Interactome & Interactomics: Mapping Protein Interaction Networks in Biology and Medicine
Life’s complexity is far greater than the sum of its parts. Over the past two decades, genomics and transcriptomics have made it possible to read biology’s “parts list” at unprecedented scale, but knowing which genes and proteins exist does not explain how they coordinate to produce function. Like listing every instrument in an orchestra without hearing the symphony, a functional understanding requires a map of how molecules “talk” to one another. Interactomics fills this gap by charting interaction networks among biomolecules—proteins, nucleic acids, metabolites, and more—turning static catalogs into dynamic, context-aware functional maps. For biomedical researchers and drug discovery scientists, understanding the interactome, protein interaction networks, and molecular interaction mapping provides a powerful route to connect genotype to phenotype.
What Is Interactomics: From “Single Proteins” to Interaction Networks
Genomics and transcriptomics have revolutionized biology by letting us read the cell’s molecular inventory—what genes exist and which RNAs are expressed. But this “what’s there” view is often not enough to explain how biological function emerges. In most real cellular processes, proteins (and other biomolecules) do not act in isolation: they assemble into complexes, form transient contacts, compete for binding partners, and relay signals through protein interaction networks (PPI networks). These interactions are inherently dynamic and context-dependent, varying across cell types, physiological conditions, and disease states. Understanding how molecules "talk" to one another is therefore essential to move beyond descriptive parts lists toward a mechanistic understanding of life. This recognition gives rise to interactomics: the discipline that systematically maps the molecular dialogue underlying cellular function.
Interactomics is the systems-level study of interactions among biomolecules—including protein-protein, protein-nucleic acid, and protein-metabolite interactions, as well as cell-cell contacts, aiming to capture how biological information is coordinated across molecular and cellular scales. The outcome of this effort is the interactome: a (contextual) map of all interactions in a cell or system—effectively a functional wiring diagram that transforms static catalogs into dynamic, mechanistic understanding (Sharifi Tabar et al., 2022). In practice, interactomics spans both physical interactions (e.g., proteins in the same complex) and broader functional associations that help explain shared pathway roles and regulatory coupling.
Mapping these interaction landscapes is important across biomedicine because interactomics can clarify how disease-associated mutations rewire signaling pathways, pinpoint critical network hubs for drug targeting, and provide a framework for anticipating both on-target effects and potential off-target risks of therapeutic candidates. By shifting biological interpretation from isolated components to integrated systems, interactomics supports a more mechanistic understanding of health and disease at the level of the interactome and protein interaction networks.
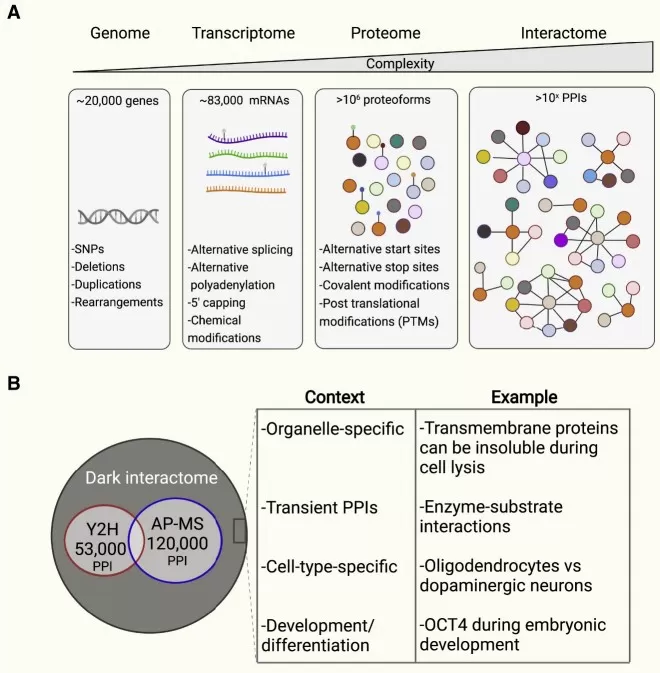
Complexity increases from the genome to the interactome in human cells
Image reproduced from Sharifi Tabar et al., 2022, Cell reports methods.
