Signal Peptides and Protein Secretion: Mechanisms, Prediction, and Engineering
Behind every successful therapeutic protein, vaccine antigen, or secreted biomarker lies a critical but often overlooked component: the signal peptide. These short N-terminal sequences function as molecular "ZIP codes," determining whether a protein will be efficiently secreted or retained intracellularly. With approximately one-third of all cellular proteins entering the secretory pathway, understanding signal peptide biology has never been more important for biotechnological innovation. This comprehensive guide explores the fundamental mechanisms of signal peptide-mediated secretion, introduces state-of-the-art prediction tools, and provides practical strategies for designing and optimizing signal peptides in recombinant protein production.
1. What Are Signal Peptides? The Gateway to Protein Secretion
Signal peptides were historically framed as disposable targeting tags, but modern structural biology, proteomics, and immunology show they are better viewed as decision elements in protein biogenesis. Signal peptides represent one of nature's most elegant solutions to the problem of protein sorting. These short amino acid sequences serve as essential targeting signals that direct nascent proteins to the endoplasmic reticulum (ER) for secretion or membrane integration (Zhang et al., 2025).
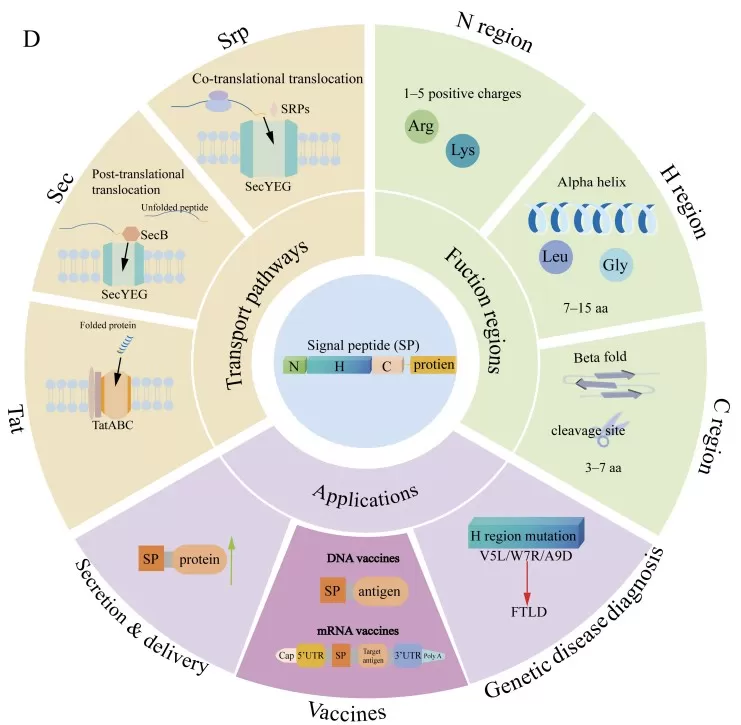
The protein translocation pathways mediated by SPs and their related application.
Image reproduced from Zhang et al., 2025, Biomolecules, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
1.1 Signal Peptide Definition and the Signal Hypothesis
A signal peptide is typically a short (often ~16–30 aa) N-terminal sequence that targets a nascent polypeptide to the secretory pathway, most commonly the ER in eukaryotes, where translocation and processing begin. The concept of signal peptides emerged from the pioneering work of Günter Blobel and colleagues in the 1970s. The seminal 1972 study by Milstein et al. observed that secreted immunoglobulin light chains lacked N-terminal sequences found in their precursors, suggesting these sequences were cleaved during transport (Zhang et al., 2025). This observation led to the "Signal Hypothesis," which proposed that proteins contain intrinsic signals for their subcellular localization—a discovery that earned Blobel the Nobel Prize in 1999. Today, we understand signal peptides as universal features across all life forms, from bacteria to complex eukaryotes.
1.2 Why Signal Peptides Matter for the Secretory Pathway
Functionally, signal peptides initiate SRP-dependent ER targeting, then enable entry into the ER environment where folding, glycosylation, disulfide formation, and quality control determine whether a protein is productively secreted or retained/degraded. This is why signal peptides are not only a cell biology concept but also a practical “front gate” for secretion efficiency and manufacturability. In omics, signal peptides sit at the intersection of genome-based secretion prediction and proteomics validation: prediction proposes candidates, while secretome/EV proteomics and N-terminomics can confirm secretion evidence and cleavage behavior.
