From "Average" to "Tissue Maps": The Evolution and Future Trends of Spatial Metabolomics
Spatial metabolomics is changing how we ask questions in biology. For years, mainstream metabolomics has excelled at telling us ‘what’ metabolites are present and ‘how much’ their levels change across experimental groups. Yet most traditional workflows begin with a step that silently erases a crucial dimension: we homogenize a tissue, extract metabolites, and generate one averaged profile for a sample that may contain many cell types, microenvironments, and physiological states. In well-mixed systems this is acceptable—but in complex tissues and diseases, the “average” can be misleading.
Spatial metabolomics offers a different promise: it keeps metabolism in its native context. Instead of turning a tissue into a single number or a single list, it turns metabolism into an interpretable ‘map’—a view of biochemical organization across tumor cores and invasive margins, across brain regions and lesion boundaries, across infection foci and host response zones, and across plant tissues with specialized functions. This spatial dimension is not decorative. In many biological systems, ‘where’ a metabolite accumulates is inseparable from ‘what it means’.
In this article, we follow a practical storyline:
(1) why spatial context is the next paradigm upgrade in metabolomics;
(2) how the technology evolved through distinct stages;
(3) where spatial metabolomics becomes irreplaceable;
(4) the most credible trends shaping the next decade;
(5) how to design a spatial metabolomics project that yields convincing, actionable deliverables.
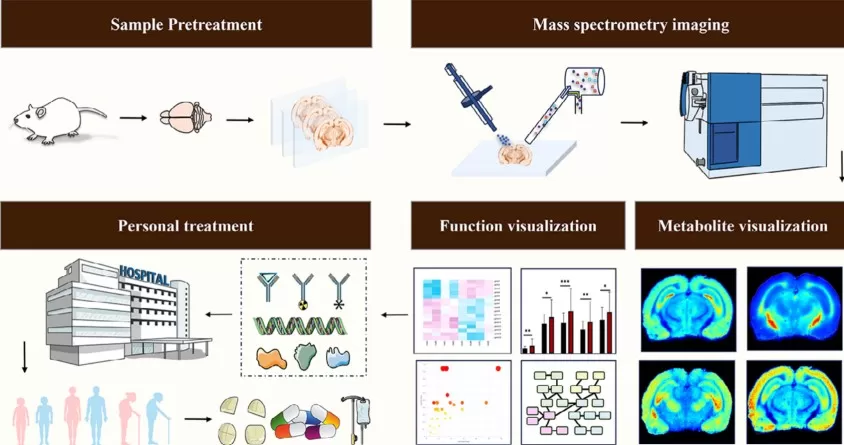
Mass spectrometry imaging analysis of metabolites
Image reproduced from Min et al., 2024, Clin Transl Med, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
1. Why “Spatial” Is the Next Paradigm Shift in Metabolomics
1.1 The “Average-Value Trap” of Bulk Metabolomics
Bulk metabolomics typically analyzes homogenized tissue extracts. This approach is powerful for sensitivity and throughput, but it collapses heterogeneous microregions into a single composite signal. If a tumor contains a hypoxic core, a well-perfused rim, necrotic patches, fibrotic compartments, and immune-rich interfaces, bulk extraction merges them into one mean profile. Metabolic gradients—oxygen, pH, nutrients, drug exposure, immune infiltration—are smoothed out. The same problem appears in many settings: inflamed tissues with patchy immune aggregates, organs with clear zonation, brain tissues with region-specific metabolism, and microbial colonization that is highly localized [1-2].
Spatial metabolomics addresses this by measuring metabolic distributions ‘in situ’. Conceptually, it upgrades metabolomics from a one-dimensional measurement (“how much?”) into a two-dimensional (and increasingly three-dimensional) representation (“how much—and where?”). The result is often a qualitative leap in interpretability: metabolism becomes linked to tissue architecture and microenvironmental structure rather than floating as context-free numbers.
