Bottom-up vs Top-down vs Middle-down Proteomics: How to Choose the Right Strategy for PTM Analysis
1. Why PTM Analysis Depends on Strategy Choice
Post-translational modifications (PTMs) multiply the functional diversity of the proteome. A single gene product can exist in many molecular forms due to PTMs, sequence variants, processing, and splicing—hence the modern push to think in terms of proteoforms rather than "one protein per gene".
But PTM proteomics is hard for reasons that are structural, not merely technical:
1) PTM site localization is probabilistic, not guaranteed.
Many PTMs (phosphorylation, acetylation, ubiquitination, methylation, glycosylation, etc.) can occur on multiple candidate residues within the same peptide/protein region. MS/MS fragment coverage and scoring determine whether you can confidently assign "which residue is modified", not just "a PTM exists".
2) Multiple PTMs can coexist on the same molecule (combinatorial PTMs / PTM crosstalk).
Biology often encodes regulation through patterns rather than single marks—classic examples include histone tails and signaling proteins with adjacent regulatory sites. Technologies that only measure isolated sites can miss rules encoded by combinations. Reviews on PTM crosstalk emphasize that capturing interactions among PTMs requires methods that preserve connectivity (which PTMs occur together) and not just site lists.
3) Coverage depth vs. protein integrity is a real trade-off.
The more you fragment proteins into peptides, the more identifications you can often achieve (depth, throughput, sensitivity)—but the more you lose information about which PTMs co-occur on the same molecule. Conversely, intact-protein approaches preserve context but pay in throughput and complexity.
That is exactly why PTM proteomics revolves around strategy choice. The three canonical strategies—Bottom-up, Top-down, and Middle-down—sit at different points in the spectrum of depth vs. integrity, and they answer different scientific questions best. A general mass-spectrometry proteomics review perspective also highlights that proteomics is an evolving toolbox whose strengths depend heavily on how workflows are matched to biological questions.
2. Bottom-up Proteomics for PTMs: The Most Mature and Widely Used PTM Strategy
2.1 How Bottom-up PTM Proteomics Works (Workflow)
Bottom-up proteomics analyzes proteins indirectly through peptides:
Protein extraction → enzymatic digestion → peptide separation (LC) → MS/MS → peptide ID → PTM site assignment
Most workflows rely on trypsin (cleaves after Lys/Arg) because it produces peptides in a length/charge range that works well with LC–MS/MS and database searching; other proteases (LysC, GluC, AspN, chymotrypsin, etc.) are often used to improve sequence coverage or access PTM-rich regions.
At the PTM level, bottom-up is typically "site-centric": you identify modified peptides and localize PTMs to specific residues when fragment evidence allows.
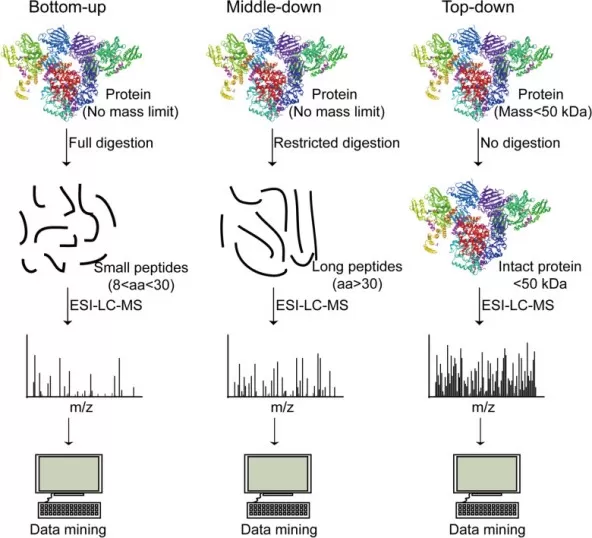
Schematic workflow of bottom-up, middle-down, and top-down proteomics.
Image reproduced from Sun, M., Zhang, X, 2022, Cell & bioscience, licensed under the Creative Commons Attribution License (CC BY 4.0).
2.2 Strengths and limitations of Bottom-up PTM Analysis
1) Strengths:
- High throughput and high sensitivity: Bottom-up is the workhorse for large cohorts, time courses, multi-condition experiments, and statistical designs where sample size matters.
