Metabolite Identification in LC-MS Metabolomics: Identification Principles and Confidence Levels
Liquid Chromatography-Mass Spectrometry (LC-MS) metabolomics enables the detection of thousands of metabolic features from complex biological samples, yet transforming these signals into confident metabolite identifications remains the field’s most persistent and rate-limiting challenge. Without rigorous identification, even the most comprehensive datasets yield little more than anonymous molecular features, limiting biological insight and risking irreproducible conclusions.
This guide provides a structured, evidence-based overview of the core principles underlying LC-MS-based metabolite identification and the standardized confidence level framework established by the Metabolomics Standards Initiative (MSI). From exact mass and MS/MS fragmentation to retention behavior and orthogonal CCS data, we detail how each line of evidence contributes to annotation certainty. Understanding this framework is essential for designing robust metabolomics studies, accurately interpreting published results, and clearly communicating the strength of your own identifications.
1. LC-MS Metabolomics: Principles and Metabolite Identification Bottleneck
Liquid chromatography-mass spectrometry (LC-MS) is the dominant analytical platform in metabolomics, offering exceptional sensitivity, resolution, and coverage across diverse chemical classes. Yet the same technological power that enables detection of thousands of metabolic features also gives rise to the field's most persistent bottleneck: confidently translating those signals into defined chemical structures.
1.1 The Power and Promise of LC-MS in Metabolomics
LC-MS achieves its analytical dominance through the synergistic integration of two distinct separation and detection principles.
- Liquid Chromatography (LC): The Separator. Before mass analysis, the complex metabolite extract undergoes LC separation. This step is crucial. It reduces ion suppression (where co-eluting compounds interfere with each other’s ionization) and, most importantly, spreads metabolites out in time based on their chemical properties (e.g., polarity, hydrophobicity). This retention time (RT) becomes the first critical dimension of identification, allowing distinction between compounds that might share an identical mass.
- Mass Spectrometry (MS): The Weigher and Fragmenter. The MS detector performs two key functions. First, the mass analyzer (often a high-resolution instrument like a Q-TOF or Orbitrap) measures the mass-to-charge ratio (m/z) of the ionized metabolites with exceptional precision (often within 5 ppm or less). This provides the exact mass, the primary key for searching molecular formulas. Second, in tandem MS (MS/MS or MS²) mode, the instrument can isolate specific ions and fragment them using collision-induced dissociation (CID), generating a characteristic fingerprint of product ions. This MS/MS spectrum holds the structural clues of the molecule.
Together, LC-MS generates a rich, multi-dimensional dataset: Retention Time, Exact Mass, and Fragmentation Pattern. This triad forms the bedrock of all metabolite identification.
 principles_1772157658_WNo_835d370.webp)
Liquid chromatography–tandem mass spectrometry (LC–MS/MS) principles
Image reproduced from Dewi et al., 2023, Journal of Analytical Science and Technology, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
1.2 The Daunting Challenge of Metabolite Identification
Despite technological advancements, confident identification remains a formidable hurdle. Here are the core challenges:
1) The Universe of Chemical Diversity: The metabolome is estimated to encompass hundreds of thousands of unique structures, spanning a vast range of chemical classes with widely differing properties. No single LC-MS method can capture them all, leading to the need for complementary analytical platforms.
2) The Isomer Problem: Perhaps the most pervasive issue. Isomers—different compounds with the same molecular formula—are ubiquitous in biology (e.g., glucose vs. fructose, or different positional isomers of lipids). They often have identical exact masses and very similar, if not indistinguishable, MS/MS spectra. Relying on MS data alone frequently cannot resolve them.
3) The Standard Dilemma: The gold standard for identification is matching data to an authentic chemical standard run on the same instrument under identical conditions. However, the commercial availability of such standards is a tiny fraction of the metabolome. For novel or rare metabolites, a standard may simply not exist.
