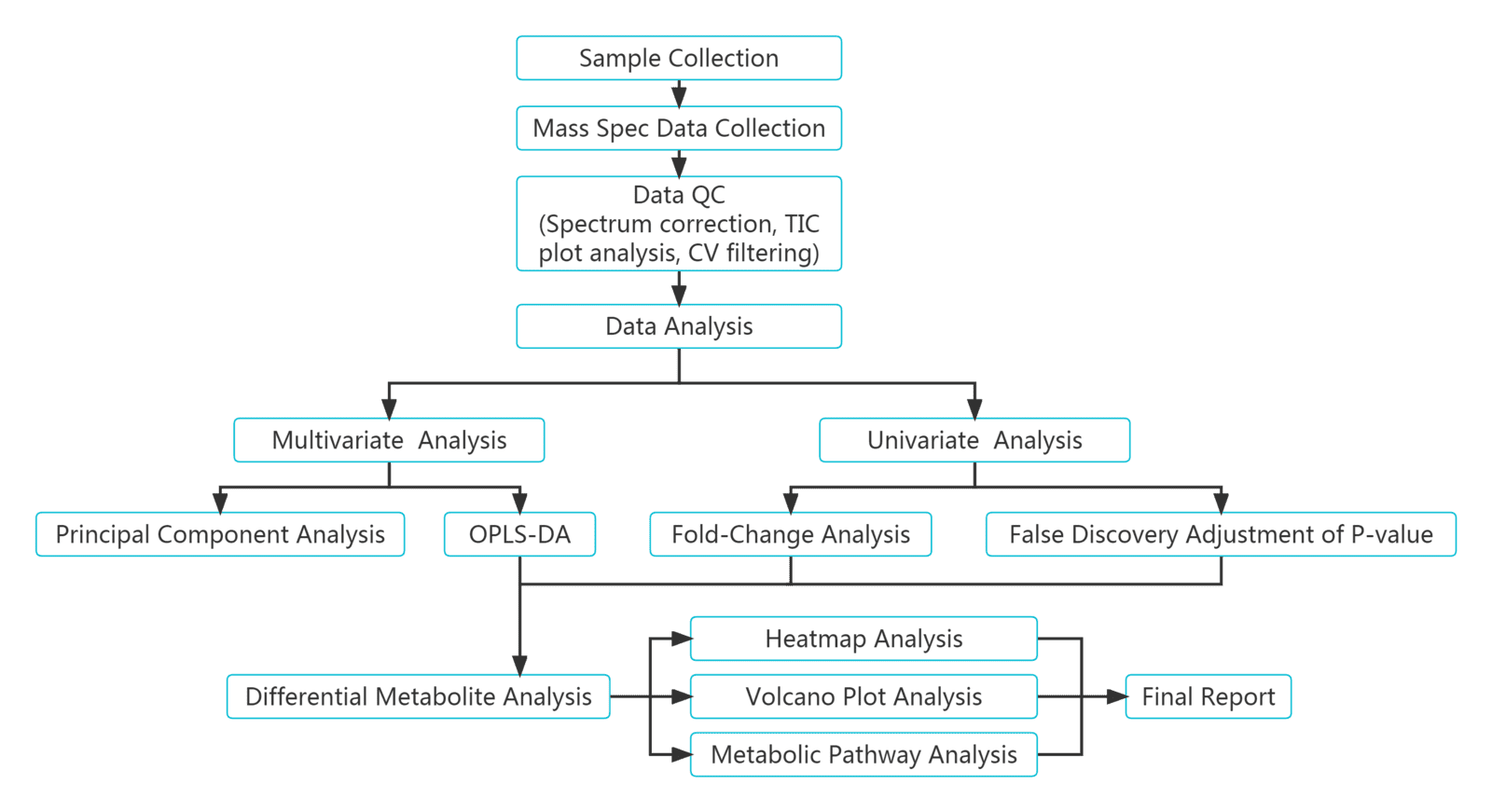
The analysis workflow of plant widely-targeted metabolomics service
Deliverables for Plant Widely-Targeted Metabolomics Service
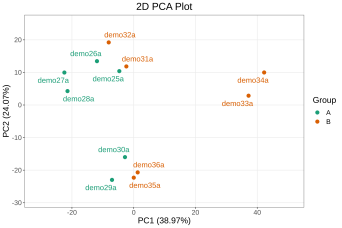
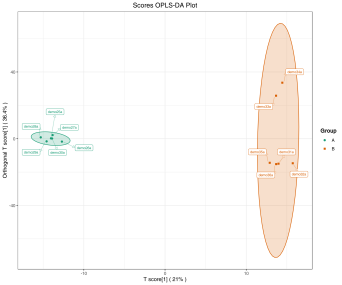
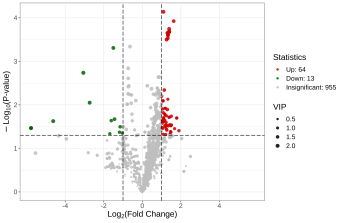
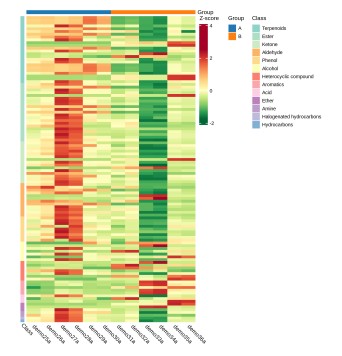
Deliverables for MetwareBio’s Plant Widely-Targeted Metabolomics Service are designed to take researchers from quantitative data review to biological interpretation with clarity and efficiency. Standard outputs include annotated metabolite lists, relative quantification data, quality control assessment, and publication-ready figures. Depending on the study design, result packages may include multivariate statistical analyses such as PCA and OPLS-DA, visualizations for differential metabolite analysis such as volcano plots and clustering heatmaps, and metabolite pathway annotation and enrichment results, supporting mechanism exploration, validation planning, and scientific publication. Contact Us for Demo