1.2 What Spatial Metabolomics Solves: Three Core Advantages
Spatial metabolomics is not simply “metabolomics with pictures.” Its value can be summarized in three core advantages:
(1) Revealing heterogeneity with biological meaning. Spatial maps allow researchers to see metabolic “hotspots” and boundaries—areas where metabolites sharply accumulate, deplete, or transition. In cancer, this can expose metabolic subregions associated with aggressiveness or resistance that bulk profiles average away. Reviews of cancer-focused spatial metabolomics consistently emphasize that intratumoral metabolic heterogeneity is not noise; it is often the biology that explains clinical behavior [3].
(2) Capturing microenvironment interactions. Metabolism is a language of interaction. Tumor cells exchange nutrients with stromal cells; immune cells can be metabolically suppressed by local metabolites; microbes can reshape host metabolism in their immediate niche. Spatial metabolomics makes these interactions visible because it measures metabolites in the actual anatomical neighborhoods where cells interact [4].
(3) Enabling translational and pharmacological insight. Drugs do not distribute uniformly. Even when systemic dosing is stable, local exposure and local metabolic consequences vary across tissue regions. Spatial metabolomics can image drug molecules, their metabolites, and endogenous metabolic changes in the same section, supporting spatial PK/PD reasoning and aiding toxicology and mechanism studies [5].
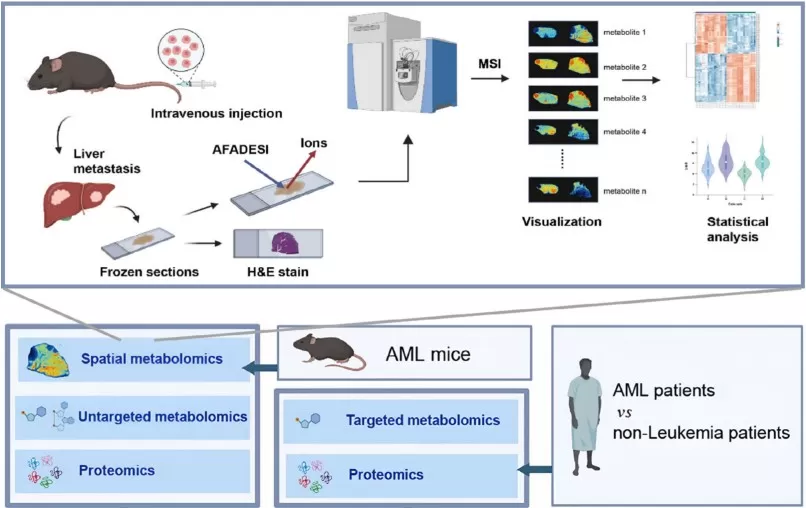
Spatial metabolomic techniques identify metabolic reprogramming features
2. Technological Evolution: How Spatial Metabolomics Advanced Step by Step
Spatial metabolomics progressed through iterative solutions to predictable bottlenecks: preserving native chemistry, improving ionization efficiency, increasing spatial resolution and coverage, strengthening metabolite identification, and scaling analysis pipelines. A useful framework is to view this evolution in four stages.
2.1 Stage 1: MSI Makes Metabolism “Visible” (From 0 to 1)
The first breakthrough was simply to make metabolites visible in tissue without homogenization. Mass spectrometry imaging (MSI) enables this by rastering across tissue sections and acquiring a mass spectrum at each pixel; computational reconstruction then produces spatial intensity maps for selected ions. Early MSI approaches demonstrated that metabolism could be visualized directly on thin sections and that such maps revealed patterns inaccessible to bulk assays [6].
At this stage, limitations were clear: sensitivity was often insufficient for broad metabolite coverage; spatial resolution was modest for many platforms; and metabolite identification remained challenging because imaging lacks chromatographic separation. Still, Stage 1 established the central proof: the metabolome is spatially organized, and mapping it produces new biological questions.
2.2 Stage 2: Higher Sensitivity, Coverage, and Resolution (From “Seeing” to “Seeing Clearly and Completely”)
Once imaging was feasible, the field focused on seeing more compounds with better detail. Improvements in ion sources, sampling strategies, and instrument performance drove a rapid expansion in detectable features and spatial fidelity.