Molecular Interaction Types and Interactome Mapping Methods
Interactomics encompasses a spectrum of interaction types spanning multiple molecular classes, from protein–protein interactions (PPIs) to protein–metabolite and protein–nucleic acid interactions. Each class differs in stability, affinity, spatial organization, and temporal dynamics—meaning no single assay can capture the full interaction landscape. As a result, interaction mapping typically relies on method combinations tailored to the biology of interest, with MS-centric strategies playing a central role in generating context-aware, mechanistically informative interaction evidence that can be assembled into an interactome.
Protein–Protein Interactions (PPIs): The Backbone of Protein Interaction Networks
PPIs form protein complexes and pathway “wiring,” making them central to any protein interaction network. Modern MS-based PPI discovery typically combines three complementary approaches (Wu et al., 2025).
- Affinity purification–MS (AP-MS): AP-MS enriches a bait protein (via an epitope tag pull-down or antibody-based co-IP) together with its co-purifying partners under relatively mild, near-physiological lysis conditions, followed by LC-MS/MS identification and quantification. Its key strength is capturing stable protein complexes and generating interpretable protein interaction network edges that reflect co-complex membership; it also supports robust comparative interactomics (e.g., treated vs control, WT vs mutant) when paired with quantitative MS designs and appropriate negative controls to enable background subtraction and reduce false positives.
- Proximity labeling proteomics: Proximity labeling uses an engineered enzyme (e.g., BioID/TurboID/APEX) fused to a bait to covalently tag nearby proteins inside living cells, after which labeled proteins are enriched (often via streptavidin) and identified by MS. Because labeling occurs in situ, this approach is particularly powerful for transient, weak, membrane-associated, or spatially restricted interactions that may dissociate during lysis in classic co-immunoprecipitation mass spectrometry workflows; it also adds a “spatial neighborhood” view that is highly relevant for organelle, signaling, and subcellular niche biology—again relying on rigorous controls and statistical filtering to manage background labeling.
- Crosslinking MS (XL-MS): XL-MS introduces chemical crosslinkers that covalently connect amino acid residues that are in close physical proximity, “freezing” interaction geometry prior to MS analysis. Unlike AP-MS or proximity labeling, XL-MS can provide structural constraints—helping distinguish direct contacts from indirect co-complex associations and localizing interaction interfaces—which is valuable for protein complex analysis and integrative structural modeling; it is especially useful when you need mechanistic insight into how complexes assemble, how mutations disrupt binding, or how conformational changes accompany signaling.
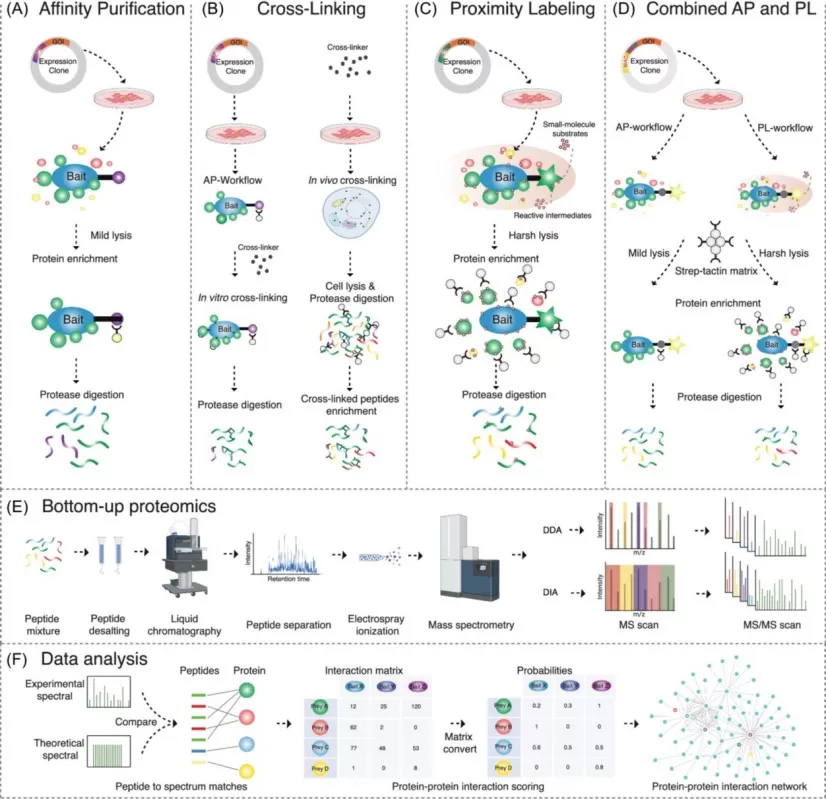
MS-based approaches to studying interactomes.
