Secretomics: Decoding the Secreted Proteome for Extracellular Communication
Every cell in the human body speaks a language—not with words, but with proteins. These extracellular proteins constitute the secretome, a complex repertoire of signaling molecules that orchestrate development, maintain homeostasis, and drive disease progression. Unlike the static genome, the cell secretome is dynamic, reflecting real-time cellular states and environmental responses. For biomedical researchers and drug discovery scientists, decoding this extracellular language offers unprecedented opportunities: biomarkers discovery in easily accessible biofluids, identification of therapeutic targets within the tumor microenvironment secretome, and mechanistic insights into cell-cell communication. Recent advances in mass spectrometry-based proteomics technologies have transformed our ability to profile these secreted proteins with depth and accuracy. This comprehensive guide explores how modern secretome analysis is unlocking new dimensions of human biology and clinical translation.
1. Secretomics and the Secreted Proteome: Definitions and Scope
The secretome encompasses the complete set of proteins actively released by cells, tissues, or organisms into the extracellular space (Knecht et al., 2023). This definition includes not only classically secreted soluble mediators but also proteins associated with extracellular vesicles (EVs) and components integrated into or interacting with the extracellular matrix. Secretomics refers to the systematic characterization of this secreted proteome using high-resolution proteomics and complementary protein-based technologies.
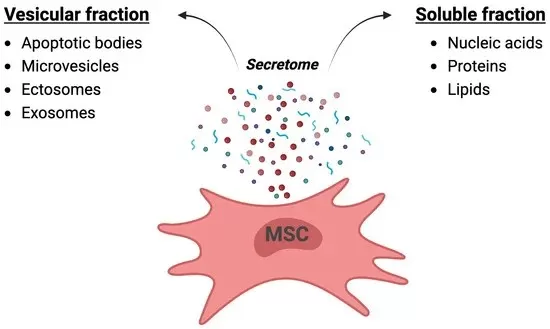
Illustration of the secretome released by mesenchymal stem cells.
Image reproduced from Kråkenes et al., 2024, International journal of molecular sciences, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
Unlike conventional proteomics, which surveys the entire intracellular protein complement, secretome analysis specifically interrogates exported proteins that mediate intercellular communication. Within this broader framework, exosome proteomics and extracellular vesicle proteomics represent focused subfields dedicated to vesicle-associated cargo, which constitutes only one fraction of the total secreted proteome. In practical terms, secretome studies are typically performed on conditioned media proteomics samples derived from cell culture supernatants and/or on purified EV populations, depending on the biological question and required resolution.
Extracellular proteins are central regulators of tissue-level physiology and pathology. Cytokines orchestrate immune responses, growth factors govern tissue development and repair, and proteases and matrix components dynamically remodel the extracellular microenvironment. In disease settings, particularly cancer, the tumor microenvironment secretome actively drives progression through immunosuppression, angiogenesis, stromal activation, and extracellular matrix remodeling (Caxali et al., 2025). This functional diversity confers substantial translational value. Because secreted proteins enter circulation and other biofluids, they are inherently accessible for minimally invasive biomarkers discovery. Moreover, their extracellular localization makes them highly amenable to therapeutic targeting, as exemplified by successful biologics against VEGF and TNF‑α. For these reasons, secretomics has become an increasingly powerful strategy to bridge mechanistic cell models with clinically actionable, patient‑relevant readouts.
2. Secretion Pathways That Shape the Cell Secretome
The routes by which proteins reach the extracellular space are remarkably diverse. Understanding these pathways is fundamental to interpreting secretome analysis data and distinguishing genuine secretion from artifacts.
2.1 Classical Secretion: Signal Peptide and ER–Golgi Export
The canonical route for protein secretion involves N-terminal signal peptides that direct nascent polypeptides to the endoplasmic reticulum, followed by Golgi-mediated trafficking and vesicular release. This pathway handles most classical signaling molecules—cytokines, hormones, growth factors, and secreted enzymes. Approximately 9% of human genes encode such classically secreted products, yet the functions of most remain unexplored (Wat et al., 2024). In conditioned media proteomics, proteins following this pathway typically exhibit characteristic signal peptide sequences and undergo glycosylation, features that inform bioinformatic prediction and experimental validation.