4) Spectral Complexity and Database Limitations: Interpreting MS/MS spectra is non-trivial. Fragmentation patterns can be complex and instrument-dependent. While public spectral libraries (e.g., MassBank, GNPS, NIST) are invaluable resources, they are incomplete and can vary in quality. Annotating a metabolite solely by spectral matching to a library entry carries inherent uncertainty unless the match is nearly perfect and supported by other evidence.
In summary, while LC-MS provides the tools to see the metabolome’s “stars,” the process of naming each one—of translating a spectral signature into a verified chemical identity—is a nuanced scientific endeavor fraught with obstacles. This reality makes a systematic framework for reporting identification confidence not just useful, but essential for scientific integrity.
2. Evidence Framework for Metabolite Identification in LC-MS
The process of moving from an unknown peak in a chromatogram to a confidently identified metabolite is a multi-stage investigative workflow. Each stage provides a different type of evidence, and together, they build a cumulative case for identity. Here, we deconstruct the core principles and the hierarchy of information used.
2.1 Evidence Tier 1: Exact Mass and Elemental Composition
The first and most fundamental clue is the exact mass of the ionized molecule ([M+H]⁺, [M-H]⁻, etc.), as measured by a high-resolution mass spectrometer (HRMS).
2.1.1 The Power of High Resolution and Mass Accuracy
High Resolution allows the instrument to distinguish between ions with very close m/z values (e.g., 301.1054 vs. 301.1128). This is critical in complex samples where signal interferences are common.
Mass Accuracy, typically reported in parts per million (ppm), determines how close the measured mass is to the true theoretical mass of a candidate formula. For example, a mass accuracy of < 5 ppm on a 500 Da molecule means the measurement is within ±0.0025 Da of the true value. This high precision dramatically narrows down the list of possible elemental compositions (C, H, N, O, P, S, etc.) that could add up to that mass. Software algorithms use this ppm error to rank plausible formulas.
2.1.2 Isotopic Abundance Patterns: The Natural “Barcode”
Beyond the monoisotopic mass (from the most abundant isotopes, e.g., ¹²C, ¹H), the isotopic pattern of a molecule provides a second, orthogonal filter. Elements like Carbon (with ¹³C), Sulfur (³⁴S), and Chlorine (³⁷Cl) have characteristic naturally abundant heavy isotopes.
The measured relative abundances of the M+1, M+2, etc., peaks in the mass spectrum are compared to the theoretical isotopic distribution of each candidate formula. A correct elemental composition must match not only the exact mass of the main peak but also the full isotopic signature. This step is particularly powerful in ruling out formulas containing elements like Sulfur or multiple Chlorine atoms, which have distinctive isotopic “fingerprints.”
At this stage, we may have a shortlist of possible molecular formulas, but we are far from a structural identification. For example, C₁₇H₂₀N₄O₆ could represent hundreds of different known metabolites.
 10 000 resolving power and zoom of the +2, +_1772157813_WNo_541d705.webp)
Theoretical isotope distribution for celecoxib at (A) 10 000 resolving power and zoom of the +2, +3, and +4 isotope peak at (B) 50 000 and (C) 500 000 resolving power (FWHM), calculated with Thermo XCalibur 3.0
Image reproduced from De Vijlder et al., 2018, Mass spectrometry reviews
2.2 Evidence Tier 2: MS/MS Spectra and Fragmentation Logic
To move from a formula to a structure, we need to break the molecule apart and examine its pieces. This is the role of MS/MS.
2.2.1 Collision-Induced Dissociation (CID) and the Generation of Fragment Ions
In a tandem MS instrument, a specific precursor ion (the metabolite of interest) is isolated and then energized, typically by colliding it with inert gas molecules like nitrogen or argon. This collision imparts internal energy, causing the ion to fragment at its weakest chemical bonds. The resulting product ions (fragments) are then mass-analyzed to produce an MS/MS (or MS²) spectrum. This spectrum is a characteristic “fragmentation fingerprint” of the original molecule under those specific collision energy conditions.