1.3 Signal Peptide vs Signal Anchor vs Transit Peptides
In the field of protein targeting, several terms—signal peptide, signal anchor, transit peptide, and nuclear localization signal—are frequently encountered and, more often than not, used interchangeably or confused. While all function as "address labels" directing proteins to specific cellular locations, they operate through fundamentally different mechanisms. A signal peptide is a short, cleavable N-terminal sequence that directs proteins into the endoplasmic reticulum for secretion; a signal anchor, by contrast, is an uncleaved transmembrane domain that embeds the protein in the lipid bilayer. Transit peptides guide proteins into mitochondria or chloroplasts and are cleaved upon arrival, whereas nuclear localization signals are internal, non-cleaved sequences that direct proteins into the nucleus through nuclear pores.
|
Feature |
Signal Peptide (Secretory) |
Signal Anchor |
Transit Peptide |
Nuclear Localization Signal |
|
Location |
N-terminal |
N-terminal or internal |
N-terminal |
Internal (anywhere) |
|
Cleavage |
Cleaved by signal peptidase |
Not cleaved |
Cleaved in organelle |
Not cleaved |
|
Target |
ER → Secretion |
ER membrane |
Mitochondria/Chloroplasts |
Nucleus |
|
Final Location |
Secreted or lumenal |
Transmembrane |
Organelle matrix |
Nucleoplasm |
|
Hydrophobicity |
High (H-region) |
Very high (transmembrane) |
Moderate, amphiphilic |
Low, basic residues |
2. Signal Peptide Structure: Architecture and Functional Diversity
The functionality of signal peptides emerges from their conserved three-domain architecture. Each region contributes distinct properties that collectively ensure efficient targeting and processing (Zhang et al., 2025).
2.1 The Tripartite Structure: N-region, H-region, and C-region
The canonical signal peptide comprises three structurally and functionally distinct regions. The N-region contains positively charged amino acids (lysine, arginine) that interact with negatively charged phospholipid head groups and the signal recognition particle (SRP). The net charge of this region influences targeting pathway selection and translocation efficiency (Kaushik et al., 2022). The H-region forms the hydrophobic core, typically 8-20 residues with high leucine, valine, and alanine content. This region inserts into the Sec61 translocon as a helical hairpin and serves as the primary recognition element for SRP (Lang et al., 2022). Finally, the C-region contains the cleavage site recognized by signal peptidase, characterized by small, neutral amino acids at positions -3 and -1 relative to the cleavage site (the "-3,-1 rule") (Zhang et al., 2025).
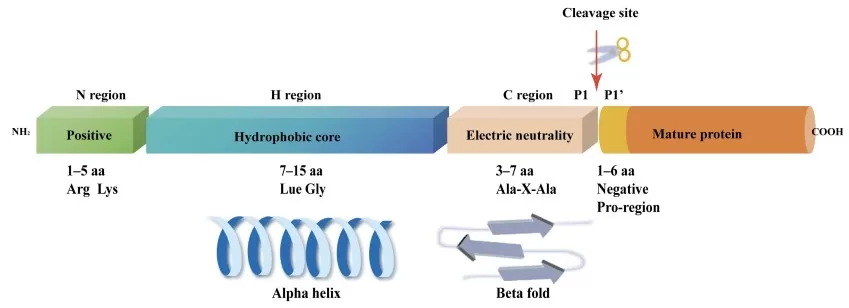
A schematic diagram of the typical structure of the SP.
Image reproduced from Zhang et al., 2025, Biomolecules, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
2.2 Sequence Conservation vs. Diversity Across Species
Despite functional conservation, signal peptides exhibit remarkable sequence diversity across species and protein families. Bacterial signal peptides tend to be shorter with less hydrophobic H-regions compared to their eukaryotic counterparts, reflecting differences in targeting machinery (Kaushik et al., 2022). Within eukaryotes, signal peptides show species-specific preferences in amino acid composition that correlate with optimal expression in different host systems—a critical consideration for heterologous protein production.