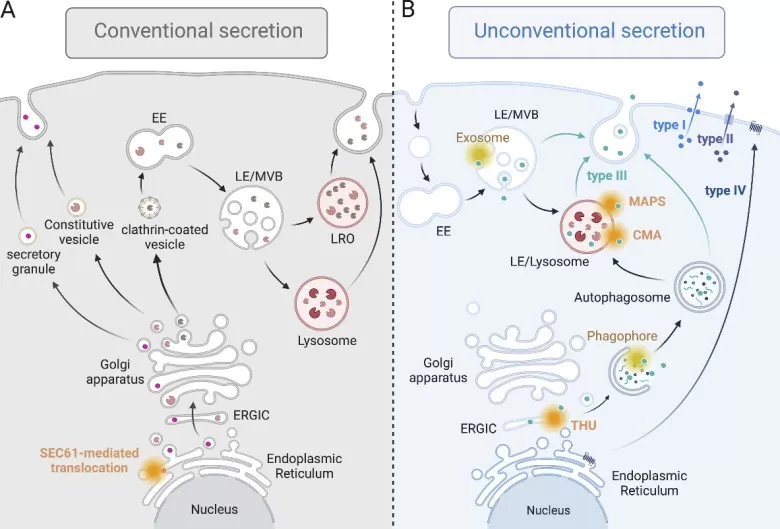
Conventional and unconventional protein secretion.
Image reproduced from Néel et al., 2024, The Journal of cell biology, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
2.2 Unconventional Secretion: Leaderless Secretory Proteins
A substantial fraction of the secreted proteome lacks identifiable signal peptides yet reproducibly appears extracellularly. These leaderless proteins employ alternative export mechanisms including direct plasma membrane translocation, lysosomal secretion, and ABC transporter-mediated release. Well-characterized examples include fibroblast growth factor 2 (FGF2), interleukin-1β, and macrophage migration inhibitory factor (MIF) (Elghetany et al., 2024). Recognition of non-classical secretion is critical for secretome analysis, as conventional signal peptide prediction algorithms will underestimate true secretory output.
2.3 Extracellular Vesicles - Mediated Secretion
Extracellular vesicle proteomics has revealed that a significant portion of the cell secretome is vesicle-encapsulated rather than freely soluble. Exosomes (30–150 nm), microvesicles (100–1000 nm), and apoptotic bodies each carry distinct protein cargoes protected from the extracellular environment. This packaging enables coordinated delivery of multiple signaling molecules, protects labile factors from degradation, and facilitates long-distance communication. For secretome analysis, distinguishing between soluble and vesicular fractions—through ultracentrifugation, size exclusion chromatography, or immunocapture—provides mechanistic insights into intercellular communication.
_1772767265_WNo_793d466.webp)
Biogenesis and components of extracellular vesicles (Evs)
Image reproduced from Singh et al., 2025, Proteomes, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
2.4 The Secretome Microenvironment: The Extracellular Matrix (ECM)
Secreted proteins do not simply diffuse away—many become immobilized within the extracellular matrix, creating a bioactive niche that regulates cell behavior. Collagens, laminins, fibronectin, and proteoglycans are major extracellular proteins that provide structural support while sequestering growth factors and presenting them to cells. The ECM thus functions as a dynamic reservoir, continuously remodeled by secreted proteases. This interplay between soluble mediators and insoluble matrix components is particularly relevant in tumor microenvironment secretome studies, where matrix remodeling drives invasion and metastasis (Caxali et al., 2025).
3. Experimental Design for Secretome Analysis
Robust secretome analysis begins with meticulous experimental design. The choice of sample type, collection conditions, and quality controls fundamentally determines data quality and biological interpretability.