One representative direction is chemistry-assisted signal enhancement. For instance, lithium-doped nanospray desorption electrospray ionization (nano-DESI) has been used to expand spatial metabolomics coverage by boosting ionization for metabolite classes that can be weak in standard modes, improving both sensitivity and breadth. Another representative milestone is scaling spatial metabolomics toward whole-organism or large-sample mapping. Ambient mass spectrometry imaging workflows have been demonstrated for mapping the global molecular landscape in whole-body zebrafish, highlighting how methodological improvements can support large-scale spatial profiling rather than only small ROIs.
Equally important, data processing matured. As experiments began generating large numbers of pixels and spectral features, robust preprocessing, denoising, normalization, and visualization became essential. Stage 2 effectively transformed MSI from a proof-of-concept into a reliable engine for discovering spatial metabolic patterns in diverse tissues.
2.3 Stage 3: MSI + Micro-Omics (From “Localization” to “Identification, Quantification, and Interpretation”)
As spatial datasets grew, two recurring concerns became unavoidable: identification (what is this peak?) and quantification (how much is it, really?). Stage 3 responded by integrating MSI with complementary microregion “deep dive” methods.
A common workflow became: do MSI first to obtain the panoramic map; select regions of interest (ROIs) based on spatial patterns; then perform deeper chemical characterization on those ROIs using orthogonal methods such as CE-MS or LC–MS/MS. A representative platform combining MSI with in-depth chemical characterization via capillary electrophoresis improved confidence in metabolite identification and strengthened mechanistic interpretation by providing richer chemical evidence than imaging alone [7-8].
In parallel, the community began developing more standardized and automated computational pipelines for annotation and quantitation. MSIpixel, for example, represents an effort toward an automated pipeline for compound annotation and quantitation in MSI experiments—an important step for reproducibility and scaling spatial metabolomics to larger studies [9].
In practice, Stage 3 is what turns a spatial map into a defensible mechanistic story: the map guides discovery, while ROI-targeted deep characterization verifies identities and supports pathway-level interpretation.
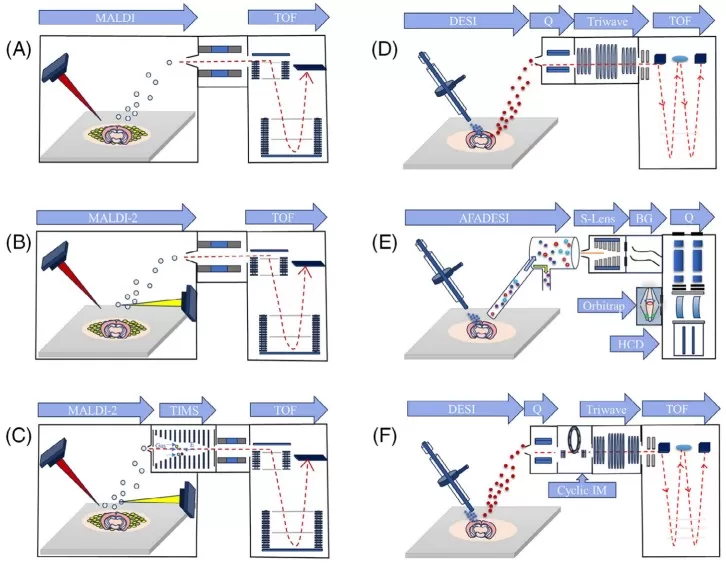
MSI instrumentation and platforms used for spatially resolved metabolomics
2.4 Stage 4: Spatial Multi-Omics Integration (From “Single Maps” to “Mechanistic Networks”)
The current frontier is integration with other spatially resolved modalities—especially spatial transcriptomics and spatial proteomics—so that metabolite patterns can be connected to gene programs, protein activity, cell phenotypes, and histology.
