PTM Proteomics Analysis Workflow: Variable Modifications, Site Localization, and FDR Control
Post-translational modifications (PTMs) shape protein activity, stability, localization, and interaction networks, making PTM-centric proteomics essential for decoding signaling and regulatory biology. However, PTM analysis is inherently challenging: modified peptides are often low-abundance, PTM-bearing spectra can be complex, and confident site localization requires rich and correctly assigned fragment-ion evidence. Conventional data-dependent acquisition (DDA) workflows further compound these issues due to stochastic precursor sampling, which can limit PTM coverage, reduce reproducibility, and introduce missing values that complicate downstream statistics. In contrast, data-independent acquisition (DIA) captures fragment information in a systematic and unbiased manner across the m/z range, enabling more consistent detection and quantification of modified peptides across samples and cohorts [1]. This article distills three practical pillars of DIA-based PTM analysis:
(i) intelligent strategies for setting variable modifications,
(ii) probability-based algorithms for PTM site localization,
Using Spectronaut as an illustrative example, the discussion connects parameter choices to underlying scoring logic, so readers can move from raw DIA files to interpretable, statistically defensible PTM site tables with high confidence.
1. DIA vs DDA: Why DIA Is Better for PTM Proteomics
DDA is widely used for global proteomics, but PTM proteomics pushes DDA to its limits. Because DDA selects precursors for fragmentation in a top-N manner, the set of fragmented ions can vary from run to run, especially in complex matrices or when PTM-bearing peptides compete with abundant unmodified peptides. In practice, this stochastic sampling increases the risk of missing low-intensity modified peptides, reduces cross-sample reproducibility, and produces sparsity that undermines quantitative comparisons in large cohorts. DIA addresses these limitations by repeatedly fragmenting all precursors within predefined isolation windows across the full mass range, thereby creating a near-comprehensive record of peptide fragmentation patterns for each run. This systematic acquisition improves quantitative completeness and supports retrospective mining—new PTM hypotheses can be tested later by re-interrogating the same DIA data with updated libraries, search settings, or localization filters. For large-scale PTM studies that prioritize reproducibility, depth, and cohort-level statistics, DIA has therefore become a preferred acquisition strategy [2]. (Learn more at: DDA vs. DIA: The Essential Guide to Label-Free Quantitative Proteomics)
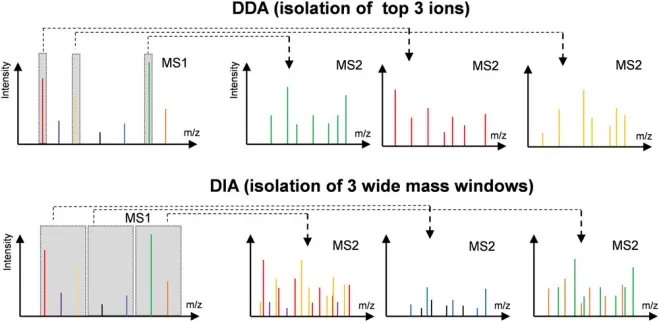
Difference in MS1 isolation windows for the DDA and DIA modes.
Image reproduced from Tian, X., Permentier, H. P., Bischoff, R, 2023, Mass spectrometry reviews, licensed under the Creative Commons Attribution License (CC BY 4.0).
2. Choosing the Right PTM Analysis Software
Software choice is not a cosmetic decision in PTM proteomics—it directly shapes identification sensitivity, site localization confidence, and the practicality of scaling to many samples. The most important differentiators include whether a workflow relies on spectral libraries or supports library-free searching, how it balances discovery of unexpected modifications versus quantification of known PTM types, and how rigorously it models fragment-ion evidence for site localization. The comparison below summarizes commonly used tools and the typical contexts in which each excels.
