Spatial Metabolomics for Biomarker Discovery in Precision Medicine
1. Why Spatial Metabolomics Is the Future of Biomarkers
Conventional metabolomics—most commonly bulk LC–MS(/MS) on homogenized tissue—has delivered powerful insights into disease-associated metabolic shifts. Yet it carries a fundamental limitation: it collapses a complex tissue into a single averaged profile, obscuring heterogeneity that is often central to pathology and therapy response [1]. In practice, a “tumor sample” is rarely just tumor cells; it can include stromal fibroblasts, endothelial cells, infiltrating immune cells, extracellular matrix, hemorrhage, necrosis, and variable perfusion. When these components are mixed, metabolic signals from small but decisive niches can be diluted or lost [1].
That blind spot matters because tissues are spatially organized ecosystems. Tumors, for example, can show striking metabolic divergence between the core vs. invasive margin, immune-infiltrated vs. immune-excluded regions, hypoxic vs. well-perfused zones, and necrotic vs. fibrotic areas—each shaped by local oxygen tension, nutrient access, pH, inflammation, and cell–cell interactions [2,3]. These microenvironments are not passive: they can drive cancer cell plasticity, immune suppression, drug resistance, and relapse dynamics [2,3].
This leads to a practical shift in biomarker thinking: A biomarker is not only “what changes”, but also “where it changes”.
Spatial metabolomics addresses that gap by preserving tissue architecture while measuring metabolite distributions in situ, enabling biomarkers that align directly with microenvironmental biology and clinical pathology [1,3].
2. What Is Spatial Metabolomics: Visualizing Metabolic Landscapes Within Tissues
Spatial metabolomics is the spatially resolved measurement of metabolite distributions within a tissue section, producing metabolic maps that link molecular variation to anatomical and pathological structure [1,4]. The best-established engine for this is mass spectrometry imaging (MSI), which generates a mass spectrum at each pixel (or sampling point) and reconstructs ion images—effectively converting invisible chemistry into interpretable spatial patterns [1,4].
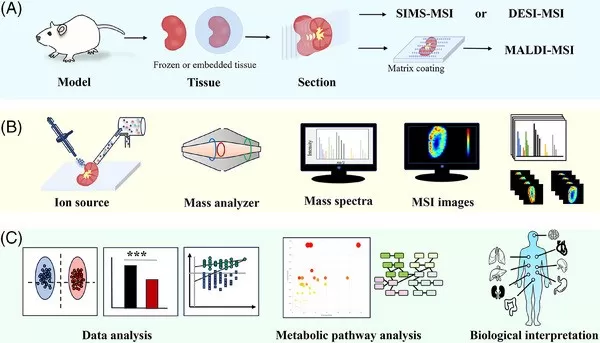
Typical workflow for spatial metabolomics.
Image reproduced from Min, X., Zhao, Y., Yu, M. et al., 2024, Clinical and translational medicine, licensed under the Creative Commons Attribution License (CC BY 4.0).
2.1 How Mass Spectrometry Imaging (MSI) works
A typical MSI-based spatial metabolomics workflow begins with tissue embedding, sectioning, and mounting to preserve tissue architecture and in situ molecular distributions. The section is then prepared in a method-dependent manner—for example, MALDI-MSI typically requires matrix deposition, while some ambient ionization approaches may involve lighter surface preparation. Next, the instrument raster-scans the tissue and acquires a mass spectrum at each pixel, generating a dense grid of spatially indexed molecular profiles. Finally, computational pipelines reconstruct selected m/z features into ion images (metabolite heatmaps) and extract region-enriched signatures, enabling systematic comparison of metabolic patterns across distinct tissue areas [1,4].
2.2 Key Technologies in Spatial Metabolomics
1) MALDI-MSI (Matrix-Assisted Laser Desorption/Ionization MSI)
MALDI-MSI relies on a UV laser to desorb molecules from the tissue surface, while a co-crystallized matrix layer absorbs the laser energy and promotes soft ionization. As the laser raster-scans across the section, a full mass spectrum is recorded at each pixel, enabling spatial maps of many analytes in parallel. MALDI-MSI is widely used for lipids and a broad range of small molecules, with ongoing improvements in spatial resolution and molecular coverage [3]. Common considerations include matrix-related effects and the persistent challenge of confident metabolite annotation in complex tissue backgrounds [1,3].
