Core Instruments and Platforms for Spatial Metabolomics: MALDI-MSI, DESI-MSI, and SIMS
Spatial metabolomics is not just metabolomics performed on tissue—it is metabolomics re-anchored to anatomy. The mission is simple to state but hard to execute: put metabolite identities and abundances (what + how much) back into the tissue coordinate system (where). Imaging mass spectrometry (MSI) is the most direct route to that goal because it measures molecules in situ, pixel by pixel, producing ion maps that can be aligned with histology and pathology.
The catch is that spatial metabolomics is defined by trade-offs. Every platform sits somewhere on a four-way tension between:
- Spatial resolution (how small a pixel can be)
- Coverage / sensitivity (how many metabolites you can detect, and how low you can go)
- Quantitative accuracy (how comparable intensities are across pixels, samples, and batches)
- Throughput / cost (how fast and scalable the workflow is)
There is no single instrument that wins on all four dimensions. That’s why platform choice is not a “best instrument” question—it’s a best match for the scientific question question. Reviews focusing on high-spatial-resolution MSI for spatial metabolomics highlight exactly this reality: pushing resolution upward often increases experimental complexity and can reduce coverage and/or sensitivity.
In this post, we break down the three “workhorse” in situ imaging technologies—MALDI-MSI, DESI-MSI, and SIMS—from ionization physics to hardware, strengths, limitations, and selection logic. The goal is to give you a practical mental map for choosing a platform for tumor margin mapping, neurotransmitter/lipid regionalization, drug distribution, isotope tracing, and more.
1. MALDI-MSI: Versatile, Coverage-First Spatial Metabolomics
Matrix-assisted laser desorption/ionization (MALDI) MSI coats the tissue with a matrix (organic crystals that absorb laser energy). A laser raster scans the sample; at each pixel, the matrix helps desorb and ionize analytes into the gas phase for MS analysis. Because MALDI is a “soft” ionization approach (relative to SIMS), it can preserve intact molecular ions for many lipid classes, metabolites, and peptides, supporting higher-confidence annotations when paired with high mass resolution and MS/MS strategies. Reviews of MALDI-MSI advances in tissues emphasize that MALDI remains the most widely used MSI method for biological samples due to its overall versatility and maturity.
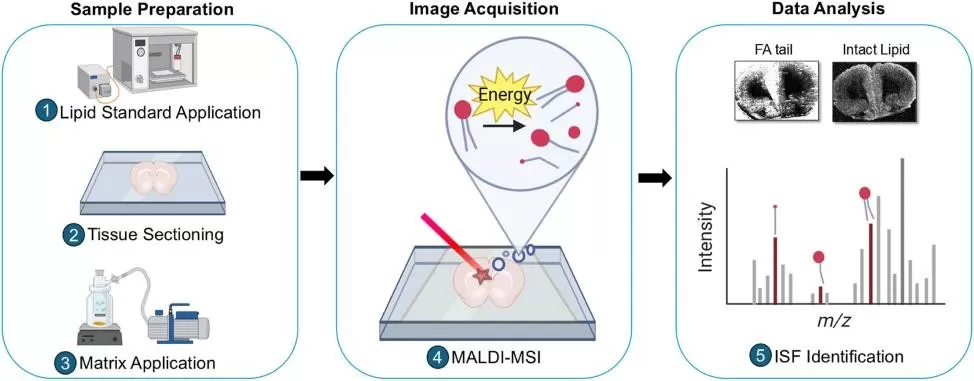
MALDI-MSI Workflow Overview.
Image reproduced from Cousineau, S. L., Sarikahya, M. H., Jurcic, K. et al., 2025, Analytica chimica acta, licensed under the Creative Commons Attribution License (CC BY 4.0).
Core instrument configurations
MALDI imaging is a workflow composed of four tightly coupled modules:
1. Ion source + laser optics (spot size and repetition rate heavily influence resolution and speed)
2. Motorized stage / raster scanning control
3. Mass analyzer (determines resolving power, accuracy, speed, and MS/MS capability)
4. Imaging software + file formats (imzML export, visualization, ROI tools, statistics)
Common analyzer pairings include:
- MALDI-TOF: often favored for high-speed imaging (large cohorts, large tissue areas)
- MALDI-Orbitrap / FTICR: favored when mass accuracy and resolving power are needed (isobars, complex lipid backgrounds, confident formula-level calls). Reviews discussing MALDI MSI for small molecules highlight how higher resolution and accuracy help address biological complexity and reduce false assignments.
