Protein Subcellular Localization: Why a Protein's "Workplace" Matters
Proteins are the fundamental drivers of cellular life, orchestrating essential biological processes, including metabolism and signaling. However, their functions are not solely dictated by their molecular structures; rather, they are closely tied to their precise locations within the cell. Recent research underscores that protein subcellular localization is meticulously regulated and plays a crucial role in determining protein function, both in health and disease [1]. This spatial regulation ensures that proteins interact with the correct partners and substrates at the right time, forming the backbone of cellular organization. For example, a protein located in the cytoplasm may facilitate metabolic reactions, while one in the nucleus may regulate gene expression. Mislocalization—such as relocating a protein from the mitochondria to the cytosol—can disrupt essential pathways, resulting in cellular dysfunction.
Understanding protein localization is critical for unraveling how cells maintain homeostasis and adapt to environmental cues. Despite this, several fundamental questions remain: How do proteins navigate to specific compartments? What happens when localization goes awry? This blog aims to explore these questions, emphasizing the significance of pinpointing a protein’s “workplace” to drive advancements in biomedical research. By examining fundamental concepts, functional relationships, disease links, and cutting-edge techniques, we will uncover how spatial proteomics—the study of protein localization and dynamics at the subcellular level—is reshaping our understanding of cell biology and paving the way for novel therapeutic strategies.
1. Fundamental Concepts of Protein Subcellular Localization
Protein subcellular localization refers to the specific cellular compartment or region in which a protein resides and carries out its functions. This positioning defines the protein’s environment and its interaction network, ultimately shaping its molecular activities. For example, a protein in the endoplasmic reticulum (ER) may assist in protein folding and quality control, while one in the mitochondria may contribute to energy production. Thus, a protein’s molecular functions are inherently tied to its subcellular context, as this environment controls its access to binding partners, substrates, and regulatory mechanisms [2]. Without accurate localization, proteins cannot perform their tasks effectively, potentially leading to cellular imbalances.
Common cellular compartments include the nucleus, cytoplasm, mitochondria, ER, Golgi apparatus, lysosomes, peroxisomes, and plasma membrane—each with specific, critical roles. The nucleus is responsible for DNA replication and transcription, acting as the cell’s control center. The cytoplasm, which houses various organelles, supports metabolic processes, protein synthesis, and cellular signaling. Mitochondria, often referred to as the “powerhouses” of the cell, are central to energy production and apoptosis regulation. The ER plays a key role in protein synthesis, lipid metabolism, and calcium storage, with the rough ER specifically involved in synthesizing proteins for secretion or membrane integration. The Golgi apparatus modifies, sorts, and packages proteins and lipids for transport to their final destinations, including secretion or incorporation into the cell membrane. Lysosomes serve as the cell’s waste disposal system, breaking down macromolecules and cellular debris. Peroxisomes are involved in detoxification and fatty acid metabolism, while the plasma membrane regulates the movement of substances in and out of the cell. These compartments are not static—proteins can shuttle between them based on cellular needs.
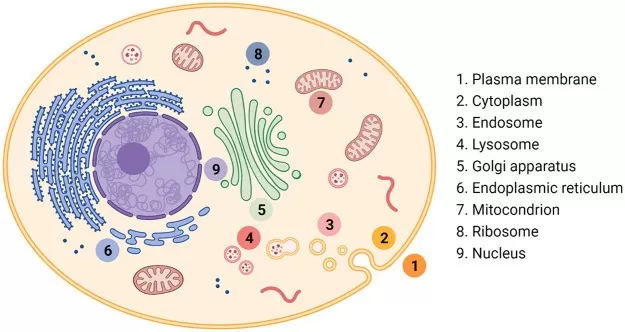
Key cellular compartments.
Image reproduced from Zhou, Z., Maxeiner, K., Ng, D. Y. W. et al., 2022, Polymer Chemistry in Living Cells, licensed under the Creative Commons Attribution License (CC BY 4.0).