2) DESI-MSI (Desorption Electrospray Ionization MSI)
DESI-MSI is an ambient ionization method: a charged solvent spray is directed onto the tissue, where droplets impact the surface, dissolve surface molecules, and eject secondary droplets carrying ionized analytes into the mass spectrometer. Because it operates under atmospheric conditions and can require relatively light sample preparation, DESI-MSI is attractive for rapid metabolic and lipid mapping while maintaining tissue integrity for parallel workflows [4]. Its practical value is often strongest when spatial chemical information needs to be obtained with minimal disruption to the tissue section [4].
3) High-Resolution and Multi-Modal Spatial Omics
High-resolution spatial metabolomics pushes MSI toward finer pixel sizes and sharper anatomical localization, but an equally important trend is multi-modal integration—computationally aligning MSI with histology and other spatial layers (e.g., transcriptomics or proteomics) to connect metabolite patterns with cell states and tissue context [5]. In this framework, MSI provides the spatial chemistry, while complementary modalities supply cellular identity and pathway regulation, making biomarker signals more interpretable and mechanistically grounded than ion images alone [5,8].
2.3 What Spatial Metabolomics Data Looks Like
Spatial metabolomics provides more than just heatmaps of metabolites; it reveals how metabolism varies across tissue regions. Reports typically start with an average mass spectrum and ion images that show the spatial distribution of key metabolites. The tissue is then divided into distinct regions based on molecular differences, with visualizations like t-SNE/UMAP making these patterns easy to interpret. Further analysis identifies which metabolites co-occur in specific areas, helping uncover coordinated metabolic programs. Finally, ROI-based differential analysis highlights metabolic differences between regions and links them to biological pathways, providing valuable insights for biomarker discovery [1,4] .
Comparison of Key Spatial Metabolomics Techniques
|
Feature |
MALDI-MSI |
DESI-MSI |
High-Resolution & Multi-Modal MSI |
|
Matrix-assisted UV laser ionization |
Ambient solvent spray ionization |
Combines MSI with other spatial omics |
|
|
Requires matrix application |
Minimal preparation, direct tissue analysis |
High-resolution, integrated multi-modal approach |
|
|
Moderate, with ongoing improvements |
Moderate, with some limitations |
High, with finer pixel and anatomical localization |
|
|
Analysis Speed |
Slower, due to matrix and laser scanning |
Faster, minimal sample prep |
Varies, typically higher throughput with integrated analysis |
|
Applications |
Lipids, small molecules, broad metabolite coverage |
Rapid profiling of metabolites and lipids |
Comprehensive analysis combining metabolism, histology, and gene expression |
|
Strengths |
High sensitivity, broad molecular range |
Fast, minimal disruption to tissue |
Enhanced data richness through multi-layer integration |
|
Challenges |
Matrix effects, complex metabolite annotation |
Limited metabolite coverage, ion suppression |
Requires complex data integration and alignment |
|
Clinical Relevance |
Well-suited for cancer, lipidomics, and complex small molecules |
Ideal for on-site, rapid clinical profiling |
Provides in-depth tissue and microenvironmental insights for biomarker discovery and disease stratification |
3. How Spatial Metabolomics Enhances Biomarker Discovery
3.1 Capturing Microenvironment-Specific Biomarkers
Bulk profiling often underestimates metabolites that are highly enriched in narrow niches—such as the tumor–immune interface, hypoxic rims, inflammatory foci, or peri-vascular regions—because these areas may represent only a small fraction of the total sample [2,3]. Spatial metabolomics preserves localization, enabling biomarkers defined by both abundance and spatial context, which is often closer to the biology driving disease behavior [2,3].
A concrete example comes from nasopharyngeal carcinoma (NPC): an integrated spatial metabolomics + spatial transcriptomics study reported that high PD-1 expression is associated with spatially organized metabolic reprogramming—including patterns in lipid metabolism, branched-chain amino acid (BCAA) metabolism, and glutamine metabolism—and linked these programs to therapeutic response [7]. This illustrates a key advantage: spatial metabolomics can identify biomarkers that are not merely “up” or “down”, but enriched in clinically meaningful microenvironments (e.g., tumor–immune junction regions) [7].