PTM Proteomics Software Comparison
|
Software Tool |
Acquisition Mode |
Core Strategy for PTM Detection |
Advantages |
Typical Use Cases |
|
DIA |
Spectral-library-based targeted extraction and quantification; supports DirectDIA |
Commercial benchmark; strong quantitative reproducibility; mature PTM workflows |
High-precision quantification of expected PTMs; cohort studies |
|
|
DIA-NN |
DIA |
Neural-network-assisted prediction and scoring; supports library-based and library-free modes |
High sensitivity and speed; strong performance in complex backgrounds |
Fast screening; large cohorts; limited library availability |
|
FragPipe (MSFragger) |
DDA / DIA |
Open search and mass-shift detection with PTM-centric annotation modules |
Powerful for unexpected PTMs; flexible discovery; automation via PTM-Shepherd |
Novel PTM discovery; multi-PTM landscapes; non-model organisms |
|
PEAKS |
DDA / DIA |
Integrated de novo sequencing combined with database search |
Useful when sequence variants/unknown peptides are common; supports discovery-oriented workflows |
Non-model organisms; engineered proteins; samples with many unknown features |
|
Mainly DDA (DIA support in newer versions) |
Database search with modification-specific scoring and LFQ |
Established ecosystem; benchmark LFQ workflows |
DDA-based PTM studies; legacy datasets and pipelines |
Spectronaut is one of the most commonly used environments for DIA-based PTM proteomics, largely because it brings modification-aware identification, localization scoring, and site-level reporting into a single, coherent workflow. Its parameterization also mirrors the decisions that matter in most PTM studies: how variable modifications define the search space, how fragment-ion evidence supports a specific site assignment, and how multi-level FDR control determines which sites remain statistically credible. The terminology may be Spectronaut-specific, but the logic behind these choices translates well across modern PTM proteomics pipelines.
3. Variable Modifications in DIA PTM Workflows
Variable modifications define which PTMs the search engine is allowed to consider. While broad PTM inclusion can increase discovery potential, every added variable modification expands the combinatorial search space, which can reduce sensitivity, increase computational burden, and elevate false positives if not paired with appropriate constraints. A practical strategy is to prioritize biologically relevant PTMs and control complexity through limits on the number of variable modifications per peptide, careful enzyme settings, and rigorous downstream filtering.
3.1 Core Parameters: Modifications and Mass Tolerance
(1) Spectral library construction: In the Library module, select the workflow for generating a spectral library and import the FASTA sequence database.
(2) Modification definition: Fixed modifications should represent processing steps that occur consistently, such as carbamidomethylation of cysteine (Carbamidomethyl, +57.021 Da, C) after iodoacetamide alkylation. Variable modifications represent dynamic biology—examples include phosphorylation (+79.966 Da on S/T/Y) or lysine acetylation (+42.011 Da on K). For complex PTM analyses, multiple variable modifications can be added simultaneously.
(3) Mass accuracy settings: For high mass accuracy instruments (e.g., Orbitrap), it is strongly recommended to set precursor and fragment mass tolerances to Dynamic. This allows Spectronaut to adjust tolerance windows dynamically based on data quality, achieving better identification performance.
(4) PTM localization criteria: Set PTM localization filters in “PTM Settings”. The core metric is Localization Probability, which quantifies confidence that a modification is assigned to a specific amino acid residue. The default threshold is typically set to 0.75; sites with probabilities above this value are classified as high-confidence “Class I” sites.
(5) Quantification consolidation: In parameters related to “PTM Workflow”, set “PTM Consolidation” to “Sum”. This setting automatically sums quantitative signals from different peptides corresponding to the same protein and the same modification site (e.g., different charge states, different enzymatic cleavage termini), and outputs quantitative results directly at the “protein ID_amino acid site” level (PTM.CollapseKey), greatly simplifying downstream data analysis.
3.2 Practical Enhancements: DirectDIA, Normalization, and Site Reporting
PTM quantification is most interpretable when it is placed in the context of total protein abundance. Recent Spectronaut versions support PTM site signal normalization using whole-proteome information, enabling a protein-level adjustment that separates changes in modification occupancy from changes in protein expression. Practically, this can be implemented by selecting protein abundance estimates derived from unmodified peptides (or from all peptides, depending on study design) as an input normalization strategy.
Library construction strategy also matters. Classical DIA PTM workflows often depend on DDA-derived libraries, which can add cost and time and may not capture all PTM-bearing peptides present in DIA runs. DirectDIA reduces this dependency by enabling library creation directly from DIA data via a FASTA-based search engine, simplifying the experimental design and enabling faster iteration for PTM panels. Regardless of the library route, PTM-aware reporting should preserve traceability: site tables should retain peptide sequences, charge states, retention times, fragment-ion evidence summaries, and localization probabilities, so that high-impact findings can be audited and validated by targeted follow-up experiments.