2.3 Sequence Variants that Shift Targeting, Cleavage, and Fate
Recent research has revealed that subtle variations in signal peptide sequence can profoundly impact protein fate. H-region hydrophobicity determines whether a protein follows co-translational (SRP-dependent) or post-translational targeting pathways. N-region charge density influences translocation efficiency, while C-region context—including propeptide sequences—affects cleavage accuracy. Importantly, mutations in the hydrophobic core represent the most common pathogenic variants in human signal peptides, often triggering the RAPP (Regulation of Aberrant Protein Production) quality control pathway and targeted mRNA degradation (Gutierrez Guarnizo et al., 2023). Approximately 25% of pathogenic mutations alter either the N-region or cleavage site, resulting in translocation deficiencies or processing failures (Gutierrez Guarnizo et al., 2023).
3: How Signal Peptides Work: ER Targeting and the Secretory Pathway
The journey of a secretory protein begins with co-translational targeting to the ER and culminates in vesicular transport to the extracellular space. Signal peptides orchestrate this journey through a series of precisely coordinated interactions. However, not all signal peptides follow the same molecular route. Based on the translocation machinery they engage, signal peptides can be broadly classified into several types—a framework now embedded in modern prediction tools like SignalP 6.0. The most common is the Sec/SPI signal peptide, which directs proteins to the general Sec translocation system for transport in an unfolded state. A variation, Sec/SPII, targets proteins for lipidation and membrane anchoring. In contrast, the Tat/SPI signal peptide guides folded proteins through the twin-arginine translocation (Tat) pathway, a mechanism prevalent in bacteria and plant chloroplasts. This chapter focuses primarily on the well-characterized Sec/SPI pathway, which underpins the majority of eukaryotic protein secretion, while noting key distinctions where relevant.
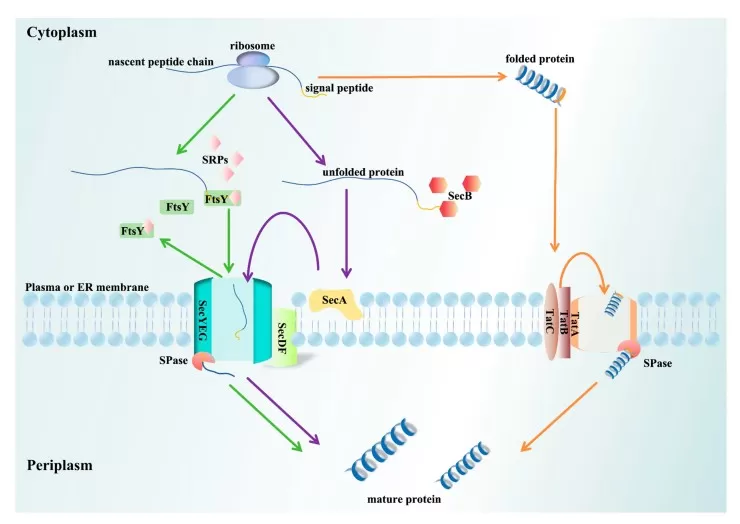
Schematic diagrams of different transport pathways.
Image reproduced from Zhang et al., 2025, Biomolecules, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
3.1 SRP-Dependent Targeting and Ribosome Docking
As the signal peptide emerges from the ribosome, its hydrophobic H-region is recognized by the signal recognition particle (SRP). This binding induces translational arrest, preventing premature folding and maintaining translocation competence. The SRP-ribosome-nascent chain complex then docks with the SRP receptor on the ER membrane, positioning the ribosome for direct transfer to the Sec61 translocon. The hydrophobicity threshold for SRP recognition determines pathway choice—proteins with moderately hydrophobic signals may bypass SRP entirely for post-translational targeting.
3.2 ER Translocation via Sec61 and Signal Peptide Insertion
Upon docking, the signal peptide inserts into the Sec61 translocon in a loop conformation, with the N-region facing the cytoplasm and the C-region entering the ER lumen. The Sec61 channel opens laterally to accommodate the signal peptide while maintaining the permeability barrier. Recent structural studies have illuminated how signal peptides actively gate the Sec61 channel, with specific features determining translocation efficiency and substrate specificity. In human cells, auxiliary components including Sec62/Sec63, TRAM1, and TRAP support translocation for challenging substrates.