Image reproduced from Liu et al., 2026, Mass spectrometry reviews, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
Protein–Metabolite Interactions: Linking Chemistry to Phenotype
Metabolites are not merely readouts; they can bind proteins to regulate enzyme activity, signaling, and stress responses. Proteome-scale protein–metabolite interaction (PMI) mapping is challenging because many bindings are transient and condition-dependent. MS-based strategies therefore often combine interaction enrichment or co-fractionation with metabolite-aware readouts, and structural-proteomics concepts such as thermal profiling or limited-proteolysis signatures to detect ligand-induced conformational changes (Wu et al., 2025). The result is a network layer that explains how metabolic state can rewire functional outcomes even when expression changes are modest.
Protein–Nucleic Acid Interactions: Building Regulatory Networks
Protein–DNA/RNA interactions connect genome information to gene regulation. For transcription factors, function depends not only on DNA occupancy but also on co-factors and chromatin-associated complexes. A practical multi-omics pattern is to pair AP MS (to define TF-associated protein partners) with ChIP seq (to map binding sites and discover composite motifs), enabling a unified view of the TF complex and its regulatory footprint (Gabele et al., 2025). This integration moves from isolated binding events to a mechanistic regulatory network that is highly actionable in disease mechanism studies and drug discovery.
Cell–Cell Interactomics: Mapping Physical Cellular Interactions
Traditional interactomics has focused on molecular partnerships within a cell—PPIs, protein–RNA, and protein–metabolite interactions. But many of the most decisive biological “conversations” happen between cells, especially in the immune system, where transient physical contacts (for example, immunological synapses) can determine activation, tolerance, or killing outcomes. A key limitation is that many spatial or imaging-based approaches are not easily scalable to liquid biopsies (blood, PBMCs) and often struggle to capture rare, short-lived contacts at population scale. This gap has driven the rise of cellular interactomics: methods designed to quantify and map physical cell–cell interactions as first-class biological features, rather than inferring communication only from ligand–receptor expression (Wheeler, 2024).
A representative breakthrough is Interact-omics, an ultra-high-scale flow cytometry–based cellular interaction mapping framework that detects physically interacting cells (PICs) while simultaneously phenotyping each partner. In essence, it uses cytometry scatter features to discriminate singlets from multicellular events and then leverages marker co-expression patterns plus clustering to annotate the interacting cell pairs across complex immune landscapes—at speed, low cost, and very high throughput. The Nature Methods study demonstrates how this approach can resolve interaction kinetics and mode of action in immunotherapy settings (including CAR-T and the CD19×CD3 bispecific blinatumomab) and can quantify organism-wide immune interaction rewiring during viral infection in vivo, enabling a genuinely network-level view of cellular immunity (Vonficht et al., 2025).