The mechanisms guiding protein localization are intricate, often involving intrinsic targeting signals. Signal peptides or specific sequences direct nascent proteins to organelles like the ER or mitochondria during synthesis. Post-translational modifications (PTMs), such as S-acylation, alter protein hydrophobicity to influence trafficking, localization, and interactions [3]. These modifications ensure that proteins respond dynamically to cellular signals, highlighting the precision of spatial regulation. Even small errors in this process can have cascading effects, disrupting cellular functions. Building on this foundation, we now explore how localization directly impacts protein functionality.
2. Relationship Between Protein Subcellular Localization and Function
A protein’s subcellular location dictates its functional repertoire, with spatial segregation enabling distinct roles. Membrane-bound proteins, for instance, often mediate cell signaling and adhesion, while nuclear proteins are involved in transcriptional regulation. This compartmentalization allows for specialized functions: membrane proteins facilitate ion transport and receptor-mediated signaling, impacting processes like cell cycle regulation and endocytosis [2]. In neurons, axonal proteins are critical for synapse maintenance and mRNA localization, with RNA-binding proteins such as TDP-43 and FUS regulating local translation for neuronal survival [4]. When these proteins are mislocalized, as seen in amyotrophic lateral sclerosis (ALS), axonal integrity is compromised, illustrating how location-specific functions are essential for cellular health.
Localization is not fixed; it is dynamic and can change in response to environmental stress or cellular states. During stress responses, proteins may translocate to specialized compartments for sequestration. For example, misfolded proteins are spatially sequestered as part of cellular quality control mechanisms, which are vital for maintaining proteome integrity and cellular health [5]. Such dynamic relocalization can occur during cell division or in pathological conditions, like cancer, where treatment-induced shifts alter protein interactions and signaling networks [2]. Studies have revealed complex architectures with single-cell variations, dynamic translocations, and proteins localizing to multiple compartments, allowing cells to adapt to stimuli like DNA damage or nutrient deprivation. This adaptability ensures cellular resilience but also introduces vulnerabilities when dysregulated. This fluidity sets the stage for examining how mislocalization contributes to disease.
3. When Localization Goes Wrong: Protein Subcellular Localization and Disease
Abnormal protein localization is a hallmark of several diseases, particularly neurodegenerative disorders and cancer. Protein misfolding, often leading to mislocalization, is associated with a growing list of human diseases, including Alzheimer’s, Parkinson’s, ALS, and various cancers [5]. In neurodegenerative diseases, misfolded proteins accumulate in intra- and extracellular inclusions, such as amyloid plaques, disrupting proteostasis—the cellular machinery that maintains protein homeostasis. For instance, the spatial sequestration of misfolded proteins is essential for cellular management, but failures in this process contribute to pathogenesis in protein aggregation-linked diseases, such as frontotemporal dementia (FTD). In cancer, altered localization can drive oncogenic signaling, as proteins that typically reside in the nucleus may shift to the cytoplasm, activating proliferation pathways or evading tumor suppression.
Understanding these mislocalizations opens up new possibilities for therapeutic strategies. Targeting specific subcellular compartments could correct abnormal protein behavior. For example, in diseases involving proteostasis collapse, enhancing quality control mechanisms in the ER or mitochondria could prevent toxic protein accumulations [5]. Similarly, drugs designed to modulate protein trafficking or PTMs (e.g., S-acylation inhibitors) could restore normal localization, as seen in experimental treatments for lipid signaling disorders [3]. Spatial proteomics studies have already leveraged comparative analyses to identify disease-specific relocalizations, which can serve as biomarkers for drug development. This knowledge not only informs compartment-specific therapies but also underscores the potential of personalized medicine, where treatments are tailored to subcellular anomalies. Advanced research methods will be essential to achieve these insights, as discussed next.
4. Techniques for Studying Protein Subcellular Localization
Accurately mapping protein localization requires a combination of classical and modern methodologies, each offering unique advantages for spatial resolution.
Classical Methods
Traditional techniques like immunofluorescence staining and subcellular fractionation remain foundational. Immunofluorescence employs antibodies against protein biomarkers to visualize their distribution within fixed cells, providing detailed spatial maps of complex tissues. Subcellular fractionation involves physically separating cellular compartments followed by biochemical analysis, enabling protein isolation from specific organelles. A robust protocol for this includes steps from fractionation to mass spectrometry (MS) sample preparation, allowing for proteome-wide profiling of protein localization across various cell lines. These methods, though straightforward, form the basis for validating spatial data.