3.1 Sample Types and Application Scenarios
Cell secretome studies typically employ conditioned medium from cultured cells, collected after defined incubation periods. Primary cells, cell lines, organoids, and tissue explants each present unique considerations. For biomarkers discovery, biofluids including plasma, serum, urine, cerebrospinal fluid, and malignant effusions provide clinically accessible windows into disease-related secretion. Critical considerations include:
- Serum-free conditioned medium: Essential to avoid interference from abundant serum proteins like albumin and immunoglobulins, which suppress detection of low-abundance secreted factors (Baby Mattamana et al., 2025)
- Collection timing: Must balance accumulation of secreted proteins against nutrient depletion and metabolic stress
- Protease inhibitors: Immediate addition prevents degradation of labile mediators
3.2 Quality Control (QC) and Contamination Assessment
Rigorous QC distinguishes genuine secretion from artifactual release. Cell viability assessment (trypan blue exclusion, LDH release) confirms that detected proteins reflect active secretion rather than passive leakage from damaged cells. Contamination control includes parallel "blank" medium controls incubated without cells to identify proteins originating from serum supplements or culture vessels. Batch controls ensure reproducibility across experimental replicates. In SILAC secretome experiments, metabolic labeling provides an additional internal control for newly synthesized versus medium-derived proteins. Methodologically, even protein quantification can mislead: Otsuka et al. describe how common assays can overestimate protein concentration in concentrated media and propose a normalization approach based on concentration rate to improve reproducibility (Otsuka et al., 2025).
3.3 Extracellular Vesicles Isolation Methods
When exosome proteomics or extracellular vesicle proteomics is the focus, isolation method choice profoundly impacts results—influencing not only which vesicles are recovered but also the purity, yield, and biological interpretability of downstream data. Different techniques exploit distinct physical or biochemical properties of vesicles, and each carries inherent trade-offs between recovery efficiency and contamination with non-vesicular components. The presence of abundant soluble extracellular proteins, lipoprotein particles, and protein aggregates can confound secretome analysis if not adequately separated from genuine vesicular cargo. Understanding these methodological nuances is essential for designing experiments that accurately reflect the biological reality of EV-mediated communication.
|
Method |
Principle |
Yield |
Purity |
Application |
|
Differential centrifugation |
Sequential spins (300×g → 2,000×g → 10,000×g → 100,000×g) |
High |
Moderate |
Preparative isolation |
|
Density gradient ultracentrifugation |
Cushion/layered gradient (iodixanol/sucrose) |
Moderate |
High |
Removal of non-vesicular contaminants |
|
Size exclusion chromatography (SEC) |
Column-based size fractionation |
Moderate |
High |
Gentle isolation preserving vesicle integrity |
|
Immunocapture |
Antibody against surface markers (CD9, CD63, CD81) |
Low |
Very high |
Subpopulation-specific analysis |
The choice among these methods should be guided by the specific research question: discovery-phase extracellular vesicle proteomics may benefit from density gradient purification to minimize false positives from co-precipitated contaminants, while functional assays requiring intact, bioactive vesicles often favor SEC's gentler separation. For biomarkers discovery in clinical samples, combining complementary methods—such as SEC followed by immunocapture—can achieve both purity and biological specificity, though at the cost of reduced throughput and yield.
4. Core Technologies for Secretome Profiling
Modern secretome analysis integrates sophisticated sample processing with advanced mass spectrometry to achieve depth, reproducibility, and biological insight.
4.1 Secretome Enrichment and Background Reduction
The fundamental challenge in conditioned media proteomics is the vast dynamic range—low-abundance signaling molecules coexist with abundant structural proteins and, in serum-containing systems, overwhelming albumin and immunoglobulin backgrounds. Optimized workflows address this through (Baby Mattamana et al., 2025):
i. Ultrafiltration (50 kDa MWCO) to concentrate large volumes while fractionating by molecular weight
ii. Depletion columns removing high-abundance serum proteins (albumin, IgG)
iii. Acetone/TCA precipitation maximizing recovery of both high- and low-molecular-weight proteins
iv. Processing filtrate and concentrate separately for albumin-rich media, or pooling for serum-free conditioned medium to capture the full secreted proteome
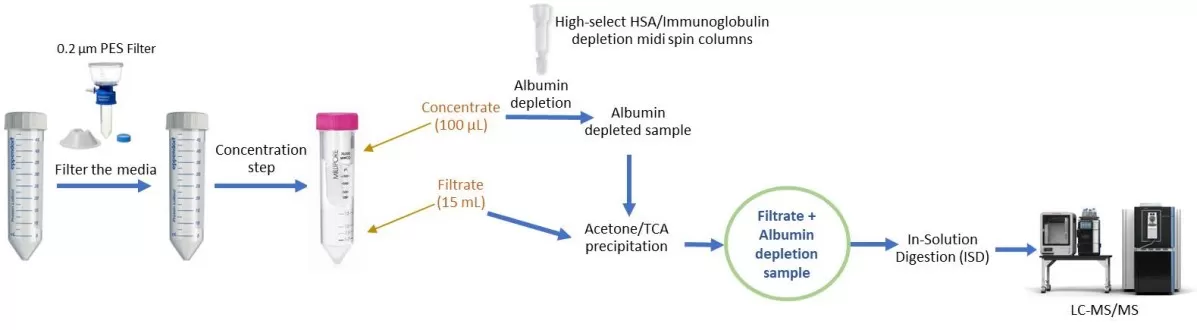
Workflow for optimized secretome preparation for LC–MS/MS analysis
Image reproduced from Baby Mattamana et al., 2025, Bio-protocol.