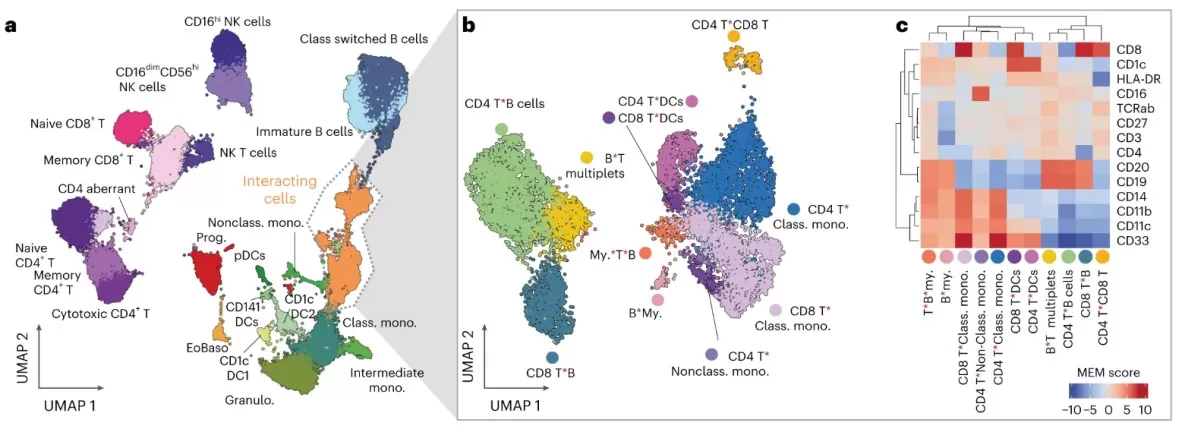
Ultra-high-scale cellular interaction mapping across complex immune landscapes.
Image reproduced from Vonficht et al., 2025, Nature methods, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
Key Interactomics Workflows and Quality Control Considerations
High-impact interactomics relies on a central principle: interaction claims are only as reliable as the controls, quantification strategy, and statistical evidence supporting them. Across AP-MS, proximity labeling proteomics, and other molecular interaction mapping workflows, rigorous experimental design and quality control are what separate biological signal from technical noise.
Experimental Design: Controls, Replicates, and Batch Discipline
Interactomics is particularly susceptible to false positives, since enrichment workflows can co-purify abundant “sticky” proteins or capture promiscuously labeled bystanders. Robust control design is therefore foundational. For AP-MS and co-immunoprecipitation mass spectrometry, recommended controls typically include mock/IgG controls, tag-only controls, and—when feasible—KO/CRISPR or binding-defective mutants that serve as biologically grounded negative controls. For proximity labeling, localization- and expression-matched controls (e.g., enzyme-only or compartment-matched references) are essential to interpret proximity signals as meaningful biological neighborhoods rather than background labeling.
Statistical power and reproducibility depend on biological replicates (not only technical repeats). Batch effects (instrument time, operator, reagent lots) are common in large studies; mitigating strategies include condition randomization across batches, consistent SOPs, and comprehensive metadata capture to enable downstream correction prior to network inference.
Sample Handling and Enrichment: Lysis, Specificity, and Background Subtraction
Sample preparation strongly determines which interaction classes are observable. Mild, non-denaturing lysis tends to preserve stable complexes but may increase nonspecific carryover, whereas stronger detergents or higher-salt conditions can reduce background yet disrupt weak or transient interactions. Method selection is therefore driven by study intent—core protein complex analysis versus dynamic signaling contacts.
Enrichment specificity is equally critical. Antibody quality, bead chemistry, incubation time, wash stringency, and buffer composition collectively shape signal-to-background performance. Practical QC typically tracks bait recovery and recovery of known interactors (when available) as process indicators. In data interpretation, background subtraction using matched controls and confidence-weighted scoring is recommended, particularly for low-abundance baits and samples prone to high nonspecific binding.
MS Acquisition Strategy: DDA vs DIA for Interactome Mapping
Both DDA and DIA proteomics can support interactomics, but they prioritize different strengths. DDA remains common for discovery AP-MS due to mature pipelines and straightforward implementation, yet it can suffer from stochastic precursor selection, reducing reproducibility across many pull-downs or experimental conditions.
By contrast, DIA can improve quantitative consistency across large interaction screens by collecting fragment data more comprehensively, which is advantageous for comparative interactomics where stable quantification is required across dozens to hundreds of samples. Trade-offs include increased computational complexity and the need for careful method optimization (e.g., gradient length, windowing scheme, and library-based versus library-free analysis). In both modes, instrument time and gradient design should match sample complexity: shorter gradients for high-throughput screens and longer gradients for deep interactome coverage with robust FDR control.