4. PTM Site Localization: From Fragment Ions to Probability
Identifying a modified peptide is not the same as localizing the modification site. Many PTMs can occur on multiple residues within a peptide, and different site assignments can produce nearly identical precursor masses. Reliable localization therefore depends on whether fragment ions provide unambiguous evidence that the mass shift resides on a specific residue. Spectronaut implements a probabilistic site localization framework that integrates confirming and refuting fragment-ion signals into a localization probability score.
4.1 Localization Workflow: Candidates and Ion Evidence
(1) Feature detection and candidate generation:
Spectronaut applies a probability-based site-localization framework that starts with three-dimensional feature detection across m/z, retention time, and intensity. This step defines coherent precursor/fragment signal traces and assembles all plausible peptide–fragment ion relationships. Based on the peptide sequence and the specified PTM rules, the algorithm then systematically enumerates every feasible modification-site arrangement (i.e., all candidate site combinations) for downstream scoring [3], as illustrated in the figure below.
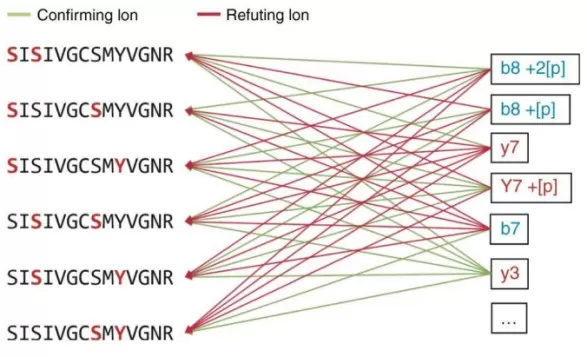
Confirming and refuting fragments for site localization.
Image reproduced from Bekker-Jensen et al., 2020, Nature Communications, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
(2) Classification of fragment-ion evidence:
For each candidate site arrangement, the algorithm interrogates the observed fragment ions and determines whether each ion supports or contradicts the proposed localization. This converts raw MS/MS evidence into site-informative constraints.
- Confirming ions: Fragment ions whose cleavage positions and observed mass shifts are consistent with a specific residue carrying the modification. In practice, when a backbone cleavage occurs such that the resulting b/y ion contains the modified residue, the expected mass shift on that ion provides direct support for that site assignment.
- Refuting ions: Fragment ions that are inconsistent with the candidate site assignment. For example, if an ion that should carry the modification under a given hypothesis is observed without the expected mass shift, or an ion that should not carry the modification is observed with a shift, that evidence weakens (refutes) the proposed localization.
(3) Multi-dimensional ion scoring:
After evidence is labeled as confirming or refuting, each fragment ion is assigned a weighted score that reflects its reliability and quantitative consistency. The scoring typically integrates three orthogonal dimensions:
Spectral feature quality: e.g., signal-to-noise ratio and peak-shape symmetry
Mass accuracy: deviation between observed and theoretical m/z
Chromatographic correlation: co-elution behavior and retention-time alignment across traces
4.2 Localization Probability: Scoring and Thresholds
With fragment-ion evidence scored, the algorithm converts candidate-level support into a residue-level probability that is directly interpretable as localization confidence.
First, compute the score for each candidate site combination. The score for a given candidate equals the total support from confirming ions minus the total contradiction from refuting ions:
Score(candidate) = Σ(score of confirming ions) − Σ(score of refuting ions).
Next, translate these candidate scores into the probability that a specific residue is modified. This is done by summing the scores of all candidates that place the modification on the residue of interest, and dividing by the sum of scores across all candidates:
PTM localization probability = Σ(candidate scores including the site) / Σ(all candidate scores).
The resulting probability ranges from 0 to 1. In many workflows, a threshold such as ≥ 0.75 is used to define high-confidence localizations (often termed Class I sites).
The figure below illustrates how candidate scores derived from spectrum matching are aggregated and normalized to yield site-level localization probabilities.