Project Experience of Plant Widely-Targeted Metabolomics Service
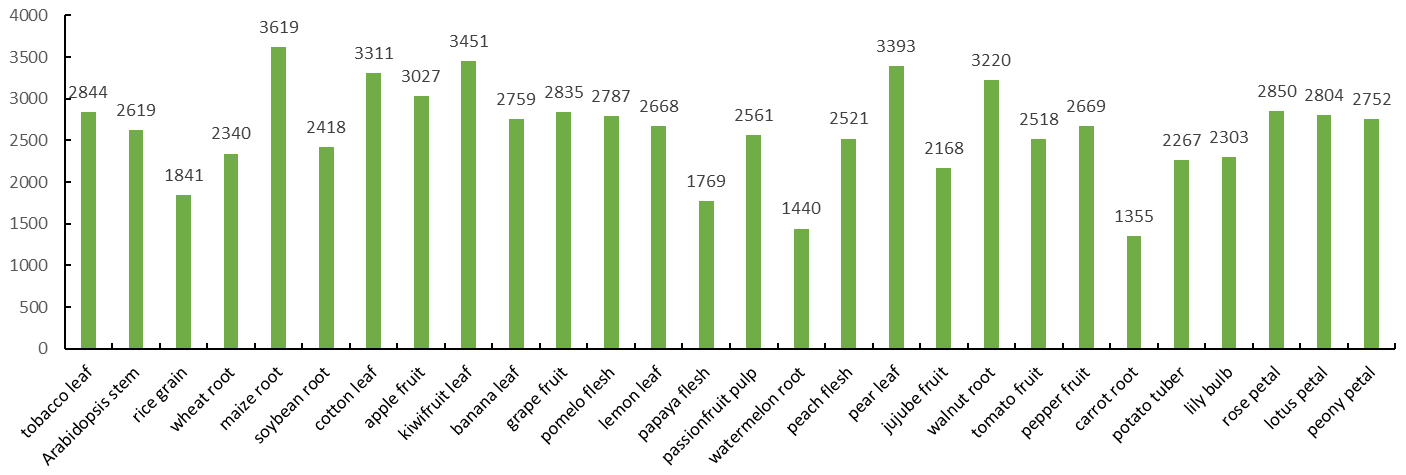
Applications of Plant Widely-Targeted Metabolomics Service
Crop Breeding and Trait-Associated Metabolite Discovery
Plant widely-targeted metabolomics is widely used in crop breeding research to compare metabolic profiles across cultivars, breeding lines, and genotypes. It helps identify trait-associated metabolites linked to yield, color, flavor, nutritional value, stress tolerance, and disease resistance, supporting marker discovery and crop improvement.
Plant Stress Response and Environmental Adaptation
Plant widely-targeted metabolomics is highly valuable for studying how plants respond to drought, salinity, temperature extremes, heavy metal toxicity, nutrient deficiency, pathogen infection, and other environmental challenges. By profiling stress-induced metabolic changes, researchers can uncover adaptive pathways, defense-related metabolites, and biochemical mechanisms underlying plant resilience.