This integration is driven by a simple reality: metabolic phenotypes often reflect upstream regulation and downstream functional consequences. Spatial multi-omics moves the field from “metabolite distributions” to “mechanistic networks in space.” One representative direction is integrating ambient ionization MSI with spatial transcriptomics on the same cancer tissues to identify RNA–metabolite correlations, enabling region-specific links between metabolic states and transcriptional programs [7]. Another representative example is the integrative spatially resolved proteomic and metabolomic imaging work in osteoarthritis, which connects spatial molecular patterns to pathological endotypes and disease progression trajectories [10]. Stage 4 also brings new computational demands: co-registration, alignment across modalities, and models that can fuse heterogeneous spatial resolutions into a coherent interpretation.
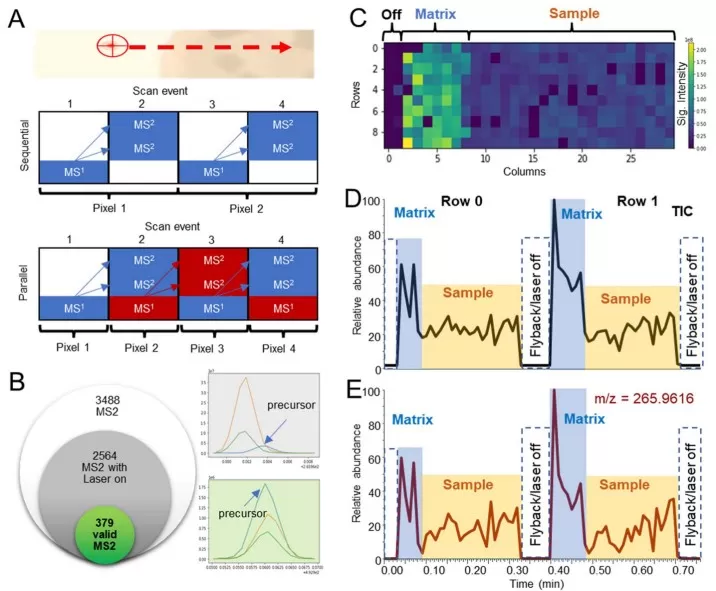
DDA principles and matrix effect in MSI
Image reproduced from Morosi et al., 2023, Brief Bioinform, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
3. Why Spatial Metabolomics Is Irreplaceable in Key Research Areas
Spatial metabolomics is most transformative in systems where location is inseparable from function. Below are five domains where spatial metabolomics has moved from “nice to have” to “hard to replace.”
3.1 Cancer and the Immune Microenvironment
Tumors are mosaics. Hypoxic zones, invasive fronts, immune-rich interfaces, and necrotic cores can exist within millimeters, each with different metabolic pressures and therapeutic vulnerabilities. Bulk homogenization hides gradients and niche behavior, masking drug-resistant compartments and immunosuppressive metabolic regions, while spatial metabolomics visualizes metabolic heterogeneity aligned with tissue architecture.
3.2 Neuroscience and Brain Disorders
The brain is highly regionalized, and many disorders have focal lesions and gradient-like penumbras. Spatial metabolomics maps neurotransmitter-related metabolites, energy metabolites, and lipids across anatomical regions and lesion margins, linking biochemistry to neuroanatomy. Broader spatial metabolomics reviews describe how this spatial view supports mechanistic interpretation of brain pathology and functional disruption [5].
3.3 Infection and Host–Microbe Metabolic Interactions
Spatial metabolomics can reveal localized metabolite production and diffusion halos, and—when integrated with other spatial assays—can connect metabolite patterns to microbial presence or host cell programs [5]. Co-localization of pathogen-associated metabolites and host defense metabolites, mapping of nutrient depletion zones, and ROI-targeted validation to confirm identities[8].
3.4 Drug Development, Distribution, and Toxicology
Drug efficacy and toxicity depend on local exposure and local metabolic response. While tissue homogenates erase drug penetration gradients and localized toxicity signatures, MSI can map parent drug, metabolites, and endogenous metabolic perturbations in the same tissue section, supporting spatial PK/PD reasoning and revealing where drugs fail to reach or where toxicity risk concentrates [5].
