Plant Metabolomics in Growth and Development: Metabolic and Hormonal Networks
Plant growth is often explained as a chain of command: hormones trigger transcriptional programs, proteins adjust enzyme activities, and organs develop. Metabolomics completes this story at the point where biology becomes chemistry. Metabolites are not just “end products”; they are the immediate substrates, signals, and constraints that determine whether cells can divide, expand, and differentiate. Because metabolite pools respond quickly to environment and internal status, they also feed back to reshape hormone biosynthesis, transport, perception, and signaling sensitivity. Metabolomics therefore offers a direct bridge from genotype and environment to phenotype, especially for developmental processes that change across time and tissues [1].
1. Plant development as a coupled system of metabolic flux and hormone signaling
Development is best viewed as a coupled system: hormones act as global coordinators, while metabolism provides the resources and local signals that decide what is feasible in each tissue. A classic example is the interplay between sugars and strigolactones (SLs). Sugars do not merely provide energy; they also act as signaling molecules that inform the plant whether branching, flowering, or senescence programs are affordable. When carbon is limiting, SL signaling can reinforce architectural restraint; when carbon is abundant, sugar signals can shift developmental decisions toward growth. Seeing this coupling requires measuring both metabolic status (e.g., sucrose, hexoses, organic acids, amino acids) and the relevant hormonal layer in the same experimental design [2].
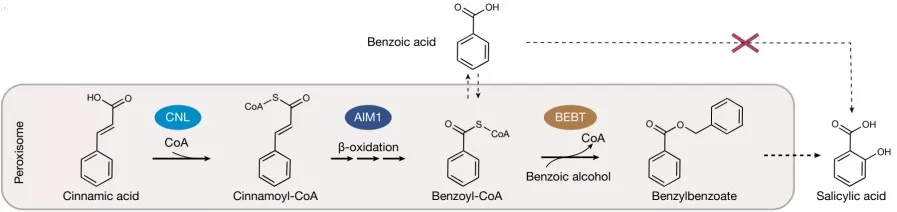
Understanding of plant SA biosynthetic pathways
Image reproduced from Wang et al., 2025, Nature, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
A mechanistic “carbon gate” for growth has been demonstrated in Arabidopsis, where glucose–TOR signaling promotes the accumulation of the brassinosteroid (BR) transcription factor BZR1, aligning growth programs with carbon availability. In developmental terms, TOR integrates carbon status with hormone outputs: when glucose is sufficient, TOR activity supports BR signaling output through BZR1, enabling expansion and anabolic metabolism; when carbon is scarce, reduced TOR activity prevents an energetically costly growth commitment. This is the kind of integrative node that metabolomics can diagnose: metabolite and hormone trajectories reveal when carbon status permits hormone-driven growth, and when feedback restrains it [3].
Beyond these “physiology-scale” hubs, core metabolites can also act as second-messenger-like links between hormone signaling and epigenetic control. Intermediates such as S-adenosylmethionine (SAM) and acetyl-CoA connect metabolic state to methylation and acetylation reactions, thereby influencing gene regulation in a developmental context. Metabolomics helps make such links testable by quantifying whether developmental transitions coincide with shifts in these key metabolic currencies [4-5].
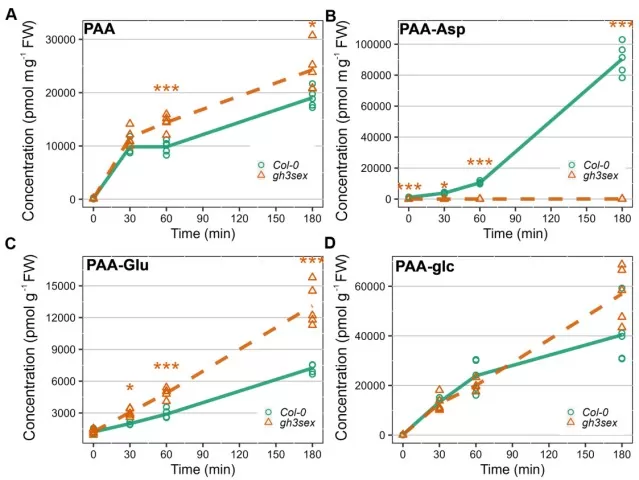
Phenylacetic acid metabolism in Arabidopsis
Image reproduced from Hladík et al., 2025, J Exp Bot, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
2. Why metabolomics is critical in developmental research
Metabolomics is uniquely powerful for plant development for four practical reasons. First, it narrows the genotype–phenotype gap by providing a biochemical readout that integrates genetics, environment, and developmental stage. This is particularly valuable in population studies or breeding contexts, where metabolite variation can point to functional loci and pathways that are not obvious from transcript abundance alone [6].