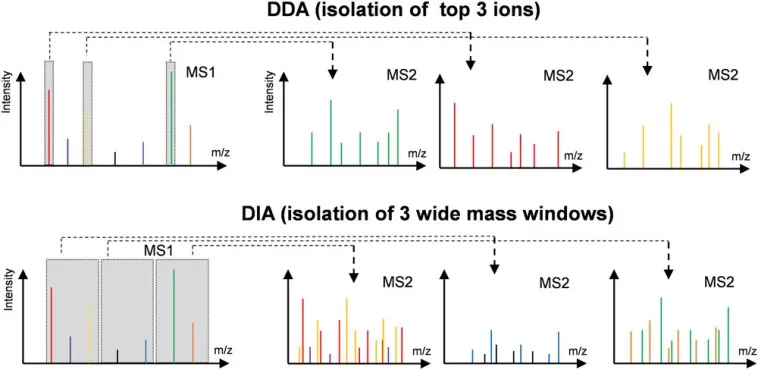
Difference in MS1 isolation windows for the DDA and DIA modes.
Image reproduced from Tian, X., Permentier, H. P., Bischoff, R, 2023, Mass spectrometry reviews, licensed under the Creative Commons Attribution License (CC BY 4.0).
Interactomics Data Analysis and Protein Interaction Network Modeling
Generating interaction data is only the starting point—its scientific and translational value depends on how confidently true interactions are separated from background, and how effectively results are converted into interpretable protein interaction networks and mechanistic hypotheses. Computational biology provides the scoring, statistics, and modeling frameworks needed to control false positives and enable reproducible network inference. It also connects interactomics to other omics layers, turning interaction lists into integrated, machine-readable knowledge that supports discovery at scale.
a) From raw MS data to a high-confidence interaction network
Interactomics analysis typically begins with peptide/protein identification (database search plus validation) and quantitative evidence (e.g., spectral counts or MS1 intensities). Because enrichment-based workflows are vulnerable to nonspecific binding, robust background subtraction against matched negative controls is essential, followed by confidence scoring to prioritize bona fide bait–prey edges and control false positives (often via FDR-aware filtering). Common scoring logic integrates enrichment magnitude, replicate consistency, and control-informed specificity into a confidence-weighted interaction graph that supports downstream complex/module discovery (Wu et al., 2025).
b) Bioinformatics prediction and structural modeling
Interaction lists gain mechanistic depth when paired with structural reasoning—particularly to distinguish direct contacts from indirect co-complex associations. AlphaFold-based multimeric modeling and related assembly strategies can propose plausible interfaces, stoichiometry, and complex topology, helping prioritize interactions for orthogonal validation and enabling more causal interpretation of network “rewiring” across conditions (Bryant et al., 2022).
c) Multi-omics integration and Linked Open Data (LOD)
Interactomics becomes substantially more actionable when linked to transcriptomics/proteomics (expression and abundance changes), pathways, variants, and phenotypes. A growing direction is to represent entities and interaction evidence using semantic web standards—RDF graphs queried by SPARQL—which harmonize identifiers across databases and create machine-readable layers for large-scale mining and machine learning.
Interactomics in Biomedicine: From Networks to Translation
Interactomics delivers practical value when interaction maps are treated as actionable biological evidence—not just descriptive networks. By measuring how complexes, pathways, and cellular neighborhoods change across perturbations, interactomics can connect molecular events to phenotype and translate discovery data into testable hypotheses.
Mechanism of Action (MoA): How perturbations rewire complexes and pathways
Interactomics is increasingly used to explain why a mutation or treatment produces a phenotype by revealing protein complex remodeling and pathway rerouting. For example, an EGFR resistance mutation (T790M) was shown to reconfigure the EGFR interactome, redirecting EGFR trafficking toward an autophagy-linked degradation route that helps cells escape targeted therapy—an MoA insight that directly suggests rational combination strategies (e.g., TKI plus autophagy inhibition) rather than relying on expression changes alone (Wu et al., 2023).
Drug Discovery and Target Safety: On-target vs off-target interaction evidence
In early discovery, interactomics-based target deconvolution can clarify drug-binding landscapes and flag potential safety liabilities. A proximity-enabled strategy (PROCID) profiled compound-associated proteins in living cells and recovered canonical dasatinib targets (ABL1/ABL2) while also identifying additional kinases consistent with broader target engagement (e.g., BTK and CSK), illustrating how interaction-aware proteomics can support on-target/off-target risk assessment and guide follow-up selectivity experiments (Kwak et al., 2022).