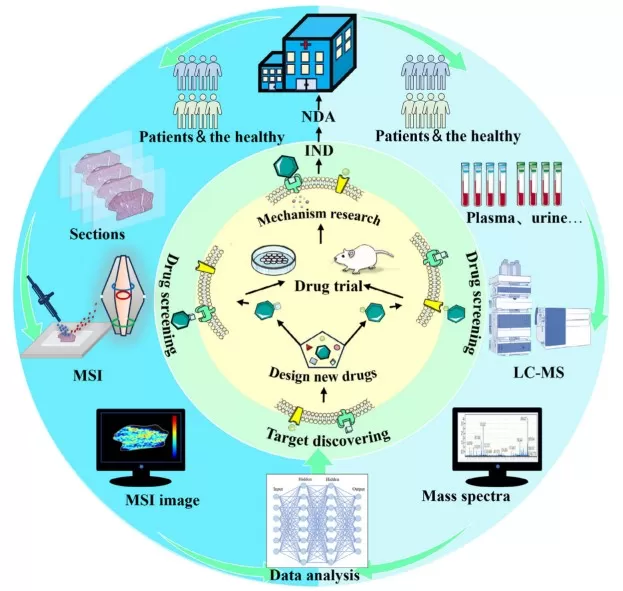
LC‒MS-based metabolomics and spatially resolved metabolomics in the new drug development process.
Image reproduced from Min et al., 2024, Clin Transl Med, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
3.5 Plants and Agriculture
Plant metabolism is compartmentalized across tissues with distinct roles (transport, defense, storage, photosynthesis). Grinding destroys compartment information and obscures tissue-specific chemistry. Spatial metabolomics reveals where secondary metabolites, phytohormones, and nutrients accumulate, directly linking metabolite location to biological function and agricultural traits [5].
4. Future Trends: Where Spatial Metabolomics Is Headed Next
Spatial metabolomics is advancing quickly, but several trends stand out as the most likely to shape near- to mid-term progress.
4.1 Higher Resolution Toward Cellular and Subcellular Mapping
The push toward single-cell and subcellular resolution will continue, constrained primarily by sensitivity and ion yield at tiny pixel sizes. Improvements in ion sources, sampling probes, and detection efficiency will be central [5].
4.2 Enhanced Sensitivity and Broader Coverage
Signal-boosting strategies—such as lithium doping in nano-DESI—illustrate how chemical and instrumental innovations can expand metabolite coverage and reduce missingness [1]. Similar strategies will proliferate.
4.3 More Reliable Metabolite Identification
Hybrid approaches combining imaging with orthogonal separation and characterization will remain critical. Platforms integrating MSI with CE-MS-like deep characterization address identification bottlenecks directly.[8] Computational tools and automated pipelines will become standard [9].
4.4 From Relative to More Quantitative Spatial Maps
Clinical translation demands quantification and reproducibility. Automated workflows for quantitation and standardized reporting will be increasingly important, building on directions represented by annotation/quantitation pipelines [9].
4.5 “Panoramic MSI + ROI Deep Dive” Becomes Routine
A pragmatic workflow will dominate: map broadly, then validate deeply in selected ROIs with orthogonal methods [8]. This reduces false discoveries and strengthens mechanistic claims.
4.6 Spatial Multi-Omics Integration Becomes the Default
Integrating metabolites with spatial transcriptomics and proteomics will increasingly be expected rather than exceptional, enabling region-specific mechanistic inference (RNA–metabolite correlations, protein–metabolite coupling) [7,10].
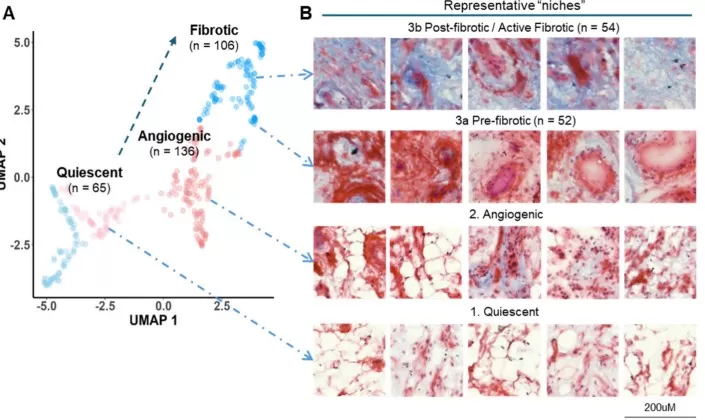
Spatially resolved proteomic and metabolomic imaging reveals synovitis endotypes
Image reproduced from Zhu et al., 2025, Theranostics, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
4.7 AI-Driven Analysis and Interpretation
As spatial datasets grow (more pixels, more modalities), AI will become essential for segmentation, clustering, co-registration, and prioritization. Automated pipelines are early indicators of this direction [9].