4.2 LC–MS/MS Strategies: DDA vs. DIA Proteomics
The choice of data acquisition mode shapes the depth and reproducibility of secretome analysis:
- Data-Dependent Acquisition (DDA) selects the most abundant precursor ions for fragmentation, offering deep coverage but stochastic sampling between runs—well-suited for discovery-phase biomarkers discovery
- Data-Independent Acquisition (DIA) fragments all precursor ions within defined mass windows, providing comprehensive, reproducible quantification across large cohorts. DIA proteomics excels for tumor microenvironment secretome studies requiring consistent measurement across many samples, though it demands robust spectral libraries for optimal data extraction
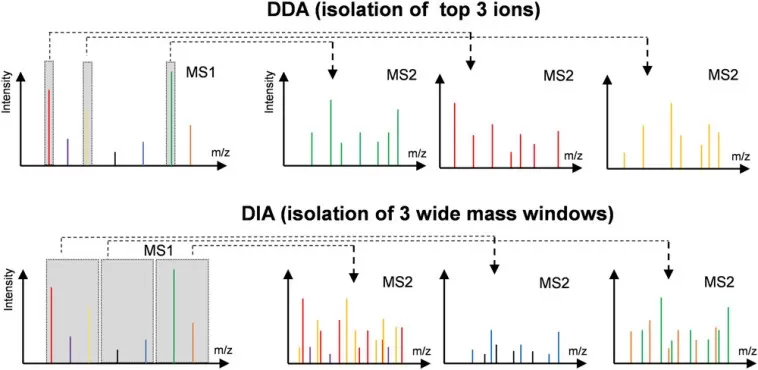
Difference in MS1 isolation windows for the DDA and DIA modes.
Image reproduced from Tian, X., Permentier, H. P., Bischoff, R, 2023, Mass spectrometry reviews, licensed under the Creative Commons Attribution License (CC BY 4.0).
4.3 Quantification Approaches: Label-Free, TMT, and SILAC Proteomics
Quantitative secretome analysis employs multiple strategies matched to experimental goals:
- Label-free quantification: Accessible and versatile, comparing peptide signals across runs; suitable for most discovery applications
- TMT proteomics (Tandem Mass Tags): Multiplexing up to 18 samples enables direct comparison with reduced run-to-run variation, ideal for time courses or perturbation studies
- SILAC secretome (Stable Isotope Labeling by Amino Acids in Cell Culture): Metabolic labeling distinguishes newly synthesized (heavy) from medium-derived (light) proteins—particularly valuable for contamination control and studying secretion dynamics
5. Interpreting Secretome Data: From Protein Identification to Biomarker Discovery
Raw protein identifications require sophisticated bioinformatic processing to yield biologically meaningful insights about the cell secretome.
5.1 Secretory Bioinformatics Annotation Framework
The first analytical step confirms and characterizes the secretory nature of identified proteins. Computational tools predict signal peptides (SignalP, Phobius) and non-classical secretion (SecretomeP). Subcellular localization annotation (UniProt, GO cellular component) distinguishes genuinely secreted proteins from contaminants released during sample processing. The extracellular proteins annotation includes both soluble mediators and membrane proteins that may appear in conditioned media proteomics through shedding or vesicle association.
5.2 Contamination/Lysis Assessment
A critical quality control step evaluates potential artifacts. Elevated proportions of intracellular markers—cytoplasmic (GAPDH, actin), nuclear (histones), or mitochondrial proteins—indicate cell lysis during collection. Contamination control metrics include comparing detected proteins against established "contaminant repositories" and assessing batch effects through principal component analysis. In SILAC secretome experiments, the ratio of heavy to light peptides provides direct evidence of active synthesis during the labeling period.