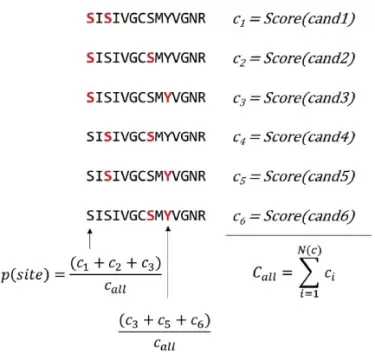

PTM localization probability Algorithm
Image reproduced from Bekker-Jensen et al., 2020, Nature Communications, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
5. FDR Control for Reliable PTM Identification and Site Reporting
PTM datasets can be deceptively persuasive: a long list of sites may look comprehensive, yet even a modest false positive rate can translate into many incorrect sites when thousands of candidates are tested. False discovery rate (FDR) control provides a statistical safeguard by estimating the expected proportion of incorrect calls among accepted identifications. In PTM proteomics, FDR must be considered not only for peptide identification, but also for modification site assignment, because incorrect localization can create biologically misleading conclusions even when the peptide itself is correctly identified.
Spectronaut uses the classic target–decoy database strategy to estimate FDR:
i. Construct a combined database containing real protein sequences (targets) and corresponding reversed or randomized sequences (decoys).
ii. Run the search against this combined database using exactly the same search parameters and scoring rules for both targets and decoys.
iii. Treat decoy hits as false positives, because any match to an artificial decoy sequence is, by definition, incorrect.
iv. Estimate the false-positive rate in target identifications by counting decoy matches, then tune the score cutoff until the decoy-to-target relationship corresponds to the preset FDR threshold.
For PTM analysis, FDR control should be applied independently at multiple reporting levels. If any layer is left uncontrolled, false positives can propagate—most importantly into the final site list—reducing the interpretability and biological reliability of PTM conclusions. Recommended stringent criteria are as follows:
- PSM-level FDR: ≤ 1% (initial filtering for each spectrum match)
- Peptide-level FDR: ≤ 1% (aggregating evidence across PSMs for the same peptide)
- Protein-level FDR: ≤ 1% (protein inference and reporting control)
- PTM site-level FDR: ≤ 1% (most critical)
Independent PTM site-level FDR control means the software treats modification sites as statistical entities in their own right and re-applies target–decoy-based estimation at the site reporting stage. This additional layer of control helps ensure that the final reported “protein_site” list reflects not only confident peptide identifications, but also statistically defensible site localizations.
6. Key Takeaways for Robust DIA PTM Proteomics
High-quality PTM proteomics relies on aligning experimental design with computational rigor. At the search-parameter level, variable modifications should be selected to match the biology and enrichment strategy, while keeping the search space controlled to protect sensitivity and statistical power. At the interpretation level, PTM site localization must be treated as a first-class result: probability-based localization scoring provides an objective way to separate confidently localized sites from ambiguous assignments. Finally, multi-level FDR control—especially site-level FDR—acts as the guardrail that keeps large PTM datasets scientifically trustworthy.
When these pillars are implemented together, DIA becomes a powerful platform for reproducible PTM discovery and quantification, supporting applications ranging from signaling-pathway mapping to cohort-scale studies that seek PTM biomarkers or treatment response markers. The practical outcome should be a PTM site table that is not only large, but also auditable: each site can be traced back to peptide evidence, fragment-ion support, localization probability, and clearly stated statistical thresholds.
References
1. Olsen, J. V., & Mann, M. (2013). Status of large-scale analysis of post-translational modifications by mass spectrometry. Molecular & cellular proteomics : MCP, 12(12), 3444–3452. https://doi.org/10.1074/mcp.O113.034181
2. Yang, Y., & Qiao, L. (2023). Data-independent acquisition proteomics methods for analyzing post-translational modifications. Proteomics, 23(7-8), e2200046. https://doi.org/10.1002/pmic.202200046
3. Bekker-Jensen, D. B., Bernhardt, O. M., Hogrebe, A., Martinez-Val, A., Verbeke, L., Gandhi, T., Kelstrup, C. D., Reiter, L., & Olsen, J. V. (2020). Rapid and site-specific deep phosphoproteome profiling by data-independent acquisition without the need for spectral libraries. Nature communications, 11(1), 787. https://doi.org/10.1038/s41467-020-14609-1
4. Tian, X., Permentier, H. P., & Bischoff, R. (2023). Chemical isotope labeling for quantitative proteomics. Mass spectrometry reviews, 42(2), 546–576. https://doi.org/10.1002/mas.21709
Next-Generation Omics Solutions:
Proteomics & Metabolomics
Ready to get started? Submit your inquiry or contact us at support-global@metwarebio.com.