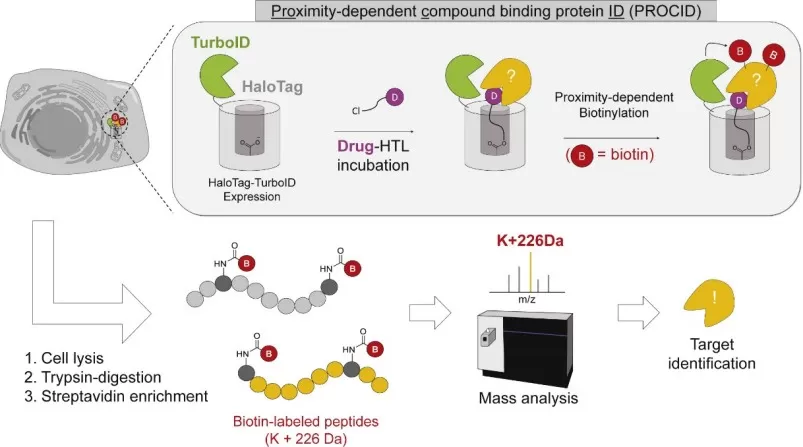
Scheme of PROCID for target-ID in live cells
Image reproduced from Kwak et al., 2022, Cell chemical biology
Disease Mechanisms and Biomarkers: Disease modules and network perturbation signatures
Disease phenotypes often reflect network-level disruption rather than single-gene effects. In a Cell study mapping the Tau interactome in human iPSC-derived neurons, neuronal activity and FTD mutations reshaped Tau’s interaction partners, highlighting synaptic vesicle biology and showing that disease-linked Tau variants reduce interactions with mitochondrial proteins; importantly, mitochondrial components connected to the Tau network were also decreased in Alzheimer’s disease brain cohorts and tracked with disease severity, supporting interactome-guided biomarker and target prioritization (Tracy et al., 2022).
Spatial and Cell-Type-Specific Interactomics: “Location defines function” in vivo
Spatial context can determine which interactions are possible and which pathways dominate. In vivo proximity labeling using a TurboID mouse strategy enabled cell-type-resolved proteomics in the native brain and revealed regional proteomic differences between neuronal and astrocytic programs that are difficult to recover with isolation-based workflows; this provides a practical foundation for extending interactomics toward compartment- and cell-type-specific interaction landscapes that better reflect physiological states (Rayaprolu et al., 2022).
The Future of Interactomics in Systems Biology
Interactomics is reshaping how biological systems are understood by shifting the focus from molecular “parts lists” to the dynamic interaction networks that actually execute function. By resolving how proteins assemble into complexes, how pathways are rewired by perturbations, and how interaction neighborhoods change across contexts, interactomics provides a practical bridge from genotype to phenotype that complements genomics, transcriptomics, and proteomics. At the same time, credible interactome discovery depends on disciplined methodology: well-chosen negative controls, robust replication, careful background subtraction, and statistically grounded confidence scoring are essential for minimizing false positives and generating interpretable protein interaction networks.
Looking ahead, several directions are accelerating the field: higher-throughput interactome mapping (including DIA-enabled scalability), deeper single-cell and cell-type-resolved interaction profiling, and improved strategies for monitoring dynamic interactions in living systems with temporal precision. These advances will increase both biological resolution and translational impact—but also raise the bar for experimental rigor and computational integration. MetwareBio supports this next phase with a professional proteomics and multi-omics service platform, offering comprehensive proteomics analysis and advanced protein complex analysis to help translate complex protein networks and proteome-wide signals into clear, decision-ready biological insights. If this aligns with your research goals, please contact us to discuss your study design and analytical needs.