Modern Technologies
Recent technological advancements have significantly improved our ability to study protein subcellular localization, with mass spectrometry (MS) emerging as a cornerstone of spatial proteomics. MS-based techniques are particularly valuable for identifying and quantifying proteins from isolated cellular compartments. By combining subcellular fractionation with MS, researchers can generate detailed proteome-wide maps of protein localization across cellular structures. MS offers deep analysis and the capacity to process complex samples, enabling the detection of thousands of proteins from specific organelles, such as the nucleus or mitochondria, in a single experiment [6].
To achieve higher spatial resolution, MS-based workflows can be coupled with Laser Microdissection (LMD), including Laser Capture Microdissection (LCM), to precisely isolate tissue regions from intact samples prior to proteomic analysis. This integration allows for localized investigations into cellular events. Additionally, single-cell sorting, through methods like fluorescence-activated cell sorting (FACS), isolates specific cell populations, enabling high-resolution proteomic profiling at the single-cell level. This is particularly valuable when studying heterogeneous tissues, where cellular variations have significant biological implications.
_1770688450_WNo_744d548.webp)
Overview of Laser Capture Microdissection (LCM).
Image reproduced from Welch, E., Tripathi, Anubhav, 2021, licensed under the Creative Commons Attribution License (CC BY 3.0).
Moreover, single-cell imaging has emerged as a complementary technique. Using technologies such as confocal microscopy and super-resolution microscopy, researchers can visualize protein distributions at cellular and subcellular resolutions. These imaging methods are critical for studying dynamic processes, like protein trafficking and interactions, in living cells. However, single-cell imaging generally offers lower proteome coverage than MS. Therefore, combining imaging with MS provides a more comprehensive approach: imaging offers spatial context, while MS provides quantitative, proteome-wide data.
Together, MS, laser capture microdissection, single-cell sorting, and single-cell imaging are revolutionizing spatial proteomics. Their integration enables unprecedented insights into how proteins are localized and how their spatial organization influences cellular function and disease progression.
Techniques for Studying Protein Subcellular Localization
|
Technique/Tool |
Description |
Purpose |
|
Immunofluorescence |
Uses antibodies and fluorescent dyes to visualize proteins in fixed cells. |
To map protein distribution in tissues or cells. |
|
Subcellular Fractionation |
Separates cellular compartments followed by mass spectrometry analysis. |
To isolate proteins from specific organelles for detailed analysis. |
|
Laser Microdissection (LMD/LCM) |
Isolates tissue sections using a laser for subsequent mass spectrometry analysis. |
To precisely localize proteins within specific tissue regions. |
|
Mass Spectrometry (MS) |
Identifies and quantifies proteins, often after fractionation. |
To create proteome-wide maps of protein localization. |
|
Single-Cell Sorting (e.g., FACS) |
Isolates specific cell populations for single-cell proteomic analysis. |
To study protein localization in heterogeneous cells. |
|
Single-Cell Imaging (e.g., Confocal & Super-Resolution Microscopy) |
High-resolution imaging techniques to visualize protein distribution in individual cells. |
To visualize protein localization and dynamics in living cells. |
Data Analysis and Integration
The large volume of spatial data generated by spatial proteomics requires sophisticated bioinformatics tools for effective interpretation. Mass spectrometry (MS)-based proteomics workflows, coupled with machine learning techniques, enable the analysis of complex datasets to predict and validate protein localizations. Quantitative proteomics methods, which can work with minute protein quantities—down to the single-cell level—offer powerful tools to track dynamic changes in protein localization. Techniques such as differential analysis provide insights into how protein localization shifts under various cellular conditions or stimuli.
Integrating imaging data with proteomics further enhances our ability to create detailed, spatially resolved profiles. For instance, combining immunostaining with MS enables researchers to map the distribution of specific proteins across cellular compartments, adding another layer of context to the proteome-wide analysis. These integrated approaches empower scientists to uncover regulatory networks, cellular signaling pathways, and disease-specific protein behavior. By aligning data from imaging and proteomics, spatial proteomics is poised to accelerate our understanding of cellular processes and offer insights into previously uncharted biological landscapes.