Spatial Metabolic Reprogramming in Tumor Microenvironment: BCAA and Glutamine Pathways.
Image reproduced from Ji, L., Wang, D., Zhuo, G. et al., 2025, Theranostics, licensed under the Creative Commons Attribution License (CC BY 4.0).
3.2 Enabling Disease Stratification and Subtype Discovery
Patients can share the same histopathological diagnosis yet exhibit different spatial metabolic architectures—differences that may map to prognosis or therapy response more faithfully than bulk averages [1,6]. Spatial metabolic signatures can therefore serve as stratification features, capturing heterogeneity that reflects real tissue organization.
In kidney disease contexts, spatial metabolomics and multi-omics integration have been described as enabling precision-medicine-oriented classification by connecting metabolite patterns to specific renal structures and disease compartments (e.g., glomeruli vs tubules; fibrotic vs inflamed regions), supporting more granular phenotyping [6]. Importantly, this kind of structural anchoring helps interpret “why” a biomarker is present and “what tissue process” it reflects [6].
3.3 Improving Translational Potential: Aligning Biomarkers with Pathological Structures
The ability to integrate spatial metabolomics with traditional pathology is a key advantage for clinical translation, as it allows metabolic maps to be co-registered with H&E and IHC staining images. This helps researchers correlate metabolic features with tissue structures such as tumor margins, immune-rich regions, necrotic cores, or fibrotic stroma. This spatial interpretability enhances clinical confidence and simplifies follow-up validation experiments (e.g., targeted LC–MS analysis on microdissected regions) [3,4,6]. However, it is important to note that co-registration with H&E and IHC is an advanced feature in spatial metabolomics and is not available in all workflows. Achieving this requires high-resolution imaging technologies and specialized data integration tools, which are typically not included in routine spatial metabolomics studies.
4. Practical Applications of Spatial Metabolomics in Precision Medicine
4.1 Diagnosis: Enhancing Pathological Interpretation and Subtyping
Spatial metabolomics can add a chemical dimension to routine tissue assessment by revealing metabolic boundaries and region-specific biochemical phenotypes that may be difficult to infer from morphology alone. In oncology, metabolite maps can help differentiate adjacent but biologically distinct areas—such as invasive margins versus tumor core, infiltrative fronts versus stromal compartments, or necrotic versus fibrotic regions—where the local metabolic state may reflect oxygenation, nutrient availability, and cell composition [3,4]. Importantly, spatial patterns can also guide how a tissue is partitioned for downstream testing: instead of treating the section as one entity, analysts can define data-driven regions and extract region-enriched features for more precise diagnostic signatures [3,4]. In nephrology, spatial metabolomics has been described as a high-dimensional diagnostic layer because it links molecular changes to renal microstructures (e.g., glomerular and tubular compartments), enabling compartment-level interpretation that is closer to the true anatomical unit of kidney pathology [6]. Taken together, these capabilities support a diagnostic workflow in which classification is informed not only by “what is present,” but by how molecular phenotypes are organized across tissue structures [3,4,6].
4.2 Predicting Treatment Efficacy: Forecasting Response to Immunotherapy and Targeted Therapy
Treatment response—particularly for immunotherapy—often depends on the metabolic constraints and resources within the tumor microenvironment. Metabolites are not merely passive readouts; they can influence immune activation, exhaustion, and cellular fitness, meaning that where metabolic suppression or enrichment occurs can be as important as whether it occurs [2]. Spatial profiling helps pinpoint metabolic stress or nutrient competition at the tumor–immune interface, avoiding the dilution effect of bulk measurements and enabling biomarker concepts that are explicitly microenvironment-aware [2]. In nasopharyngeal carcinoma (NPC), spatially resolved metabolic reprogramming linked to PD-1–associated biology has been connected to response and resistance patterns, illustrating how spatial metabolic signatures can function as predictive biomarkers rather than simple global “up/down” differences [7]. Practically, this supports response-prediction strategies that combine regional metabolic features (e.g., interface-associated signals or hypoxia-associated patterns) with patient stratification—providing a more mechanistic basis for predicting who benefits from immunotherapy or targeted regimens [2,7].