Second, metabolomics reveals hormone–metabolite interaction hubs rather than one-way cascades. For instance, salicylic acid (SA) biosynthesis is not only a defense pathway; its flux intersects with primary metabolism (phenylalanine supply, shikimate pathway activity, redox state), which can change with developmental stage. Metabolomics can quantify precursor–product relationships and help distinguish whether SA increases are driven by precursor availability, enzyme regulation, or broader metabolic reprogramming across tissues [2].
Third, metabolomics strengthens multi-omics integration because metabolite changes are biochemically interpretable anchors for network models. When combined with transcriptomics and proteomics, metabolite patterns help prioritize which pathways are truly active, and which gene expression changes are likely consequential. Computational frameworks for systems metabolomics emphasize that robust integration benefits from pathway- and network-based approaches grounded in metabolites [7].
Fourth, spatial context often decides causality in development. The same metabolite can indicate growth in one zone and defense in another, and bulk extraction can average away decisive gradients. Spatially resolved approaches, or at minimum careful tissue dissection matched to developmental anatomy, can therefore prevent misinterpretation and reveal “where” metabolic control is exerted [8-9].
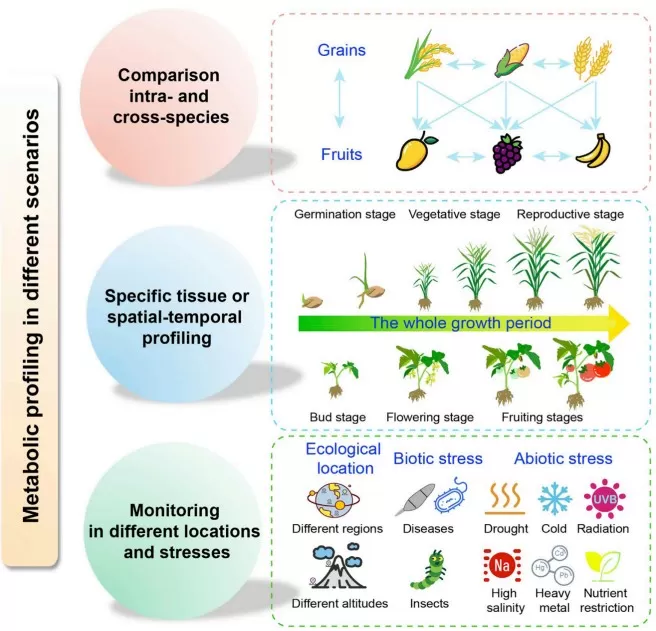
Metabolic profiling in different scenarios
3. Technical strategies in developmental metabolomics: from global scanning to precision targeting
A strong developmental metabolomics study typically progresses from broad coverage to targeted validation. Widely targeted metabolomics (large-panel MRM) is especially useful for developmental series because it balances breadth with cross-batch quantitative stability—critical when comparing multiple stages, tissues, or genotypes. It supports consistent measurement of central metabolites (sugars, amino acids, organic acids, lipids) and specialized metabolites that drive developmental traits such as color, waxes, and volatiles [1, 10].
Hormone profiling is the complementary “signaling layer.” Parallel quantification of ABA, auxin, cytokinin, gibberellins, jasmonates, SA, ethylene-related metabolites, BRs, and SL-associated markers turns descriptive metabolite lists into mechanistic hypotheses about regulatory directionality. This is particularly important in developmental phenotypes that reflect coordinated shifts across multiple hormone classes rather than a single dominant signal [11].