Reference
1. Sharifi Tabar, M., Parsania, C., Chen, H., Su, X. D., Bailey, C. G., & Rasko, J. E. J. (2022). Illuminating the dark protein-protein interactome. Cell reports methods, 2(8), 100275. https://doi.org/10.1016/j.crmeth.2022.100275
2. Liu, X., Abad, L., Chatterjee, L., Cristea, I. M., & Varjosalo, M. (2026). Mapping protein-protein interactions by mass spectrometry. Mass spectrometry reviews, 45(1), 69–106. https://doi.org/10.1002/mas.21887
3. Wu, S., Zhang, S., Liu, C. M., Fernie, A. R., & Yan, S. (2025). Recent Advances in Mass Spectrometry-Based Protein Interactome Studies. Molecular & cellular proteomics: MCP, 24(1), 100887. https://doi.org/10.1016/j.mcpro.2024.100887
4. Wheeler M. A. (2024). Interactions between immune cells recorded. Nature, 627(8003), 277–279. https://doi.org/10.1038/d41586-024-00426-9
5. Vonficht, D., Jopp-Saile, L., Yousefian, S., Flore, V., Simó Vesperinas, I., Teuber, R., Avanesyan, B., Luo, Y., Röthemeier, C., Grünschläger, F., Fernandez-Vaquero, M., Fregona, V., Ordoñez-Rueda, D., Schmalbrock, L. K., Deininger, L., Yamachui Sitcheu, A. J., Gu, Z., Funk, M. C., Mikut, R., Heikenwälder, M., … Haas, S. (2025). Ultra-high-scale cytometry-based cellular interaction mapping. Nature methods, 22(9), 1887–1899. https://doi.org/10.1038/s41592-025-02744-w
6. Tian, X., Permentier, H. P., & Bischoff, R. (2023). Chemical isotope labeling for quantitative proteomics. Mass spectrometry reviews, 42(2), 546–576. https://doi.org/10.1002/mas.21709
7. Bryant, P., Pozzati, G., Zhu, W., Shenoy, A., Kundrotas, P., & Elofsson, A. (2022). Predicting the structure of large protein complexes using AlphaFold and Monte Carlo tree search. Nature communications, 13(1), 6028. https://doi.org/10.1038/s41467-022-33729-4
8. Wu, P. S., Lin, M. H., Hsiao, J. C., Lin, P. Y., Pan, S. H., & Chen, Y. J. (2023). EGFR-T790M Mutation-Derived Interactome Rerouted EGFR Translocation Contributing to Gefitinib Resistance in Non-Small Cell Lung Cancer. Molecular & cellular proteomics : MCP, 22(9), 100624. https://doi.org/10.1016/j.mcpro.2023.100624
9. Kwak, C., Park, C., Ko, M., Im, C. Y., Moon, H., Park, Y. H., Kim, S. Y., Lee, S., Kang, M. G., Kwon, H. J., Hong, E., Seo, J. K., & Rhee, H. W. (2022). Identification of proteomic landscape of drug-binding proteins in live cells by proximity-dependent target ID. Cell chemical biology, 29(12), 1739–1753.e6. https://doi.org/10.1016/j.chembiol.2022.10.001
10. Tracy, T. E., Madero-Pérez, J., Swaney, D. L., Chang, T. S., Moritz, M., Konrad, C., Ward, M. E., Stevenson, E., Hüttenhain, R., Kauwe, G., Mercedes, M., Sweetland-Martin, L., Chen, X., Mok, S. A., Wong, M. Y., Telpoukhovskaia, M., Min, S. W., Wang, C., Sohn, P. D., Martin, J., … Gan, L. (2022). Tau interactome maps synaptic and mitochondrial processes associated with neurodegeneration. Cell, 185(4), 712–728.e14. https://doi.org/10.1016/j.cell.2021.12.041
11. Rayaprolu, S., Bitarafan, S., Santiago, J. V., Betarbet, R., Sunna, S., Cheng, L., Xiao, H., Nelson, R. S., Kumar, P., Bagchi, P., Duong, D. M., Goettemoeller, A. M., Oláh, V. J., Rowan, M., Levey, A. I., Wood, L. B., Seyfried, N. T., & Rangaraju, S. (2022). Cell type-specific biotin labeling in vivo resolves regional neuronal and astrocyte proteomic differences in mouse brain. Nature communications, 13(1), 2927. https://doi.org/10.1038/s41467-022-30623-x
Next-Generation Omics Solutions:
Proteomics & Metabolomics
Ready to get started? Submit your inquiry or contact us at support-global@metwarebio.com.