4.3 Optimizing Therapeutic Strategies: Targeting Local Metabolic Dependencies in Precision Medicine
Beyond prediction, spatial metabolomics can directly inform treatment strategy by identifying localized metabolic dependencies—pathways that are disproportionately dysregulated in specific regions and may represent intervention opportunities. This is particularly relevant when a pathway is not uniformly altered across the tissue but becomes pronounced only in certain microenvironments (for example, at invasive fronts, peri-vascular niches, or immune-excluded territories), where selective pressure and resource limitation shape biology [3,4]. In NPC, the reported spatial reprogramming involving lipid, BCAA, and glutamine-related metabolism—supported by follow-up validation—illustrates how spatial findings can be translated into mechanistic hypotheses and therapeutic targeting directions [7]. More broadly, spatial multi-omics work in glioblastoma reinforces that tumor–host interdependence is spatially organized, suggesting that effective intervention may require understanding where the relevant metabolic interactions occur, not just which genes or metabolites differ overall [8]. In translational practice, spatial metabolomics can therefore act as a “target discovery compass,” prioritizing region-defined pathways for downstream confirmation and functional testing [7,8].
In a practical pipeline, spatial metabolomics discoveries can feed into: targeted LC–MS assays for absolute quantification in selected regions, orthogonal validation in tissue-matched compartments, mechanistic studies (e.g., pathway perturbation or nutrient modulation), and iterative refinement toward clinically robust biomarkers [1,6,7].
5. Challenges and Future Trends in Spatial Metabolomics
5.1 Challenges in Spatial Metabolomics: Throughput, Identification, and Integration
Throughput and speed
High spatial resolution typically increases acquisition time, computational load, and data-management complexity. These constraints can be nontrivial when scaling to large cohorts or clinical timelines [4,5].
Metabolite identification (especially isomers)
Assigning confident metabolite identities in MSI remains challenging due to complex backgrounds, ion suppression, adduct formation, and the difficulty of distinguishing isomers/isobars. Identification is often the rate-limiting step for turning spatial features into mechanistic biomarkers [1,5].
Multi-modal integration and analytics
While multi-modal spatial omics is powerful, integrating MSI with other spatial layers introduces algorithmic challenges—registration accuracy, resolution mismatch, batch effects, and interpretability of fused features [5,6].
5.2 Future Trends in Spatial Metabolomics: Single-Cell, 3D Imaging, and Clinical Application
Single-cell and subcellular resolution
As spatial metabolomics continues to evolve, single-cell and subcellular resolution will become increasingly important. This will allow for an even more detailed mapping of metabolic interactions within individual cells, uncovering how cellular microenvironments influence metabolic behavior. It will also enable researchers to examine rare cell populations, such as tumor stem cells or infiltrating immune cells, that may play crucial roles in disease progression but are often masked in bulk analyses [2,3]. Single-cell resolution will provide unprecedented insights into cellular heterogeneity and the ways in which different cells within the same tissue interact metabolically, ultimately advancing our understanding of disease at the most granular level.

From Bulk to Single-Cell Analysis: Advancements in Understanding Cellular Heterogeneity.
Image reproduced from Lee, S., Vu, H. M., Lee, J. H. et al., 2023, Biology, licensed under the Creative Commons Attribution License (CC BY 4.0).
3D spatial metabolomics
Current spatial metabolomics typically works with thin tissue sections, providing a 2D snapshot of metabolic variation. However, 3D spatial metabolomics is an emerging frontier that aims to overcome this limitation. By reconstructing metabolic profiles across tissue volumes, this technology will allow for a more accurate representation of organ-level architecture and metabolic gradients. It will be particularly beneficial in understanding complex, dynamic processes like tumor invasion, where the full depth of tissue needs to be considered to capture the full scope of metabolic alterations. This shift towards three-dimensional mapping will also help improve drug delivery models and guide the development of new therapies by revealing how drugs interact with the tissue at different depths [4].