2.2.2 Deciphering the Fingerprint: Neutral Losses and Diagnostic Ions
Expert interpretation of MS/MS spectra can reveal structural motifs. Key clues include:
- Neutral Losses: The mass difference between the precursor ion and a major fragment often corresponds to the loss of a neutral molecule. A loss of 18 Da suggests H₂O (common in alcohols, acids); 162 Da may indicate a hexose sugar (like in flavonoids or glycosides); 44 Da could be CO₂ (from carboxylic acids).
- Diagnostic Product Ions: Specific fragment ions can point directly to a core structure. For example, the presence of a phosphatidylcholine head group fragment at m/z 184.07 is a hallmark of PCs in lipidomics. A phenyl ion at m/z 77 is common in aromatic compounds.
 Product ion spectrum and (B) proposed fragmentation scheme of haloperidol_1772157968_WNo_473d674.webp)
(A) Product ion spectrum and (B) proposed fragmentation scheme of haloperidol
Image reproduced from De Vijlder et al., 2018, Mass spectrometry reviews
While manual interpretation is possible for some classes, identification today heavily relies on spectral library matching. The experimental MS/MS spectrum is computationally compared against databases of reference spectra. A high spectral similarity score (e.g., dot product score) provides strong, though not conclusive, evidence for identity.
2.3 Evidence Tier 3: Orthogonal Evidence-Retention Time, CCS, and Beyond
To address the isomer problem and add independent validation, we must utilize separation-based evidence.
2.3.1 Retention Time (RT): The First Orthogonal Parameter
A metabolite’s interaction with the LC column under standardized conditions is highly reproducible. The retention time is a physicochemical property related to its structure (polarity, hydrophobicity). In identification, RT serves two key purposes:
- Validation: If an authentic chemical standard is available, matching the unknown’s RT to the standard’s RT under identical analytical conditions provides near-definitive proof.
- Prediction/Indexing: In the absence of a standard, predicted RT models (based on molecular descriptors) or publicly available retention time index systems can be used to assess the plausibility of a candidate. A large discrepancy between predicted and observed RT can rule out a formula or structure that matched well by MS alone.
2.3.2 The Emerging Power of Ion Mobility (IM)
Ion Mobility-Mass Spectrometry (LC-IM-MS) introduces a powerful new separation dimension occurring in the gas phase, milliseconds after LC and before MS detection.
Ions are propelled through a drift tube filled with an inert gas under a weak electric field. Larger ions experience more collisions and drift more slowly than compact ones. This measured drift time is converted into a collision cross-section (CCS) value—a reproducible, physicochemical descriptor of the ion’s size and shape.
The CCS value is highly specific for a given molecule and isomer. Matching an unknown’s experimental CCS to a reference CCS (from a standard or a validated database) provides an additional, orthogonal layer of confidence that is exceptionally effective in distinguishing challenging isomers (e.g., lipids, glycans) that have identical masses and very similar MS/MS spectra. It represents the new frontier in high-confidence metabolite annotation.
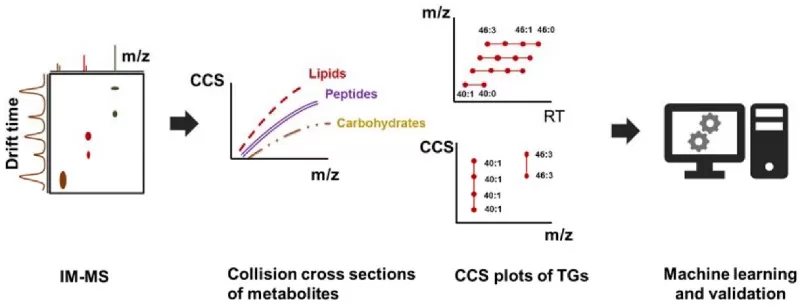
Ion mobility can be used as an additional orthogonal approach to resolve complex mixtures.