3.3 Signal Peptidase Cleavage and Cleavage-Site Determinants
Signal peptidase (SPase) recognizes the C-region cleavage site, cleaving between positions -1 and +1 to release the mature protein into the ER lumen. The enzyme exhibits strong preference for small, neutral residues at the -3 and -1 positions, typically alanine, glycine, or serine. Cleavage failure results in retained signal peptides, leading to mislocalization, ER retention, and ultimately ER stress or proteasomal degradation. Non-specific cleavage events have also been documented, with SPase occasionally processing at alternative sites.
3.4 ER Folding, Quality Control, and Secretion Output
The signal peptide's role extends beyond targeting—it sets the stage for subsequent events in the secretory pathway. Upon entry into the ER, proteins encounter chaperones (BiP, calnexin/calreticulin) that facilitate folding, while glycosylation enzymes modify asparagine residues. Properly folded proteins proceed through the Golgi apparatus to secretion; misfolded proteins are retrotranslocated and degraded by ERAD (ER-associated degradation). The final secretome composition reflects integrated decisions at each step—targeting efficiency, folding success, and quality control stringency.
4. Signal Peptide Prediction: Bioinformatics Approaches and Best Practices
Computational prediction of signal peptides has become essential for genome annotation, target selection, and protein engineering. However, interpreting prediction results requires understanding tool capabilities and limitations.
4.1 Common Signal Peptide Prediction Tools
Modern prediction tools leverage machine learning to achieve impressive accuracy. SignalP 6.0 utilizes deep neural networks to not only predict signal peptides but also classify them into Sec/SPI (standard secretory), Sec/SPII (lipoprotein), and Tat/SPI (twin-arginine translocation) types (Zhang et al., 2025). Phobius combines signal peptide and transmembrane domain prediction using hidden Markov models, reducing false positives from transmembrane helices. TargetP 2.0 extends prediction to multiple subcellular localizations, including mitochondria and chloroplasts. Tool outputs typically include probability scores, cleavage site positions, and classification information.
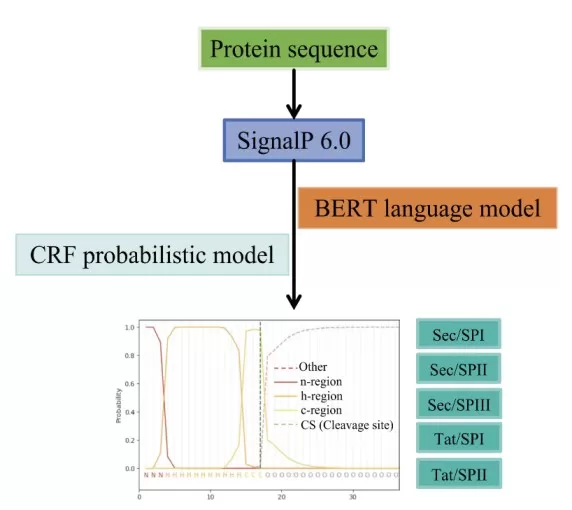
The workflow of SP prediction using SignalP 6.0.
Image reproduced from Zhang et al., 2025, Biomolecules, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
4.2 Confidence Interpretation and False-Positive Control
High prediction scores do not guarantee true secretion. Transmembrane helices at the N-terminus (signal anchors) frequently score as false-positive signal peptides due to structural similarities. Low-complexity regions and extreme amino acid compositions can also trigger false predictions. Best practice employs cross-validation with multiple tools and integration of complementary evidence: SignalP for classification, Phobius for transmembrane discrimination, and TargetP for organelle targeting assessment. SignalP 6.0's ability to distinguish signal peptide types provides additional confidence in prediction specificity.
4.3 From Prediction to Evidence: Secretome Proteomics and N-Terminomics
In silico predictions require experimental validation. Secretome analysis of culture supernatants by mass spectrometry provides direct evidence of protein secretion, though careful controls for cell lysis are essential. N-terminomics approaches (e.g., COFRADIC, TAILS) can experimentally identify cleavage sites, confirming signal peptidase processing. Subcellular fractionation followed by proteomic analysis localizes proteins to specific compartments. The Paenibacillus secretome study exemplifies this integrated approach, combining SignalP 4.1 predictions with nanoLC-MS/MS to validate 657 secreted proteins and classify their functions (Kumari et al., 2023).