- Excellent compatibility with PTM enrichment: Many PTMs are sub-stoichiometric. Enrichment (IMAC/TiO₂ for phosphopeptides; anti-K-ε-GG for ubiquitin remnants; anti-acetyl-lysine; lectin/HILIC/other for glyco-enrichment, etc.) pairs naturally with peptide-level workflows.
- A mature computational ecosystem: Search engines, localization scoring, FDR control, quantification pipelines, and standardized reporting are far more developed for peptide-centric proteomics than for intact proteoforms. Broad proteomics reviews often describe how the field built large-scale catalogs and quantitative assays largely on the back of bottom-up methods.
2) Limitations:
- Loss of protein context ("connectivity problem"): Digestion breaks the link between distant PTM sites. If phosphorylation at site A only matters when acetylation at site B is present on the same molecule, peptide-only evidence may not be able to prove that co-occurrence.
- Combinatorial PTMs are difficult to reconstruct: You may identify many PTM sites but still not know which combinations coexist on the same proteoform—especially when sites lie on different peptides after digestion. PTM crosstalk reviews repeatedly note that measuring PTM interactions demands methods that can preserve or infer connectivity more directly.
- Protein isoforms and proteoforms can be ambiguous: Bottom-up often faces "protein inference" issues: peptides can map to multiple isoforms, and PTM patterns that define distinct proteoforms can be invisible once the protein is chopped up.
2.3 Typical use cases for Bottom-up PTM Proteomics
Bottom-up is often the best default when your goals look like:
- Large-scale PTM profiling (phosphoproteomics, acetylproteomics, ubiquitinomics) across many samples
- Comparative and quantitative studies (disease vs control, perturbation screens, pathway activation, dose response)
- Biomarker discovery where statistical power and throughput dominate
- Systems biology that needs breadth and quantification more than full proteoform resolution
3. Top-down Proteomics for PTMs: Proteoform-Level, Intact-Protein Analysis
3.1 How Top-down PTM Proteomics Works
Top-down proteomics avoids digestion and measures intact proteins directly:
Protein/proteoform separation → intact MS (precursor) → fragmentation of whole-protein ions → proteoform identification and characterization
Top-down is inseparable from the proteoform concept: it aims to characterize the specific molecular forms of proteins—including combinations of PTMs—rather than inferring them from peptide fragments. The term "proteoform" was proposed to unify how we describe the many molecular forms produced from one gene, including PTM-defined forms.
A widely cited review of top-down proteomics frames the field as proteoform-centric and contrasts it with bottom-up’s peptide-centric strengths and weaknesses.
3.2 Strengths and limitations of Top-down Proteomics
1) Strengths:
- Preserves full PTM combinations on a single molecule: This is the top-down superpower. You can directly observe that this proteoform carries phosphorylation at X, acetylation at Y, and methylation at Z—together.
- Discriminates isoforms and PTM-defined variants: When separation and MS resolving power are sufficient, top-down can distinguish proteoforms that differ by small mass shifts or different modification patterns—critical for mechanistic interpretation and structure–function relationships.
- Strong alignment with functional and structural biology: If your question is "Which specific proteoform interacts, localizes, or drives phenotype?", top-down provides a more direct molecular answer than a site list.
2) Limitations:
- Higher instrument and workflow demands: Intact proteins are harder to ionize, separate, fragment, and interpret than peptides. Sample handling (solubility, fractionation, detergents), chromatography, and MS method optimization are more demanding.
- Lower throughput for complex proteomes: Proteome-wide top-down in highly complex samples remains challenging; the practical throughput is often lower than peptide workflows.
- Bioinformatics complexity: Proteoform identification involves larger search spaces and more complicated scoring, deconvolution, and ambiguity handling than standard peptide searches. Reviews emphasize that while top-down capabilities are advancing, analysis and standardization remain major bottlenecks.