Clinical translation
For spatial metabolomics to make a significant impact in clinical settings, clinical translation will rely on the standardization of protocols, from sample collection and tissue handling to data analysis. Ensuring consistent quality control (QC) across samples, as well as high annotation confidence, will be crucial for generating reproducible, interpretable results. Additionally, computational pipelines that integrate and analyze multi-modal spatial data (including metabolomics, histology, and transcriptomics) must be robust, scalable, and capable of delivering clinically actionable insights that are directly aligned with pathology workflows. As these areas continue to improve, spatial metabolomics will become an increasingly reliable tool for diagnosis, treatment planning, and monitoring response to therapy in the clinical space [1,6].
References:
1. Min, X., Zhao, Y., Yu, M., Zhang, W., Jiang, X., Guo, K., Wang, X., Huang, J., Li, T., Sun, L., & He, J. (2024). Spatially resolved metabolomics: From metabolite mapping to function visualising. Clinical and translational medicine, 14(11), e70031. https://doi.org/10.1002/ctm2.70031
2. Berrell, N., Sadeghirad, H., Blick, T., Bidgood, C., Leggatt, G. R., O'Byrne, K., & Kulasinghe, A. (2024). Metabolomics at the tumor microenvironment interface: Decoding cellular conversations. Medicinal research reviews, 44(3), 1121–1146. https://doi.org/10.1002/med.22010
3. Planque, M., Igelmann, S., Ferreira Campos, A. M., & Fendt, S. M. (2023). Spatial metabolomics principles and application to cancer research. Current opinion in chemical biology, 76, 102362. https://doi.org/10.1016/j.cbpa.2023.102362
4. Ma, X., & Fernández, F. M. (2024). Advances in mass spectrometry imaging for spatial cancer metabolomics. Mass spectrometry reviews, 43(2), 235–268. https://doi.org/10.1002/mas.21804
5. Santos, A. A., Delgado, T. C., Marques, V., Ramirez-Moncayo, C., Alonso, C., Vidal-Puig, A., Hall, Z., Martínez-Chantar, M. L., & Rodrigues, C. M. P. (2024). Spatial metabolomics and its application in the liver. Hepatology (Baltimore, Md.), 79(5), 1158–1179. https://doi.org/10.1097/HEP.0000000000000341
6. Sharma, K., Hansen, J., Susztak, K., Eberlin, L., Anderton, C. R., Alexandrov, T., & Iyengar, R. (2026). Spatial metabolomics and multiomics integration for breakthroughs in precision medicine for kidney disease. Nature reviews. Nephrology, 22(2), 152–164. https://doi.org/10.1038/s41581-025-01007-3
7. Ji, L., Wang, D., Zhuo, G., Chen, Z., Wang, L., Zhang, Q., Wan, Y., Liu, G., & Pan, Y. (2025). Spatial Metabolomics and Transcriptomics Reveal Metabolic Reprogramming and Cellular Interactions in Nasopharyngeal Carcinoma with High PD-1 Expression and Therapeutic Response. Theranostics, 15(7), 3035–3054. https://doi.org/10.7150/thno.102822
8. Ravi, V. M., Will, P., Kueckelhaus, J., Sun, N., Joseph, K., Salié, H., Vollmer, L., Kuliesiute, U., von Ehr, J., Benotmane, J. K., Neidert, N., Follo, M., Scherer, F., Goeldner, J. M., Behringer, S. P., Franco, P., Khiat, M., Zhang, J., Hofmann, U. G., Fung, C., … Heiland, D. H. (2022). Spatially resolved multi-omics deciphers bidirectional tumor-host interdependence in glioblastoma. Cancer cell, 40(6), 639–655.e13. https://doi.org/10.1016/j.ccell.2022.05.009
9. Lee, S., Vu, H. M., Lee, J. H., Lim, H., & Kim, M. S. (2023). Advances in Mass Spectrometry-Based Single Cell Analysis. Biology, 12(3), 395. https://doi.org/10.3390/biology12030395
Next-Generation Omics Solutions:
Proteomics & Metabolomics
Ready to get started? Submit your inquiry or contact us at support-global@metwarebio.com.