Image reproduced from Blaženović et al., 2018, Metabolites, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
3. Identification Confidence Levels in LC-MS Metabolomics
The confidence level associated with a metabolite identification is a critical determinant of its utility in biological interpretation. In the absence of absolute structural confirmation, an unqualified claim of “identification” provides insufficient scientific context, as it fails to distinguish between verification via authentic standards and tentative spectral library matching. This ambiguity directly impacts the reproducibility and comparability of findings. To address this, the Metabolomics Standards Initiative (MSI) introduced a standardized framework for reporting identification confidence levels, which has since been widely adopted or endorsed by major journals in the field. The framework establishes a transparent, evidence-based lexicon for conveying the strength of each annotation, enabling rigorous assessment of data quality and supporting robust cross-study integration. The following levels represent a hierarchy of confidence, from the highest experimental verification to a simple observation.
Level 1: Confidently Identified Compound (The Gold Standard)
Level 1 represents the highest confidence identification, achieved by matching the experimental data from an unknown metabolite to that of an Authentic Chemical Standard analyzed within the same laboratory, on the identical analytical platform, and under the exact same experimental conditions. This rigorous, multi-parameter verification—spanning exact mass, MS/MS spectrum, and retention time (and CCS if applicable)—provides definitive confirmation of the metabolite's structure, offering the utmost confidence short of techniques like NMR. Consequently, Level 1 is the standard for targeted quantification and serves as the gold standard for verifying key discoveries in untargeted studies whenever a pure reference compound is available.
Evidentiary Proof Required:
i. Exact Mass Match: Measured m/z matches the theoretical m/z of the standard within a strict pre-defined error (e.g., < 5 ppm).
ii. MS/MS Spectral Match: The full MS/MS fragmentation pattern (or at least all major diagnostic ions) of the unknown matches that of the standard. A high spectral similarity score is required.
iii. Retention Time (RT) Match: The chromatographic retention time of the unknown aligns with that of the standard (typically within a narrow window, e.g., ± 0.1 min or 2%).
iv. (If available) CCS Match: In LC-IM-MS workflows, the experimental Collision Cross Section value matches that of the standard.
Level 2: Putatively Annotated Compound
Level 2 denotes a putative annotation, where the evidence—typically a high-quality match to an experimental MS/MS spectrum from a public or commercial spectral library—points to a specific compound or a very limited set of structural isomers, but confirmation via an authentic chemical standard analyzed in parallel is lacking. This is the most common confidence level achieved in untargeted metabolomics for novel biomarker discovery; while the spectral evidence is strong and clearly points to a defined molecular structure, the possibility of a closely related isomer that is not distinguishable by the available MS/MS data cannot be entirely ruled out. Transparent reporting at this level should always specify the spectral library used and the corresponding match score.
Evidentiary Proof Required:
i. Exact Mass & Library MS/MS Match: The measured exact mass and, crucially, the MS/MS spectrum show a high-confidence match to a reference spectral library. This could be a public repository (e.g., MassBank, GNPS, HMDB) or a well-curated commercial library (e.g., NIST, mzCloud).
ii. Lack of Standard RT/CCS: The identification lacks the confirmatory RT or CCS match from an in-house standard run on the same system. However, predicted RT or CCS may be consistent, adding supportive evidence.
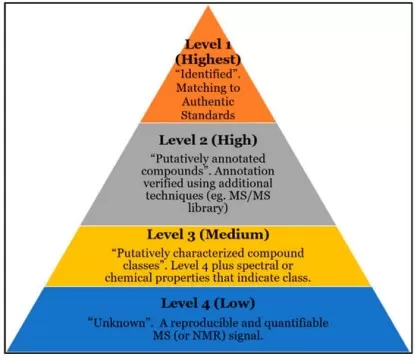
Currently accepted levels of confidence in metabolomics compound identification.