Spatial metabolomics (including imaging and tissue-resolved workflows) adds a third dimension by mapping metabolite distributions onto tissue coordinates. In wheat, drought-associated cuticle traits illustrate why this matters: β-diketones and related wax components can accumulate in specific epidermal layers, and these localized patterns contribute to water-loss control and stress adaptation. Without spatial or tissue-aware sampling, such structure–function links can be missed or diluted [9].
Finally, multi-omics integration is how correlation becomes mechanism. Metabolomics combined with genomics, transcriptomics, and proteomics can link “enzyme → metabolite flux → hormone balance → phenotype.” Reproducible pipelines also matter when datasets scale; standardized processing and mining tools help ensure that developmental conclusions are not artifacts of batch effects or missingness [12].
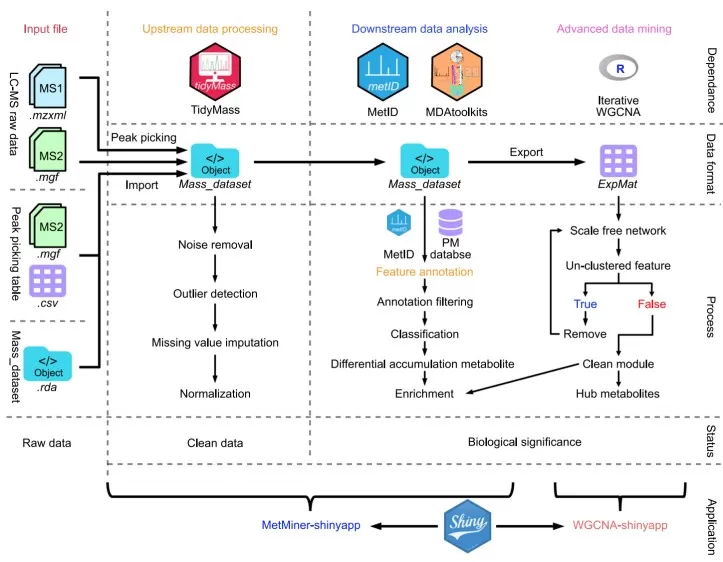
The architecture of MetMiner pipeline
Image reproduced from Wang et al., 2024, J Integr Plant Biol, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
4. Applications of metabolomics across the plant life cycle: five case-driven playbooks
Germination and Seedling Establishment
Germination is a rapid commitment decision: seeds must switch from dormancy maintenance to intense energy demand. A practical hypothesis is that ABA and GA provide the core push–pull signal, while sugar availability and energy status determine whether the hormonal instruction can be executed. In Arabidopsis, the glucose–TOR–BZR1 axis offers a concrete example of carbon gating: when glucose supports TOR activity, BR signaling output through BZR1 is promoted and growth proceeds; when carbon is limited, this gate restricts growth. A publishable design uses a dense time course (imbibition → radicle emergence → cotyledon expansion) with widely targeted metabolites plus hormone profiling to capture short-lived pulses that would be missed in two-timepoint comparisons [3].
Root Architecture and Branching
Root branching is shaped by local developmental programs that integrate nutrient signals and hormone gradients, especially auxin. Post-translational regulation of auxin transport and cross-talk with other hormones can rewire where auxin accumulates, which in turn changes lateral root initiation. Metabolomics adds the missing layer by measuring the chemical environment that accompanies these hormonal shifts, including amino acids, organic acids, and phenylpropanoid-related metabolites that can influence signaling and microbe interactions. A representative mechanistic example is the endophytic fungus Serendipita indica, which alters auxin distribution partly through changes in auxin transport and conjugation, promoting host growth. Pairing hormone profiling with metabolite panels helps identify whether auxin homeostasis is shifting via conjugation dynamics, precursor supply, or downstream metabolic rewiring across root zones [8].
Floral Transition and Organ Development
Flowering is not only an environmental timing decision; it is also a resource decision. When sugar status is favorable, plants can execute reproductive development without compromising survival. A practical metabolomics design profiles carbohydrates and central carbon metabolites in source leaves alongside hormone panels in apices and developing floral tissues, then uses network analysis to identify which metabolic states predict the transition. Because flowering involves tissue-specific decisions at meristems, incorporating tissue-resolved sampling (or spatial methods when feasible) can clarify whether key metabolites accumulate at the decision site or merely reflect systemic status [3].