5. Engineering Signal Peptides for Recombinant Protein Production
For recombinant protein production, signal peptide selection is often the difference between milligram and gram-per-liter yields. Rational design and high-throughput screening strategies now enable systematic optimization.
5.1 Selecting Optimal Signal Peptides for Expression Systems
Signal peptide choice must align with host system biology. In mammalian cells, native human signal peptides or optimized variants (e.g., Igκ, human albumin) typically perform well, with considerations for glycosylation and authentic processing. For yeast systems, the S. cerevisiae MFα signal peptide remains the gold standard in Pichia pastoris, though engineered variants have demonstrated superior performance. In E. coli, signal peptides such as PelB, OmpA, and PhoA direct proteins to the periplasmic space, enabling oxidative folding and simplified purification. The key selection criteria include host compatibility, cleavage accuracy, secretion efficiency, product quality (folding, glycosylation), and scalability.
5.2 Key Sequence Features for Enhanced Secretion Efficiency
Rational engineering targets specific signal peptide features. H-region hydrophobicity modulates targeting pathway choice—increasing hydrophobicity enhances SRP-dependent co-translational translocation but risks improper membrane insertion or ER stress at extremes. N-region positive charge optimization improves SRP interaction and targeting fidelity. C-region cleavage site engineering ensures efficient processing, preventing signal peptide retention and ER retention. Adjacent sequences (propeptides) influence translation kinetics and cotranslational folding, affecting final product quality. The recent development of fully synthetic signal peptide libraries enables systematic exploration of sequence space (Jeon et al., 2024).
5.3 Experimental Validation: From in silico Design to Functional Readouts
A robust validation workflow integrates computational design with experimental verification. Initial in silico screening using SignalP and Phobius confirms signal peptide presence and eliminates candidates with transmembrane domains. Expression constructs are generated and evaluated through quantitative secretion assays (ELISA, Western blot of supernatants vs. lysates). Cleavage site confirmation by N-terminal sequencing or mass spectrometry validates processing fidelity. Product quality assessment includes glycosylation analysis, aggregation testing, and functional activity assays. The successful engineering of E2-Spy antigen in P. pastoris exemplifies this approach: hybrid signal peptide cSP3 increased production 3.38-fold, gene dosage optimization added another 4.38-fold improvement, and molecular chaperone co-expression (SSA1, YDJ1, BiP, LHS1) contributed an additional 1.92-fold increase, culminating in a 6.18-fold overall enhancement (Li et al., 2024).
6: Signal Peptides in Disease and Translational Omics
Signal peptide research now extends far beyond protein production, intersecting with disease mechanisms, therapeutic development, and cutting-edge analytical methods.
6.1 Secretory Pathway Dysregulation in Disease: Signal Peptides as Mechanistic Clues
Mutations in signal peptides cause human disease through multiple mechanisms. A comprehensive analysis of human genome variants identified over 11,000 pathogenic signal peptide mutations affecting more than 3,300 genes encoding secreted and membrane proteins. Most pathogenic mutations alter the hydrophobic core, potentially activating RAPP quality control and mRNA degradation. The remaining 25% affect N-regions or cleavage sites, causing translocation deficiencies or processing inhibition (Gutierrez Guarnizo et al., 2023). These findings position signal peptides as critical nodes in disease pathology, with implications for diagnostics and therapeutic intervention. Secretome analysis of pathogenic bacteria similarly reveals virulence factors and immune modulators dependent on functional signal peptides for secretion (Kumari et al., 2023).