5.3 Functional and Mechanistic Analysis
With a curated secreted proteome list, functional interpretation proceeds through:
- Pathway enrichment (KEGG, Reactome, GO biological process) identifying overrepresented biological themes
- Protein-protein interaction networks revealing functional modules and hub proteins
- Upstream regulator analysis predicting transcriptional or signaling programs driving observed secretion
- Cell-cell communication inference integrating ligand-receptor pairs with recipient cell expression data
5.4 From Discovery to Validation
Candidate biomarkers discovery requires orthogonal validation. Targeted mass spectrometry approaches (PRM/MRM) provide quantitative, multiplexed measurement of selected proteins across many samples. Immunoassays (ELISA, Luminex) offer higher throughput for validated candidates. Prioritization considers biological plausibility, statistical significance, fold-change magnitude, and technical reproducibility.
6. Applications of Secretomics in Biomedicine and Drug Development
The power of modern secretome analysis is best illustrated through concrete applications spanning basic biology to clinical translation.
6.1 Tumor and Tumor Microenvironment (TME)
The tumor microenvironment secretome orchestrates cancer progression through multiple mechanisms. A recent study integrating secretome analysis and interactome mapping in colorectal cancer identified region-specific biomarkers: FGFR4, FLT1, and WNT5A emerged as key players modulating fibroblast recruitment and cell division, with expression patterns varying by tumor location. This work further predicted dovitinib and nintedanib as potential therapeutic agents targeting multiple identified markers (Caxali et al., 2025). Such approaches demonstrate how secretome analysis can reveal both diagnostic biomarkers and actionable therapeutic targets.
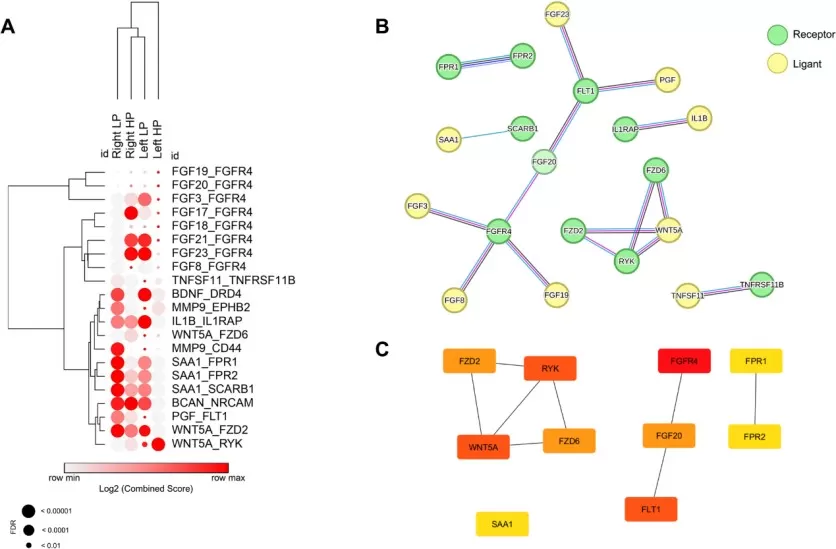
A Interactome pairs that were altered in specific components across various regions of the large intestine. B Protein-protein interaction network of the altered genes. C Top 10 genes with the highest number of interactions in the protein-protein network
Image reproduced from Caxali et al., 2025, Cell Communication and Signaling, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
6.2 Inflammation/Infection and Immunity
The immune cell secretome comprises complex cytokine networks that coordinate host defense but can drive pathology when dysregulated. In malignant pleural effusions across multiple epithelial cancers, multiplex analysis revealed eleven consistently elevated analytes including CXCL10/IP10 (672.3 pM), CCL2/MCP1 (562.9 pM), sIL-6Rα (403.1 pM), and IL-6 (137.6 pM) (Donnenberg et al., 2024). These mediators promote chemotaxis, epithelial-mesenchymal transition, and immunosuppression, with the IL-6/sIL-6Rα axis emerging as a central pathogenic driver and therapeutic target.