Fruit/grain development and quality formation
Quality traits—flavor, color, aroma, nutrition—are chemical phenotypes determined by how carbon and nitrogen are partitioned during development. Bitter gourd provides a clear example of how metabolomics becomes actionable: comprehensive resources integrating genome-scale information with metabolite profiling can connect fruit traits to biosynthetic pathways and candidate genes, enabling metabolite-assisted breeding or functional validation. In practice, the most informative designs use a dense developmental series (not just “early vs late”), include multiple genotypes to capture natural variation, and quantify both primary metabolites (sugars, acids, amino acids) and specialized metabolites (pigments, volatiles). The output is a metabolite–gene association map that links developmental stage to quality formation trajectories [5].
Senescence and nutrient remobilization
Senescence is a regulated allocation program rather than simple deterioration. It involves coordinated hormone dynamics (notably SA and ethylene, with interactions from other hormones) and large-scale shifts in primary metabolism that move carbon and nitrogen from aging leaves to sinks. Metabolomics can identify senescence markers such as amino acid surges, lipid turnover products, and organic acid changes, and can help disentangle whether stress accelerates senescence through hormone shifts, metabolic depletion, or both. In soybean, mass-spectrometry-based metabolomics has been used to decode signaling and defense responses under biotic stress, offering patterns that can be aligned with developmental progression to distinguish defense-induced metabolic reprogramming from age-driven remobilization [10].
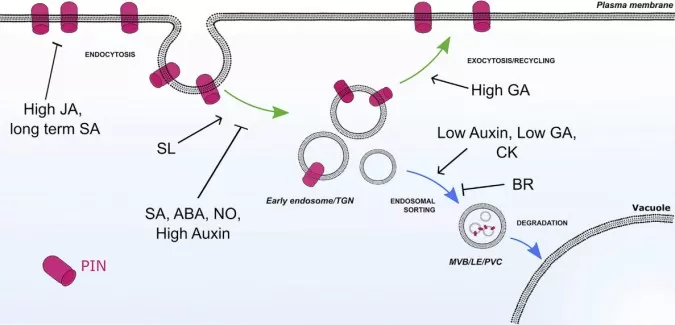
Hormonal Regulation of PIN Trafficking in Plant Cells
Image reproduced from Semeradova et al., 2020, Plant Commun, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
5. Conclusion: from metabolite profiles to mechanistic insights in plant development
Metabolomics has become a central pillar of systems plant biology because development is executed by dynamic metabolite pools and fluxes embedded in bidirectional hormone networks. When metabolite panels, hormone profiling, spatial context, and multi-omics integration are designed around developmental transitions, the result is not merely “many metabolites changed,” but a mechanistic narrative with testable control nodes—such as carbon gating through TOR, auxin homeostasis shaped by transport and conjugation, or tissue-localized wax remodeling under drought. A network-first approach turns developmental metabolomics into a practical engine for discovery and trait improvement [6].
References
1. Rai M, Dutta M, Saito K, Rai A. A deep dive into plant metabolomics: Milestones, technologies, and translational impact. Plant Physiol. 2025 Nov 26;199(4):kiaf408. doi: 10.1093/plphys/kiaf408.
2. Wang Y, Song S, Zhang W, Deng Q, Feng Y, Tao M, Kang M, Zhang Q, Yang L, Wang X, Zhu C, Wang X, Zhu W, Zhu Y, Cao P, Chen J, Pan J, Feng S, Chen X, Dai H, Song S, Yang J, Zhao T, Cao F, Tao Z, Shen X, Last RL, Hu J, Yu J, Fan P, Pan R. Deciphering phenylalanine-derived salicylic acid biosynthesis in plants. Nature. 2025 Sep;645(8079):208-217. doi: 10.1038/s41586-025-09280-9. Epub 2025 Jul 23.
3. Zhang Z, Zhu JY, Roh J, Marchive C, Kim SK, Meyer C, Sun Y, Wang W, Wang ZY. TOR Signaling Promotes Accumulation of BZR1 to Balance Growth with Carbon Availability in Arabidopsis. Curr Biol. 2016 Jul 25;26(14):1854-60. doi: 10.1016/j.cub.2016.05.005.