Developmental Profiling and Tissue-Specific Metabolism
Plant widely-targeted metabolomics enables systematic analysis of metabolic variation across tissues and developmental stages, including leaves, roots, stems, flowers, fruits, and seeds. It supports the investigation of tissue-specific metabolite accumulation, developmental regulation, and dynamic pathway changes during plant growth and reproduction.
Quality Evaluation and Natural Product Research
Plant widely-targeted metabolomics service is also well suited for plant quality assessment and natural product studies involving medicinal plants, functional crops, fruits, vegetables, and herbal materials. It helps characterize metabolites associated with nutritional quality, taste, aroma, color, and bioactive properties, supporting quality differentiation, active compound discovery, and product value evaluation.
Case Studies of Plant Widely-Targeted Metabolomics Service
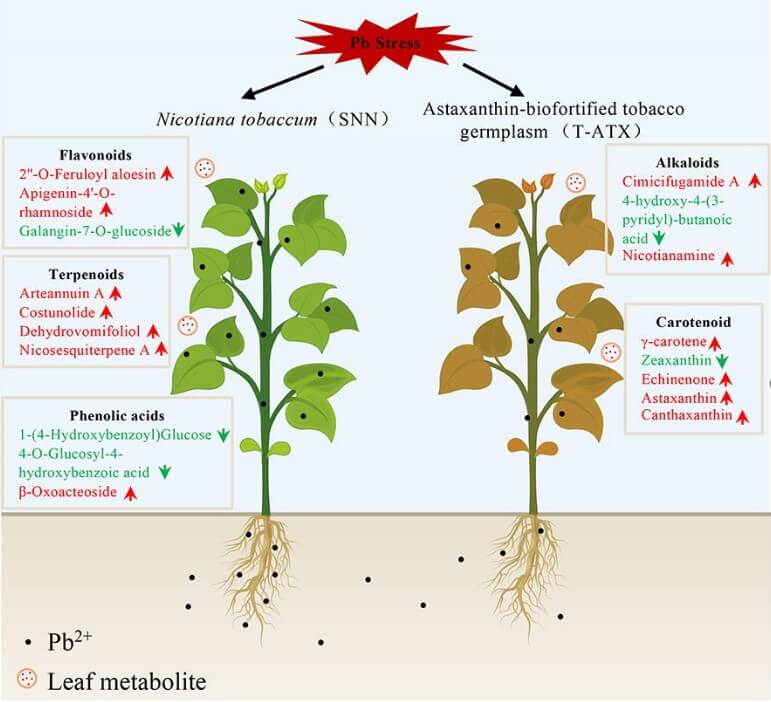
Case Study 1 | Mechanism Discovery for Heavy Metal Stress Tolerance in Tobacco
Published in Journal of Advanced Research (2026), this study investigated how endogenous astaxanthin improves tobacco tolerance to lead stress. Using MetwareBio’s plant widely-targeted metabolomics service based on UPLC-MS/MS platform, the authors identified 1804 metabolites and revealed metabolic changes associated with antioxidant defense, detoxification pathways, carotenoid metabolism, Pb transport, and hormone signaling. Together with physiological and transcriptomic evidence, these results explained the improved growth, lower Pb accumulation, and reduced Pb translocation observed in astaxanthin-biofortified tobacco. This case highlights the value of our Plant Widely-Targeted Metabolomics Service for stress-response metabolomics, mechanism-focused plant research, and pathway-level interpretation in environmental adaptation studies and phytoremediation research.
Case Study 2 | Metabolic Fingerprinting of Jujube Cultivars Across Growing Sites
Published in Plants (2023), this study profiled mature fruits from 11 jujube cultivars grown at three field sites in New Mexico to determine how cultivar and environment shape fruit metabolomes. Using MetwareBio’s LC-MS/MS-based plant widely-targeted metabolomics analysis, the researchers detected 1,315 compounds and showed that cultivar was the dominant factor, while location had a secondary effect. Pairwise comparisons enabled cultivar fingerprinting and revealed characteristic metabolite differences, including cultivar-specific compounds such as sanjoinine A. This case highlights the application value of our Plant Widely-Targeted Metabolomics Service in metabolic fingerprinting, germplasm evaluation and characterization, quality-related trait analysis, and multi-site comparative plant studies for cultivar discrimination and authenticity verification.