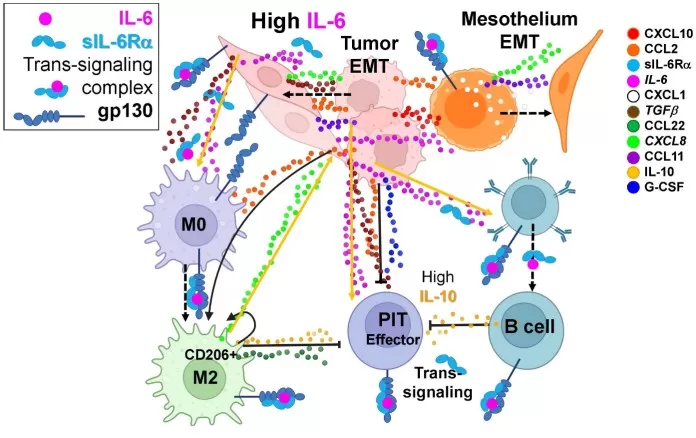
The IL-6 axis interacts with multiple cytokines to promote tumor and mesothelial EMT and suppress anti-tumor immunity.
Image reproduced from Donnenberg et al., 2024, Frontiers in Immunology, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
6.3 Stem Cells and Regenerative Medicine
Mesenchymal stem cell (MSC) therapies increasingly focus on paracrine mechanisms rather than engraftment. The MSC cell secretome contains complex mixtures of growth factors, immunomodulatory cytokines, and extracellular vesicles that mediate tissue repair. In a recent study, human umbilical cord MSC secretome analysis combined with nanoemulsion propolis reduced osteoclastogenesis in LPS-induced osteolysis with hyperglycemia. Treatment inhibited TRAP and cathepsin K levels while enhancing OPG expression, demonstrating therapeutic potential for inflammatory bone loss (Nugraha et al., 2025).
6.4 Drug Mechanisms and Biomarker Development
Secretome analysis is increasingly integrated into drug development pipelines. Pharmacoproteomic profiling of 46 drugs in hepatocyte models revealed distinct clusters of non-canonical secretion corresponding to specific cellular insults—lysosomal alterations from cationic-amphiphilic drugs, mitochondrial damage linked to Rab7-dependent protein secretion, and ER stress reducing basal secretion (Knecht et al., 2024). Such approaches identify secreted markers for aberrant drug action, supporting mechanism-of-action studies, pharmacodynamic biomarker development, and safety assessment.
7. Future Directions of Secretomics and Extracellular Proteomics
The secretome has emerged as a fundamental executor of cellular function—the dynamic protein repertoire through which cells orchestrate tissue homeostasis, coordinate immune responses, and drive disease pathogenesis. Advances in conditioned media proteomics, DIA proteomics, and extracellular vesicle proteomics have transformed our capacity to decode this extracellular language, enabling unprecedented depth in biomarkers discovery and mechanistic insight into intercellular communication. These technological developments, coupled with rigorous experimental design addressing contamination control, have established secretome analysis as an indispensable pillar of modern biomedical research.
Looking forward, the field is poised for transformative evolution. Spatial biology approaches—combining laser capture microdissection with ultrasensitive proteomics—will soon resolve tumor microenvironment secretome heterogeneity at single-cell resolution. Integration of multi-omics data, particularly linking secretome analysis with transcriptomics and metabolomics, promises systems-level understanding of secretory networks in health and disease. As computational tools mature to predict secretion signals and model cell–cell communication, the secreted proteome will increasingly inform clinical decision-making—from liquid biopsy-based diagnostics to rationally designed therapeutics targeting extracellular mediators. The coming decade will witness the cell secretome transition from discovery science to routine clinical application, fundamentally reshaping precision medicine.