Advances in bioinformatics are also facilitating the integration of multi-omics data, where proteomics data is combined with genomics, transcriptomics, and other layers of molecular information. This systems-level approach allows researchers to better understand the interplay between protein localization and other molecular processes, providing a more comprehensive view of cellular function and dysfunction in disease states.
5. Future Outlook: Where Spatial Proteomics Is Headed
In summary, protein subcellular localization is a pivotal factor in cellular function, influencing everything from metabolism to disease progression. As we’ve seen, understanding a protein’s “workplace” is critical—not just for basic science but also for advancing medical research. The dynamic nature of protein localization, including its shifts in response to stress or disease, plays a vital role in maintaining proteostasis and cellular fitness.
Looking ahead, spatial proteomics is set to transform the field of cell biology and medical research. We are entering an exciting era where this field will integrate seamlessly with other biological disciplines, providing unbiased, systems-level insights into cellular processes. Future advancements will likely focus on increasing resolution and throughput. For example, the combination of multiplexed imaging with artificial intelligence (AI) for real-time analysis of tissue architecture holds great potential. This approach could revolutionize diagnostics and clinical applications, such as biomarker discovery for body fluids and the development of targeted therapies for diseases caused by mislocalization.
As technologies such as single-cell and spatial omics continue to mature, they will offer unprecedented maps of cellular heterogeneity. These advancements will deepen our understanding of biology, unlocking new opportunities for personalized medicine. Ultimately, spatial proteomics promises to bridge the gap between basic research and clinical practice, accelerating progress toward precision medicine by providing the insights needed for more effective, tailored treatments.
References:
1. Lundberg, E., & Borner, G. H. H. (2019). Spatial proteomics: a powerful discovery tool for cell biology. Nature reviews. Molecular cell biology, 20(5), 285–302. https://doi.org/10.1038/s41580-018-0094-y
2. Arslan, T., Pan, Y., Mermelekas, G., Vesterlund, M., Orre, L. M., & Lehtiö, J. (2022). SubCellBarCode: integrated workflow for robust spatial proteomics by mass spectrometry. Nature protocols, 17(8), 1832–1867. https://doi.org/10.1038/s41596-022-00699-2
3. Azizi, S. A., Kathayat, R. S., & Dickinson, B. C. (2019). Activity-Based Sensing of S-Depalmitoylases: Chemical Technologies and Biological Discovery. Accounts of chemical research, 52(11), 3029–3038. https://doi.org/10.1021/acs.accounts.9b00354
4. Piol, D., Robberechts, T., & Da Cruz, S. (2023). Lost in local translation: TDP-43 and FUS in axonal/neuromuscular junction maintenance and dysregulation in amyotrophic lateral sclerosis. Neuron, 111(9), 1355–1380. https://doi.org/10.1016/j.neuron.2023.02.028
5. Sontag, E. M., Samant, R. S., & Frydman, J. (2017). Mechanisms and Functions of Spatial Protein Quality Control. Annual review of biochemistry, 86, 97–122. https://doi.org/10.1146/annurev-biochem-060815-014616
6. Dong, Z., Wu, C., Chen, J., Jiang, W., Piatkevich, K. D., Zhu, Y., & Guo, T. (2025). Filter-aided expansion proteomics for the spatial analysis of single cells and organelles in FFPE tissue samples. Nature protocols, 10.1038/s41596-025-01256-3. Advance online publication. https://doi.org/10.1038/s41596-025-01256-3.
7. Zhou, Z., Maxeiner, K., Ng, D. Y. W., & Weil, T. (2022). Polymer Chemistry in Living Cells. Accounts of chemical research, 55(20), 2998–3009. https://doi.org/10.1021/acs.accounts.2c00420
8. Welch, E. & Tripathi, Anubhav. (2021). Preparation of Tissues and Heterogeneous Cellular Samples for Single-Cell Analysis. https://www.intechopen.com/chapters/78769
Next-Generation Omics Solutions:
Proteomics & Metabolomics
Ready to get started? Submit your inquiry or contact us at support-global@metwarebio.com.