4.8 Standardization and Clinical Translation
Standard operating procedures, reference standards, and quality-control metrics will determine whether spatial metabolomics becomes routine in pathology and drug development. Broad reviews emphasize that standardization is a prerequisite for reliable translation [5].
5. How to Run a Successful Spatial Metabolomics Project
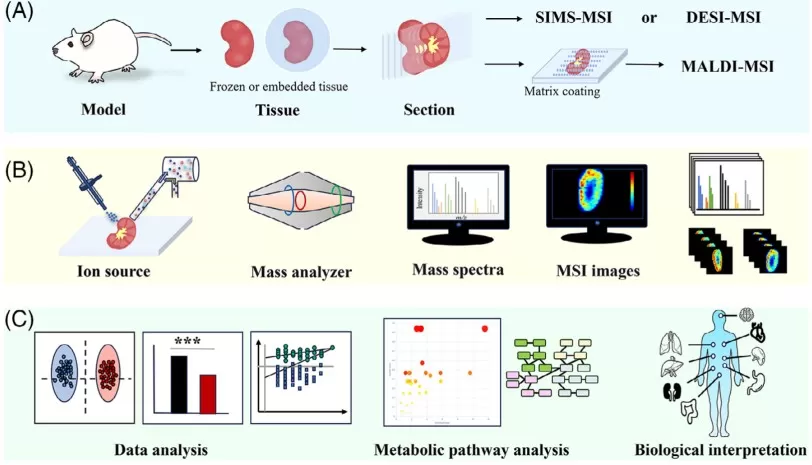
Typical workflow for spatially resolved metabolomics
Image reproduced from Min et al., 2024, Clin Transl Med, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
Spatial metabolomics projects succeed when they begin with a clear question, match the question to the correct technology, and plan validation and deliverables early.
5.1 Define the Question and Choose the Appropriate Workflow
Start with the spatial scale you must resolve: organ-level zonation, microenvironment-level niches, or near-cellular resolution. Define the chemistry: polar metabolites, lipids, xenobiotics, or untargeted coverage. Use this to choose the best MSI workflow and whether you need integration with spatial transcriptomics/proteomics [6,7].
5.2 Preserve Spatial Chemistry: Sample Preparation Is the Highest-Leverage Step
Metabolites can diffuse and degrade. Freeze quickly, keep cold, minimize thaw cycles, section consistently, and plan adjacent sections for multi-omics alignment when necessary. The best downstream analysis cannot rescue poor spatial preservation.
5.3 Plan Identification and Validation Up Front
Untargeted MSI will yield many features. Decide in advance how you will confirm top findings: ROI selection, orthogonal MS/MS, or micro-omics characterization methods [8]. Automated annotation pipelines can support consistent reporting across samples [9].
5.4 Define Deliverables Clearly
A strong deliverable package typically includes: high-quality spatial heatmaps for key metabolites (with normalization clarity), ROI-based quantitative comparisons, confident metabolite IDs (supported by MS/MS or orthogonal evidence), pathway and biological interpretation tied to histology, and, when relevant, integrated spatial multi-omics overlays [7,10].
Conclusion: A Map-Based Future for Metabolism
Spatial metabolomics is transforming metabolomics from an averaging science into a mapping science. Its core contribution is a change in what questions we can credibly ask—and answer. By revealing metabolic heterogeneity, microenvironment interactions, and localized pharmacology, it makes metabolism interpretable in the same coordinate system as histology and tissue architecture. The technology has progressed through clear stages: from making metabolites visible, to improving sensitivity and resolution, to strengthening identification and quantification, and now to integrating with spatial transcriptomics and proteomics to build mechanistic networks.