4. Hladík P, Brunoni F, Žukauskaitė A, Zatloukal M, Bělíček J, Kopečný D, Briozzo P, Ferchaud N, Novák O, Pěnčík A. Phenylacetic acid metabolism in land plants: novel pathways and metabolites. J Exp Bot. 2025 Aug 21;76(12):3427-3443. doi: 10.1093/jxb/eraf092.
5. Zhang Y, Wang X, Wang Y, Niu Y, Han X, Tan J, Ma X, Zhang Q, Liu J, Zhang Z, Gao W, Liang W, Li L, Liu Z, Zhang X, He H, Yang Y, Yu R. Telomere-to-telomere genome assembly and comprehensive mutation library facilitate bitter gourd breeding and functional genomics research. Plant Commun. 2026 Feb 16:101779. doi: 10.1016/j.xplc.2026.101779.
6. Shen S, Zhan C, Yang C, Fernie AR, Luo J. Metabolomics-centered mining of plant metabolic diversity and function: Past decade and future perspectives. Mol Plant. 2023 Jan 2;16(1):43-63. doi: 10.1016/j.molp.2022.09.007.
7. Krumsiek J, Bartel J, Theis FJ. Computational approaches for systems metabolomics. Curr Opin Biotechnol. 2016 Jun;39:198-206. doi: 10.1016/j.copbio.2016.04.009.
8. González Ortega-Villaizán A, King E, Patel MK, Pérez-Alonso MM, Scholz SS, Sakakibara H, Kiba T, Kojima M, Takebayashi Y, Ramos P, Morales-Quintana L, Breitenbach S, Smolko A, Salopek-Sondi B, Bauer N, Ludwig-Müller J, Krapp A, Oelmüller R, Vicente-Carbajosa J, Pollmann S. The endophytic fungus Serendipita indica affects auxin distribution in Arabidopsis thaliana roots through alteration of auxin transport and conjugation to promote plant growth. Plant Cell Environ. 2024 Oct;47(10):3899-3919. doi: 10.1111/pce.14989.
9. Bi H, Luang S, Li Y, Bazanova N, Morran S, Song Z, Perera MA, Hrmova M, Borisjuk N, Lopato S. Identification and characterization of wheat drought-responsive MYB transcription factors involved in the regulation of cuticle biosynthesis. J Exp Bot. 2016 Oct;67(18):5363-5380. doi: 10.1093/jxb/erw298.
10. Silva E, Belinato JR, Porto C, Nunes E, Guimarães F, Meyer MC, Pilau EJ. Soybean Metabolomics Based in Mass Spectrometry: Decoding the Plant's Signaling and Defense Responses under Biotic Stress. J Agric Food Chem. 2021 Jul 7;69(26):7257-7267. doi: 10.1021/acs.jafc.0c07758.
11. Wang X, Liang S, Yang W, Yu K, Liang F, Zhao B, Zhu X, Zhou C, Mur LAJ, Roberts JA, Zhang J, Zhang X. MetMiner: A user-friendly pipeline for large-scale plant metabolomics data analysis. J Integr Plant Biol. 2024 Nov;66(11):2329-2345. doi: 10.1111/jipb.13774.
12. Semeradova H, Montesinos JC, Benkova E. All Roads Lead to Auxin: Post-translational Regulation of Auxin Transport by Multiple Hormonal Pathways. Plant Commun. 2020 Apr 22;1(3):100048. doi: 10.1016/j.xplc.2020.100048.
13. Rekhter D, Lüdke D, Ding Y, Feussner K, Zienkiewicz K, Lipka V, Wiermer M, Zhang Y, Feussner I. Isochorismate-derived biosynthesis of the plant stress hormone salicylic acid. Science. 2019 Aug 2;365(6452):498-502. doi: 10.1126/science.aaw1720.
Next-Generation Omics Solutions:
Proteomics & Metabolomics
Ready to get started? Submit your inquiry or contact us at support-global@metwarebio.com.