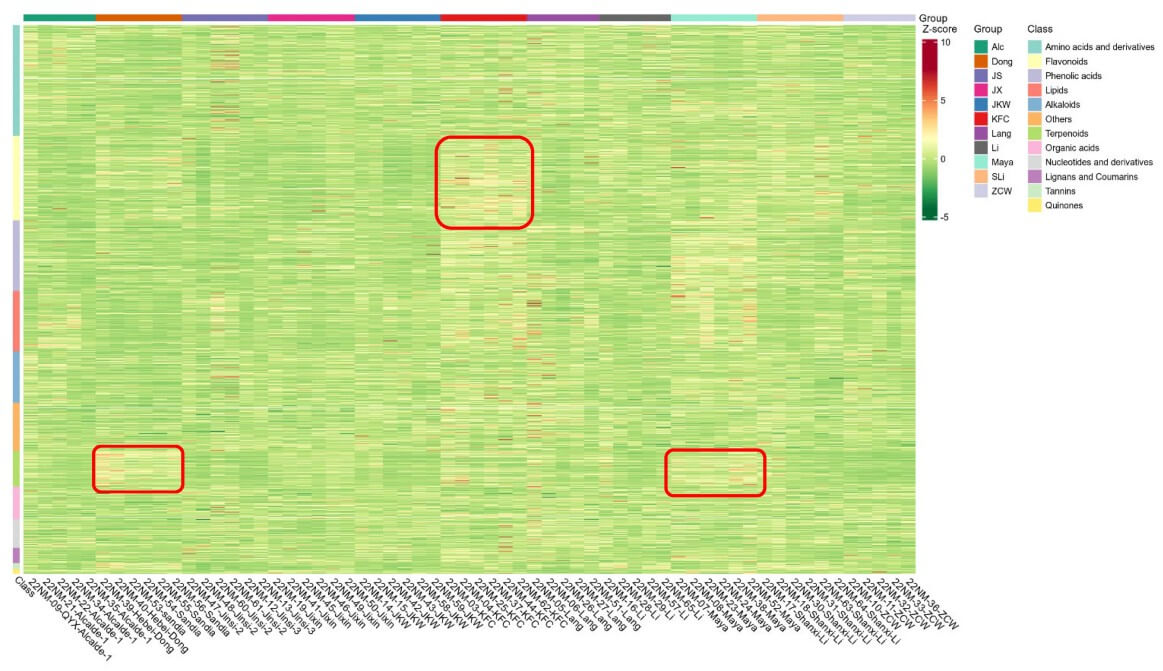

Reference:
1. Du, Z., Liang, M., Wang, X., Liu, Y., Du, S., Shi, D., Sun, Y., Ji, C., Zhang, C., Cui, H., Li, R., & Xue, J. (2026). Astaxanthin biofortification enhances tobacco tolerance to lead stress through boosting antioxidant defense, reducing Pb accumulation, and modulating detoxification pathways. Journal of Advanced Research, 82, 189–211. https://doi.org/10.1016/j.jare.2025.07.038
2. Yao, S., Sapkota, D., Hungerford, J. A., & Kersten, R. D. (2023). Jujube Fruit Metabolomic Profiles Reveal Cultivar Differences and Function as Cultivar Fingerprints. Plants, 12(12), 2313. https://doi.org/10.3390/plants12122313
Sample Requirements for Plant Widely-Targeted Metabolomics Service
| Category | Sample Type | Recommended Amount | Minimum Amount |
| Plant Tissue | Fresh plant tissue (stem, shoot, node, leaf, root, flower, fruit, callus) | 600 mg | 100 mg |
| Freeze-dried plant tissue / powder | 600 mg | 100 mg | |
| Dry samples (seeds, old roots, old stems) | 600 mg | 300 mg | |
| Tobacco samples | 600 mg | 200 mg | |
| Plant-Derived Liquid | Root exudates | 10 mL | 3 mL |
| Extracts, tissue fluid, phloem sap, leaf apoplastic fluid, plant secretions | 500 μL | 200 μL | |
| Juice | 5 mL | 3 mL | |
| Oil / essential oil | 500 μL | 100 μL | |
| Others | Plant glandular trichomes | 50 mg | 15 mg |
- For sample types not listed above, please contact us to obtain specific sample requirements and handling recommendations.
- At least three biological replicates are recommended for each group to ensure reliable statistical analysis and biological interpretation.
FAQs on Plant Widely-Targeted Metabolomics Service
What is the difference between plant widely-targeted metabolomics and untargeted metabolomics?
Plant widely-targeted metabolomics and untargeted metabolomics differ mainly in analytical strategy and instrumentation approach.