Reference
1. Kråkenes, T., Sandvik, C. E., Ytterdal, M., Gavasso, S., Evjenth, E. C., Bø, L., & Kvistad, C. E. (2024). The Therapeutic Potential of Exosomes from Mesenchymal Stem Cells in Multiple Sclerosis. International journal of molecular sciences, 25(19), 10292. https://doi.org/10.3390/ijms251910292
2. Knecht, S., Eberl, H. C., Kreisz, N., Ugwu, U. J., Starikova, T., Kuster, B., & Wilhelm, S. (2023). An Introduction to Analytical Challenges, Approaches, and Applications in Mass Spectrometry-Based Secretomics. Molecular & cellular proteomics : MCP, 22(9), 100636. https://doi.org/10.1016/j.mcpro.2023.100636
3. Caxali, G. H., Aal, M. C. E., Osvaldo, C. W. G., Oliveira, J. S., Bidinotto, L. T., Carvalho, R. F., & Delella, F. K. (2025). Integrating the secretome and interactome to identify novel biomarkers and therapeutic targets in colorectal cancer. Cell communication and signaling : CCS, 23(1), 428. https://doi.org/10.1186/s12964-025-02424-4
4. Wat, L. W., & Svensson, K. J. (2024). Novel secreted regulators of glucose and lipid metabolism in the development of metabolic diseases. Diabetologia, 67(12), 2626–2636. https://doi.org/10.1007/s00125-024-06253-x
5. Elghetany, M. T., Pan, J. L., Sekar, K., Major, A., Mf Su, J., Adesina, A., Hui, K. M., Li, X. N., & Teo, W. Y. (2024). Re-programming by a six-factor-secretome in the patient tumor ecosystem during nutrient stress and drug response. iScience, 27(10), 110932. https://doi.org/10.1016/j.isci.2024.110932
6. Néel, E., Chiritoiu-Butnaru, M., Fargues, W., Denus, M., Colladant, M., Filaquier, A., Stewart, S. E., Lehmann, S., Zurzolo, C., Rubinsztein, D. C., Marin, P., Parmentier, M. L., & Villeneuve, J. (2024). The endolysosomal system in conventional and unconventional protein secretion. The Journal of cell biology, 223(9), e202404152. https://doi.org/10.1083/jcb.202404152
7. Singh, M., Tiwari, P. K., Kashyap, V., & Kumar, S. (2025). Proteomics of Extracellular Vesicles: Recent Updates, Challenges and Limitations. Proteomes, 13(1), 12. https://doi.org/10.3390/proteomes13010012
Baby Mattamana, B., Gajjela, R., K C, J., Parish, R. A., & Faull, P. A. (2025). Optimized Secretome Sample Preparation From High Volume Cell Culture Media for LC-MS/MS Proteomic Analysis. Bio-protocol, 15(24), e5542. https://doi.org/10.21769/BioProtoc.5542
8. Otsuka, T., Hatano, A., Matsumoto, M., & Matsui, H. (2025). A robust protocol for proteomic profiling of secreted proteins in conditioned culture medium. Biology methods & protocols, 10(1), bpaf068. https://doi.org/10.1093/biomethods/bpaf068
9. Tian, X., Permentier, H. P., & Bischoff, R. (2023). Chemical isotope labeling for quantitative proteomics. Mass spectrometry reviews, 42(2), 546–576. https://doi.org/10.1002/mas.21709
10. Donnenberg, V. S., Luketich, J. D., Popov, B., Bartlett, D. L., & Donnenberg, A. D. (2024). A common secretomic signature across epithelial cancers metastatic to the pleura supports IL-6 axis therapeutic targeting. Frontiers in immunology, 15, 1404373. https://doi.org/10.3389/fimmu.2024.1404373
11. Nugraha, A. P., Yang, K., Pramusita, A., Putranti, N. A. R., Luthfi, M., Ramadhani, N. F., Sitalaksmi, R. M., Situmorang, P. C., Artasasta, M. A., Alhasyimi, A. A., Riawan, W., Nugraha, A. P., Shariff, K. A., Tanimoto, K., & Hong, G. (2025). Human umbilical cord mesenchymal stem cells secretome and nanoemulsion propolis combination ameliorate osteoclastogenesis in lipopolysaccharide-induced osteolysis in hyperglycemia rats. Journal of dental sciences, 20(4), 2181–2188. https://doi.org/10.1016/j.jds.2025.07.003
12. Knecht S, Kalxdorf M, Korbeń J, Mathieson T, Sevin DC, Kuster B, Kasprowicz R, Sakatis MZ, Eberl HC, Bantscheff M. (2024) Pharmacoproteomic profiling identifies secreted markers for aberrant drug action.bioRxiv. https://doi.org/10.1101/2024.10.16.618637
Next-Generation Omics Solutions:
Proteomics & Metabolomics
Ready to get started? Submit your inquiry or contact us at support-global@metwarebio.com.