3.3 Typical use cases for Top-down Proteoform Analysis
Top-down is particularly valuable for:
- Low-to-moderate complexity systems (purified proteins, complexes, limited panels, enriched fractions)
- Isoform/proteoform-specific biology (e.g., proteoform-specific signaling, disease-associated variants, PTM-defined functional states)
- Detailed structural/functional characterization where the "whole molecule" matters
- Proteoform mapping projects where discovering intact variation is the goal
4. Middle-down Proteomics for PTMs: A Practical Balance Between Depth and Integrity
4.1 How Middle-down Proteomics Works
Middle-down proteomics sits between bottom-up peptides and top-down intact proteins. The idea is to generate longer peptides (often several kDa) that retain more PTM connectivity than typical tryptic peptides, while being more manageable than intact proteins.
This is typically achieved through:
- Limited digestion (controlled proteolysis)
- Alternative proteases that yield longer fragments (e.g., GluC, AspN in certain configurations, or tailored strategies)
- Chemical cleavage in some contexts
Middle-down has become especially prominent in histone PTM analysis, where dense PTM patterns on N-terminal tails are central to biological interpretation.
4.2 Strengths and limitations of Middle-down PTM Analysis
1) Strengths:
- Partial preservation of combinatorial PTM information: If multiple PTMs lie on the same long fragment, you can observe co-occurrence patterns that bottom-up would split across peptides.
- Often more feasible than full top-down: Middle-down fragments can be easier to separate, fragment, and identify than intact proteins, offering a "sweet spot" for some PTM-dense regions.
- Particularly strong for histone biology: Histone tails (high PTM density, combinatorial regulation) are a canonical target where middle-down provides meaningful connectivity without full intact-protein complexity. Comparative studies have examined the accuracy of middle-down vs bottom-up for histone PTM quantification.
2) Limitations:
- Less standardized and less widely supported computationally: Compared with classic tryptic bottom-up pipelines, middle-down often needs customized methods for fragmentation, deconvolution, and database/search settings.
- Narrower adoption and fewer "plug-and-play" workflows: While powerful in certain domains (histones, PTM-dense regions, specific domains), it is not yet as universally turnkey as bottom-up.
- Still challenging in very complex proteomes: Long fragments can complicate chromatography and MS/MS interpretation, especially without targeted fractionation strategies.
4.3 Typical use cases (Histone Tails, PTM Clusters)
Middle-down is often the best fit for:
- Histone PTMs and epigenetics (capturing PTM patterns on tails)
- High-density PTM regions where connectivity matters (clusters, repeated motifs, regulatory tails)
- Domain-focused studies where you care about PTM combinations within a functional region more than full-length proteoforms
- Bridging studies that want more context than bottom-up but more throughput than top-down
5. How to Choose: Bottom-up vs Middle-down vs Top-down for Your PTM Study
Comparison of Bottom-Up, Top-Down, and Middle-Down Proteomics
|
Dimension |
Bottom-up |
Middle-down |
Top-down |
|
Primary analysis level |
Peptide / PTM site |
Long peptide (multi-kDa fragment) |
Intact protein / proteoform |
|
PTM site localization |
Often strong (when fragments support it) |
Good within long fragments; depends on fragmentation coverage |
Can be strong, but depends heavily on intact fragmentation and interpretation |
|
Combinatorial PTM interpretation |
Limited (connectivity often lost) |
Moderate–strong within the same fragment |
Strongest (proteoform-level combinations preserved) |
|
Throughput |
Highest |
Medium |
Lowest (in complex settings) |
|
Tolerance to sample complexity |
Best |
Moderate |
Most sensitive to complexity |
|
Instrument/workflow demands |
Lowest barrier |
Medium barrier |
Highest barrier |
|
Best "default" for large cohorts |
Yes |
Sometimes |
Rarely |
|
Typical "signature" applications |
Global phospho/acetyl/ubiquitin profiling |
Histone tails, PTM clusters, domain-centric PTM crosstalk |
Proteoform mapping, isoform/variant resolution, mechanistic proteoform biology |
A useful way to choose is to ask what you must not lose:
- If you must not lose throughput (many samples, strong statistics), choose bottom-up.
- If you must not lose PTM connectivity within a region (histone tails, PTM clusters), choose middle-down.
- If you must not lose the intact proteoform definition (isoforms/variants/complete PTM patterns on one molecule), choose top-down.
There is no universally "best" method—only the best match to your biological question, sample constraints, and resources.