- Untargeted metabolomics uses high-resolution mass spectrometry (HRMS), primarily in full-scan DDA mode, to acquire broad spectral signals and support exploratory metabolite discovery without predefined target lists. It is well suited for detecting unknown or unexpected metabolites, but may offer lower quantification accuracy and weaker reproducibility across batches.
- Plant widely-targeted metabolomics combines DDA-based broad metabolite detection with MRM-based targeted quantification on two MS platforms. This dual-mode strategy delivers both discovery capability and quantitative precision in one LC-MS/MS workflow, making it more suitable for studies that require stronger metabolite identification and annotation, more robust quantitative comparison, and higher reproducibility for large-scale sample cohorts.
What are the advantages of plant widely-targeted metabolomics over untargeted metabolomics?
Compared with untargeted metabolomics, plant widely-targeted metabolomics offers more accurate metabolite identification, more reliable quantification, and better reproducibility. It is especially well suited for large-scale plant metabolomics studies, multi-batch sample analysis, and projects that require follow-up targeted validation of selected metabolites.
What quality control measures are used in Plant Widely-Targeted Metabolomics?
Our Plant Widely-Targeted Metabolomics Service includes multiple quality control measures to ensure data reliability and reproducibility, including blank controls, solvent controls, pooled QC samples, internal standards, and randomized injection order. These controls help monitor background interference, extraction stability, instrument performance, and analytical consistency throughout the plant metabolomics workflow.
What data quality assessment results are included in the Plant Widely-Targeted Metabolomics report?
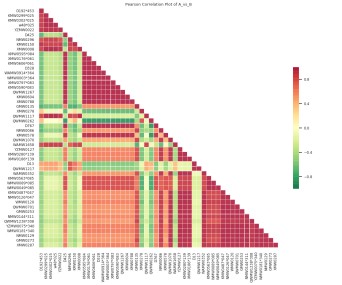
Our Plant Widely-Targeted Metabolomics Service report can include multiple data quality assessment results, such as TIC plots of QC samples, TIC overlap of QC samples, XIC multi-peak plots from MRM detection, CV distribution, PCA, sample clustering, and sample correlation analysis. These evaluations help assess data stability, reproducibility, and overall dataset quality in plant metabolomics studies.
How are missing values handled in plant metabolomics data analysis?
In our plant metabolomics workflow, missing values are typically imputed using one-fifth of the minimum detected value. This commonly used strategy helps maintain dataset integrity for downstream statistical analysis while minimizing distortion in plant metabolite profiling results.
How do you handle batch effects when samples are submitted in multiple batches over time?
For plant widely-targeted metabolomics projects in which samples are submitted in multiple batches over time, we use bridging QC samples for cross-batch signal correction to improve comparability across runs. Combined with standardized sample processing and instrument monitoring, this strategy supports more reliable data integration for large-scale plant metabolomics analysis.
How many biological replicates are recommended for plant metabolomics analysis?
We recommend at least three biological replicates per group to support more reliable statistical analysis in plant metabolomics studies. Additional replicates may further improve confidence when sample variability is high or the experimental design is complex.
Does the service support multi-omics integration analysis?
Yes. In addition to plant widely-targeted metabolomics, we support integrated multi-omics analysis with:
- Transcriptomics (RNA-seq) — linking gene expression changes with metabolite abundance variations
- Proteomics — connecting protein abundance with enzymatic activity and metabolic flux
- Microbiome / Microbiomics — investigating host-microbe metabolic interactions in rhizosphere and phyllosphere contexts
- mGWAS (metabolome Genome-Wide Association Studies) — connecting metabolite quantitative trait loci (mQTLs) with genetic markers for trait association and candidate gene discovery
These integrative approaches enable deeper systems-level insight into plant traits, metabolic pathways, and biological mechanisms underlying complex agronomic phenotypes.
What are the recommended sample preparation, storage, and shipping conditions for plant metabolomics?
For plant metabolomics sample preparation, freshly collected samples should be gently cleaned with clean lint-free paper to remove visible dust, soil, and other surface impurities, then immediately snap-frozen in liquid nitrogen for at least 5 minutes to rapidly quench metabolism. Samples should be stored at -80°C for long-term preservation before shipment. During transportation, samples should be packed with sufficient dry ice to maintain continuous low-temperature conditions throughout transit, helping preserve sample integrity and native metabolite profiles for reliable plant widely-targeted metabolomics analysis.
Next-Generation Omics Solutions:
Proteomics & Metabolomics
Ready to get started? Submit your inquiry or contact us at support-global@metwarebio.com.