Image reproduced from Reisdorph et al., 2019, Metabolites, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
Level 3: Putatively Characterized Compound Class
This level of confidence is assigned when the available evidence supports assignment to a specific compound class or chemical family, but falls short of pinpointing a precise molecular structure. The annotation relies on diagnostic chemical evidence extracted from the data—such as characteristic neutral losses (e.g., 162 Da indicating a hexose loss, suggestive of a glycoside), diagnostic fragment ions (e.g., ‘m/z’ 184 for phosphatidylcholines), or consistent physicochemical properties like mass defect patterns or retention time behavior specific to a lipid class or other chemical category. Thus, a metabolite may be reported as “a triacylglycerol,” “a sulfated steroid,” or “a dipeptide containing leucine/isoleucine.” While this level provides valuable biological context and is commonly the highest confidence achievable for lipids, novel metabolites, or compounds poorly represented in spectral libraries, it explicitly stops short of full structural identification.
Level 4: Unknown (Yet Distinct) Molecular Feature
At this level, the feature is defined solely by its exact mass and, in some cases, its isotopic pattern, which together enable the prediction of one or more plausible elemental compositions. No interpretable MS/MS spectrum is available—either because it was not acquired, or because the acquired fragmentation data yielded no match to libraries and no recognizable diagnostic pattern. These features are reliably detected and quantified across samples and may exhibit statistically significant changes, making them potentially biologically relevant; however, their chemical identity remains completely unknown. In publications and reports, such unknowns are typically annotated by their precise ‘m/z’ value and retention time (e.g., ‘m/z’ 301.1054 @ RT 8.7 min). They represent candidates for downstream targeted purification, orthogonal analysis, or structural elucidation using advanced techniques such as NMR.
This tiered system provides a common language for the entire metabolomics community. It forces transparency, prevents overstatement, and guides the reader in interpreting the biological significance of results. A pathway analysis based on Level 1 identifications carries far more weight than one based solely on Level 3 annotations.
4. Improving Annotation Confidence: Key Determinants and Best Practices
The theoretical framework of identification confidence levels provides a clear target, but achieving these levels in real-world experiments is contingent upon a cascade of technical, analytical, and strategic decisions. No amount of post-acquisition processing can compensate for poor instrument performance, non-standardized methods, or fragmented analytical workflows. Conversely, even high-quality raw data will fail to yield reliable annotations if interrogated with inadequate reference libraries or uni-dimensional matching strategies. This chapter examines the key factors that govern achievable confidence levels and outlines practical strategies for maximizing annotation reliability throughout the experimental lifecycle.
4.1 Key Drivers of LC-MS Identification Confidence
The upper limit of identification confidence is established first by instrument capability and method standardization. High-resolution mass spectrometers capable of sub-5-ppm mass accuracy are non-negotiable for constraining elemental composition candidates from exact mass measurements; instruments with lower resolving power generate exponentially larger formula sets, rendering downstream filtering inefficient or impossible. Beyond hardware, chromatographic reproducibility directly determines whether retention time can be leveraged as orthogonal evidence. Column chemistry, gradient profile, and temperature control must be rigorously standardized to enable cross-batch comparison or reference standard matching. Equally critical is the standardization of MS/MS acquisition parameters, particularly collision energy regimes, as fragmentation spectra generated under non-standardized conditions exhibit poor inter-laboratory reproducibility and often fail to match library reference spectra.
Parallel to instrumentation, the quality and provenance of reference spectral data establish the definitive ceiling for annotation confidence. Experimentally acquired, curator-validated spectral libraries—such as NIST, MassBank, or GNPS—constitute the only reliable basis for Level 2 putative identifications. In contrast, exclusive reliance on in-silico predicted fragmentation introduces substantial and often unquantifiable uncertainty, as theoretical models remain imperfect predictors of real-world dissociation behavior under specific instrumental conditions.
4.2 Best Practices for Robust LC-MS Metabolite Annotation
While hardware and reference resources constrain the theoretical maximum confidence, the analytical strategy employed determines how closely any given experiment approaches that ceiling. Uni-dimensional approaches that rely solely on exact mass or single-library spectral matching consistently underperform relative to multi-evidence integration workflows. Systematic alignment of exact mass, isotopic fidelity, MS/MS spectral similarity, retention time index or predicted retention behavior, and—where available—experimental or predicted collision cross-section (CCS) values dramatically increases the posterior probability of correct assignment. Implementation of such integrated workflows requires deliberate bioinformatics pipeline design, incorporating parameterized filters for mass error, spectral match thresholds, and retention time deviation tolerances that are applied sequentially or in parallel to rank candidate identifications by cumulative evidence weight.