The next phase will be defined by reliable identification, more quantitative reporting, scalable computation, tighter multi-omics integration, and standardization that enables translation. In that future, “metabolic maps” may become as routine and informative as histology—adding molecular function to morphology and helping us see biology in its native, spatially organized reality.
References:
1. Nguyen K, Carleton G, Lum JJ, Duncan KD. Expanding Spatial Metabolomics Coverage with Lithium-Doped Nanospray Desorption Electrospray Ionization Mass Spectrometry Imaging. Anal Chem. 2024 Nov 19;96(46):18427-18436. doi: 10.1021/acs.analchem.4c03553.
2. Sun Y, Zhou Z, Xie B, Wang Z, Abliz Z. Spatially Resolved Metabolomics Method for Mapping the Global Molecular Landscape of Whole-Body Zebrafish (Danio rerio) Using Ambient Mass Spectrometry Imaging. Anal Chem. 2023 Jun 20;95(24):9164-9172. doi: 10.1021/acs.analchem.2c05047.
3. Planque M, Igelmann S, Ferreira Campos AM, Fendt SM. Spatial metabolomics principles and application to cancer research. Curr Opin Chem Biol. 2023 Oct;76:102362. doi: 10.1016/j.cbpa.2023.102362.
4. Wheeler K, Gosmanov C, Sandoval MJ, Yang Z, McCall LI. Frontiers in Mass Spectrometry-Based Spatial Metabolomics: Current Applications and Challenges in the Context of Biomedical Research. Trends Analyt Chem. 2024 Jun;175:117713. doi: 10.1016/j.trac.2024.117713.
5. Min X, Zhao Y, Yu M, Zhang W, Jiang X, Guo K, Wang X, Huang J, Li T, Sun L, He J. Spatially resolved metabolomics: From metabolite mapping to function visualising. Clin Transl Med. 2024 Nov;14(11):e70031. doi: 10.1002/ctm2.70031.
6. Guo X, Wang X, Tian C, Dai J, Zhao Z, Duan Y. Development of mass spectrometry imaging techniques and its latest applications. Talanta. 2023 Nov 1;264:124721. doi: 10.1016/j.talanta.2023.124721.
7. Godfrey TM, Shanneik Y, Zhang W, Tran T, Verbeeck N, Patterson NH, Jackobs FE, Nagi C, Ramineni M, Eberlin LS. Integrating Ambient Ionization Mass Spectrometry Imaging and Spatial Transcriptomics on the Same Cancer Tissues to Identify RNA-Metabolite Correlations. Angew Chem Int Ed Engl. 2025 Jun 10;64(24):e202502028. doi: 10.1002/anie.202502028.
8. Mavroudakis L, Golubova A, Lanekoff I. Spatial metabolomics platform combining mass spectrometry imaging and in-depth chemical characterization with capillary electrophoresis. Talanta. 2025 May 1;286:127460. doi: 10.1016/j.talanta.2024.127460.
9. Morosi L, Miotto M, Timo S, Carloni S, Bruno E, Meroni M, Menna E, Lodato S, Rescigno M, Martano G. MSIpixel: a fully automated pipeline for compound annotation and quantitation in mass spectrometry imaging experiments. Brief Bioinform. 2023 Nov 22;25(1):bbad463. doi: 10.1093/bib/bbad463.
10. Zhu L, Diao X, Yuan C, Lau CML, Wang J, Wu W, Mobasheri A, Houard X, Wen C, Cai Z. Integrative spatially resolved proteomic and metabolomic imaging reveals synovitis endotypes implicated in osteoarthritis progression. Theranostics. 2025 Aug 30;15(18):9729-9741. doi: 10.7150/thno.117788.
Next-Generation Omics Solutions:
Proteomics & Metabolomics
Ready to get started? Submit your inquiry or contact us at support-global@metwarebio.com.