6. Future Trends in PTM Proteomics: DIA, AI, and Multi-Strategy Workflows
PTM proteomics is moving toward integration—of strategies, acquisition modes, and computation—because no single approach solves everything.
1) Multi-strategy designs will become more common
A pragmatic pattern is emerging:
- Bottom-up for discovery and cohort-scale quantification
- Middle-down for PTM-pattern validation in dense regions (e.g., histones)
- Top-down for definitive proteoform resolution on key targets
In other words: use bottom-up to find where the action is, then use middle-/top-down to determine what exact molecular forms are responsible.
2) DIA is expanding PTM-scale quantification (and pushing software innovation)
Data-independent acquisition (DIA) is increasingly used for PTM workflows—especially phosphoproteomics—because it can improve consistency and reduce missing values across samples. For example, DIA-based phosphoproteomics strategies and benchmarks have been published in high-impact venues, demonstrating large-scale site-level quantification with systematic computational approaches.
As DIA PTM data grows, open software is also maturing around site-level reporting and quantification (including tools designed specifically for DIA phosphoproteomics reporting).
Key implication: DIA will likely make population-scale PTM proteomics more robust, while raising the bar for PTM site localization models and scoring methods.
3) AI/deep learning will increasingly shape PTM identification and localization
Deep learning has already reshaped peptide-spectrum prediction and spectral library generation. A flagship example is Prosit, which demonstrated proteome-wide prediction of peptide fragment spectra and boosted identification and DIA library workflows.
The next wave focuses more explicitly on modified peptides and PTM-aware prediction/localization (including emerging work aimed at improving PTM site localization and multiply modified peptide detection).
Key implication: Better predictions + better rescoring can increase confidence in PTM localization and enable identification of PTMs that were previously "too rare" or "too ambiguous", especially when paired with DIA.
4) Stronger links between structure, function, and phenotype
The deeper motivation behind top-down and middle-down is not merely "more complete MS data," but more biologically interpretable molecular entities—proteoforms and PTM patterns that can be connected to function.
PTM crosstalk reviews emphasize that understanding how PTMs work together requires approaches that measure interactions and combinatorial patterns, not just independent sites.
That conceptual direction naturally pushes the field toward integrative workflows that preserve more connectivity while still scaling to complex biology.
References:
1. Sun, M., & Zhang, X. (2022). Current methodologies in protein ubiquitination characterization: from ubiquitinated protein to ubiquitin chain architecture. Cell & bioscience, 12(1), 126. https://doi.org/10.1186/s13578-022-00870-y
2. Smith, L. M., Kelleher, N. L., & Consortium for Top Down Proteomics (2013). Proteoform: a single term describing protein complexity. Nature methods, 10(3), 186–187. https://doi.org/10.1038/nmeth.2369
3. Toby, T. K., Fornelli, L., & Kelleher, N. L. (2016). Progress in Top-Down Proteomics and the Analysis of Proteoforms. Annual review of analytical chemistry (Palo Alto, Calif.), 9(1), 499–519. https://doi.org/10.1146/annurev-anchem-071015-041550
4. Sidoli, S., Lin, S., Karch, K. R., & Garcia, B. A. (2015). Bottom-up and middle-down proteomics have comparable accuracies in defining histone post-translational modification relative abundance and stoichiometry. Analytical chemistry, 87(6), 3129–3133. https://doi.org/10.1021/acs.analchem.5b00072
5. Kitata, R. B., Choong, W. K., Tsai, C. F., Lin, P. Y., Chen, B. S., Chang, Y. C., Nesvizhskii, A. I., Sung, T. Y., & Chen, Y. J. (2021). A data-independent acquisition-based global phosphoproteomics system enables deep profiling. Nature communications, 12(1), 2539. https://doi.org/10.1038/s41467-021-22759-z
6. Leutert, M., Entwisle, S. W., & Villén, J. (2021). Decoding Post-Translational Modification Crosstalk With Proteomics. Molecular & cellular proteomics : MCP, 20, 100129. https://doi.org/10.1016/j.mcpro.2021.100129
Next-Generation Omics Solutions:
Proteomics & Metabolomics
Ready to get started? Submit your inquiry or contact us at support-global@metwarebio.com.