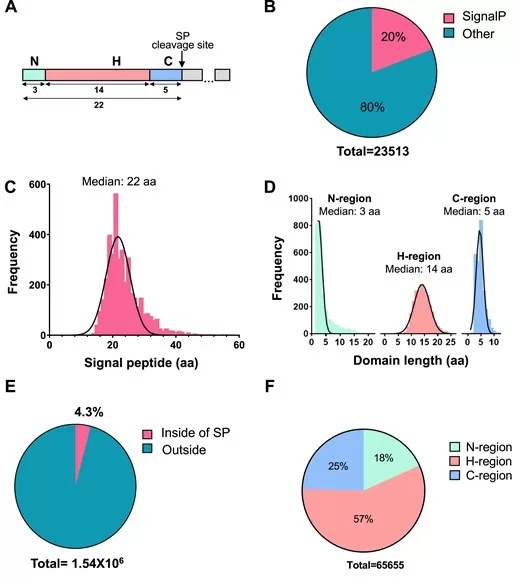
Detection of signal peptides and signal peptide missense variants at the whole human genome.
Image reproduced from Gutierrez Guarnizo et al., 2023, NAR genomics and bioinformatics, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
6.2 Therapeutic Proteins, Vaccines, and Cell Therapy: Signal Peptide as a Design Lever
Signal peptide engineering directly impacts therapeutic product development. In recombinant protein production, optimized signal peptides enhance yields of monoclonal antibodies, fusion proteins, and enzymes. For vaccine development, signal peptides ensure proper antigen presentation and secretion, critical for immunogenicity. The E2-Spy antigen for classical swine fever virus vaccine achieved 168.3 mg/L production through combined signal peptide engineering, gene dosage, and chaperone co-expression—a 6.18-fold improvement enabling practical vaccine manufacturing (Li et al., 2024). In cell therapy applications, engineered signal peptides direct therapeutic factor secretion from modified cells, enabling localized delivery and reduced systemic toxicity.
6.3 Emerging Methods: N-terminomics, Secretome Profiling, and Spatial/Single-Cell Context
Advanced methodologies are transforming signal peptide research. N-terminomics (e.g., COFRADIC, TAILS) enables proteome-wide identification of signal peptidase cleavage sites, validating predictions and revealing processing heterogeneity. Quantitative secretome profiling using SILAC or TMT labeling tracks secretion dynamics under physiological or pathological conditions. The integration of single-cell RNA sequencing with signal peptide prediction reveals cell-type-specific secretory signatures and intercellular communication networks. Spatial transcriptomics adds tissue context, localizing secretory activity within complex microenvironments.
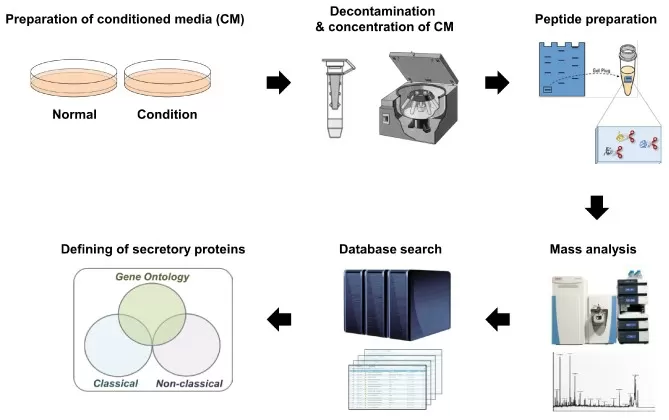
General workflow of cell secretome investigations.
Image reproduced from Song et al., 2019, International journal of molecular sciences, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
6.4 Post-Targeting Biology of Signal Peptides: Immune Signaling and Translational Opportunities
Although many cleaved signal peptides are rapidly degraded after ER targeting and signal peptidase processing, accumulating evidence shows that some signal peptide fragments persist and can be biologically meaningful. A well-established case is HLA-E presentation of MHC class I leader (signal) peptides, which are sensed by CD94/NKG2 receptors on NK cells and tune immune inhibition/activation, linking signal peptide processing to immune surveillance (Huisman et al., 2023; Middelburg et al., 2023). Translationally, reviews summarize reports of circulating signal-peptide–derived fragments such as BNPsp (17–26) after myocardial infarction as exploratory biomarker candidates (Ono, 2024), while a 2025 study proposed bacterial signal peptides as ligands for formyl peptide receptors (FPRs) in glioblastoma for imaging/targeting concepts (Al Taleb et al., 2025). These examples are context-dependent, but they broaden the view of signal peptides beyond disposable “address tags.”