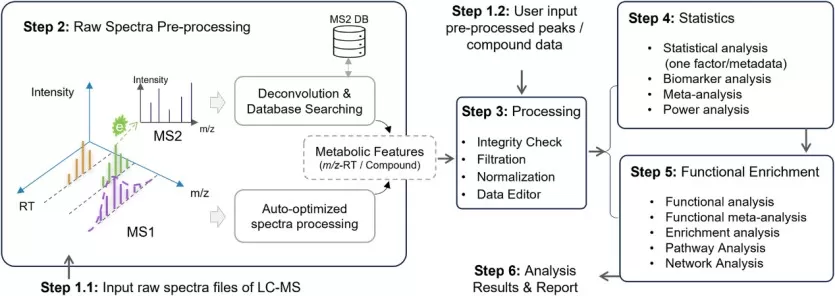
LC-MS untargeted metabolomics workflow based on MetaboAnalystR 4.0.
Image reproduced from Pang et al., 2024, Nature communications, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
At the interpretive stage, the assigned confidence level must function as an explicit guide for biological reasoning. Metabolites designated as Level 1 or Level 2 carry sufficient structural specificity to support pathway mapping, mechanistic hypothesis generation, and biomarker nomination. Level 3 annotations, while useful for indicating broad chemical class involvement, should be interpreted with explicit acknowledgment of their structural ambiguity. Transparent reporting—including quantitative disclosure of the number and proportion of features assigned to each confidence tier—is not merely an editorial requirement but a fundamental component of scientific rigor, enabling reviewers and readers to calibrate their trust in downstream biological claims precisely to the strength of the underlying analytical evidence.
Reference
1. Dewi, K.R., Ismayati, M., Solihat, N.N. et al. Advances and key considerations of liquid chromatography–mass spectrometry for porcine authentication in halal analysis. J Anal Sci Technol 14, 13 (2023). https://doi.org/10.1186/s40543-023-00376-3
2. De Vijlder T, Valkenborg D, Lemière F, Romijn EP, Laukens K, Cuyckens F. A tutorial in small molecule identification via electrospray ionization-mass spectrometry: The practical art of structural elucidation. Mass Spectrom Rev. 2018;37(5):607-629. doi:10.1002/mas.21551
3. Blaženović I, Kind T, Ji J, Fiehn O. Software Tools and Approaches for Compound Identification of LC-MS/MS Data in Metabolomics. Metabolites. 2018;8(2):31. Published 2018 May 10. doi:10.3390/metabo8020031
4. Reisdorph NA, Walmsley S, Reisdorph R. A Perspective and Framework for Developing Sample Type Specific Databases for LC/MS-Based Clinical Metabolomics. Metabolites. 2019;10(1):8. Published 2019 Dec 21. doi:10.3390/metabo10010008
5. Pang Z, Xu L, Viau C, et al. MetaboAnalystR 4.0: a unified LC-MS workflow for global metabolomics. Nat Commun. 2024;15(1):3675. Published 2024 May 1. doi:10.1038/s41467-024-48009-6
Read more
- LC-MS VS GC-MS: What's the Difference
- LC vs. HPLC vs. UHPLC: Tracing the Evolution of Chromatographic Techniques
- Mastering Chromatography: Everything You Need to Know
- Metabolomics Batch Effects
- Analytical vs. Semi-Preparative vs. Preparative HPLC: A Strategic Guide to Precision, Scale, and Efficiency
- A Comprehensive Guide to Quantitative Lipidomics: Methodologies, Workflows, and Applications
Next-Generation Omics Solutions:
Proteomics & Metabolomics
Ready to get started? Submit your inquiry or contact us at support-global@metwarebio.com.