Reference
1. Zhang, S., He, Z., Wang, H., & Zhai, J. (2025). Signal Peptides: From Molecular Mechanisms to Applications in Protein and Vaccine Engineering. Biomolecules, 15(6), 897. https://doi.org/10.3390/biom15060897
2. Kaushik, S., He, H., & Dalbey, R. E. (2022). Bacterial Signal Peptides- Navigating the Journey of Proteins. Frontiers in physiology, 13, 933153. https://doi.org/10.3389/fphys.2022.933153
3. Lang, S., Nguyen, D., Bhadra, P., Jung, M., Helms, V., & Zimmermann, R. (2022). Signal Peptide Features Determining the Substrate Specificities of Targeting and Translocation Components in Human ER Protein Import. Frontiers in physiology, 13, 833540. https://doi.org/10.3389/fphys.2022.833540
4. Li, B., Zheng, Y., Zhao, S., Zhang, Y., & Li, D. (2024). High-Level Secretory Production of Recombinant E2-Spy Antigen Protein via Combined Strategy in Pichia pastoris. Fermentation, 10(2), 99. https://doi.org/10.3390/fermentation10020099
5. Gutierrez Guarnizo, S. A., Kellogg, M. K., Miller, S. C., Tikhonova, E. B., Karamysheva, Z. N., & Karamyshev, A. L. (2023). Pathogenic signal peptide variants in the human genome. NAR genomics and bioinformatics, 5(4), lqad093. https://doi.org/10.1093/nargab/lqad093
6. Kumari, K., Sharma, P. K., Ma, Y., & Singh, R. P. (2023). Secretome of Paenibacillus sp. S-12 provides an insight about its survival and possible pathogenicity. Folia microbiologica, 68(4), 559–569. https://doi.org/10.1007/s12223-023-01032-4
7. Jeon, E. J., Lee, S. M., Hong, H. S., & Jeong, K. J. (2024). Design of fully synthetic signal peptide library and its use for enhanced secretory production of recombinant proteins in Corynebacterium glutamicum. Microbial cell factories, 23(1), 252. https://doi.org/10.1186/s12934-024-02516-9
8. Song, P., Kwon, Y., Joo, J. Y., Kim, D. G., & Yoon, J. H. (2019). Secretomics to Discover Regulators in Diseases. International journal of molecular sciences, 20(16), 3893. https://doi.org/10.3390/ijms20163893
9. Huisman, B. D., Guan, N., Rückert, T., Garner, L., Singh, N. K., McMichael, A. J., Gillespie, G. M., Romagnani, C., & Birnbaum, M. E. (2023). High-throughput characterization of HLA-E-presented CD94/NKG2x ligands reveals peptides which modulate NK cell activation. Nature communications, 14(1), 4809. https://doi.org/10.1038/s41467-023-40220-1
10. Middelburg, J., Ghaffari, S., Schoufour, T. A. W., Sluijter, M., Schaap, G., Göynük, B., Sala, B. M., Al-Tamimi, L., Scheeren, F., Franken, K. L. M. C., Akkermans, J. J. L. L., Cabukusta, B., Joosten, S. A., Derksen, I., Neefjes, J., van der Burg, S. H., Achour, A., Wijdeven, R. H. M., Weidanz, J., & van Hall, T. (2023). The MHC-E peptide ligands for checkpoint CD94/NKG2A are governed by inflammatory signals, whereas LILRB1/2 receptors are peptide indifferent. Cell reports, 42(12), 113516. https://doi.org/10.1016/j.celrep.2023.113516
11. Ono K. (2024). Signal Peptides and Their Fragments in Post-Translation: Novel Insights of Signal Peptides. International journal of molecular sciences, 25(24), 13534. https://doi.org/10.3390/ijms252413534
12. Al Taleb, Z., Hierlmeier, I., Heilmann, H., Jung, M., Bartholomä, M., & Bufe, B. (2025). Assessing the potential of bacterial signal peptides for radiopharmaceutical applications. Scientific reports, 15(1), 32479. https://doi.org/10.1038/s41598-025-18831-z
Next-Generation Omics Solutions:
Proteomics & Metabolomics
Ready to get started? Submit your inquiry or contact us at support-global@metwarebio.com.