Examples of current platforms:
Commercial ecosystems exist across multiple vendors; for example, Bruker describes the timsTOF fleX as combining MALDI imaging with an integrated workflow and analysis tools. Thermo Fisher also describes high-resolution imaging workflows built around Orbitrap systems coupled to atmospheric-pressure MALDI-style sources (e.g., AP-SMALDI integrations) for high mass accuracy imaging.
What MALDI-MSI is best at
- Broad molecular coverage: lipids, many metabolites, peptides, and drug-related molecules (matrix and method dependent)
- Strong software ecosystem and community protocols: critical for spatial workflows that require reproducibility, alignment, and scalable analysis
- Balanced resolution: often in the 10–50 µm range for many workflows, with specialized approaches pushing lower (but typically with trade-offs)
Where MALDI-MSI struggles
- Sample preparation dominates image quality: matrix selection, crystal size, and coating uniformity strongly affect spatial fidelity and ion yield. MALDI reviews repeatedly emphasize sample prep as a primary determinant of performance.
- Ion suppression and tissue heterogeneity: intensity is not purely concentration; it is “concentration × ionization efficiency”, which varies by microenvironment.
- Quantitation is non-trivial: it’s possible, but it requires deliberate design (internal standards, normalization, and often tissue-matched calibration strategies).
When MALDI-MSI is the best choice
- Tumor heterogeneity and margin biology where you want metabolites + lipids together
- Brain region mapping (lipids, energy metabolites, small molecules—depending on method)
- Pathology-driven ROI analysis where mature co-registration workflows matter
2. DESI-MSI: Ambient, Minimal Prep, Clinically Friendly
Desorption electrospray ionization (DESI) MSI is an ambient ionization technique. A charged solvent spray impacts the tissue surface, dissolves analytes in a thin film, and ejects secondary droplets carrying ions into the MS inlet. Because it runs at atmospheric pressure and usually requires no matrix, DESI is often positioned as a workflow-friendly option for clinical and translational settings. Waters’ DESI documentation also highlights this “direct-from-surface” nature and compatibility with histopathology workflows.
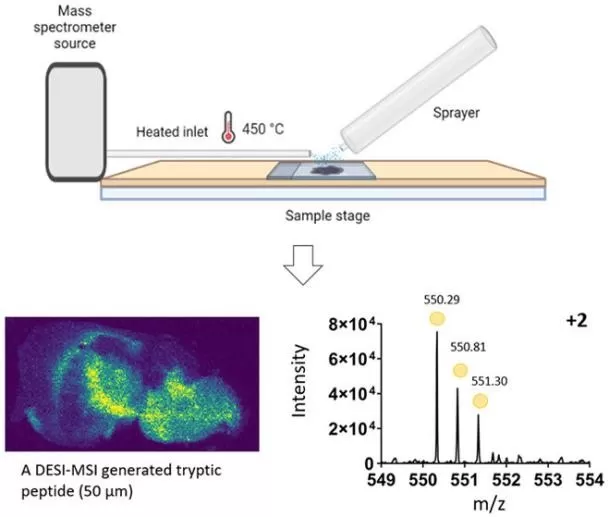
DESI-MSI Workflow Overview.
Image reproduced from Bottomley, H., Phillips, J., Hart, P., 2024, Journal of the American Society for Mass Spectrometry, licensed under the Creative Commons Attribution License (CC BY 4.0).
Core instrument configurations
- DESI source: spray geometry, solvent composition, flow rate, nebulizing gas, and stage height are key
- High-resolution MS is common (Q-TOF or Orbitrap-class analyzers) because complex tissues benefit from high mass accuracy for annotation and cleaner images.
Examples of current platforms:
Waters positions DESI XS as a DESI imaging source designed for stability and high-throughput MSI and notes typical pixel sizes and workflow integration features.
What DESI-MSI is best at
- Speed and simplicity: no matrix coating step reduces hands-on time and variability.
- Ambient operation: practical for clinical samples and rapid-turnaround studies.
- Drug and metabolite localization: DESI is frequently used for exogenous compounds and metabolic phenotyping where minimal manipulation is desired (method dependent).
Where DESI-MSI struggles
- Spatial resolution ceiling: DESI resolution is often limited by spray footprint and solvent spreading. Vendors and protocols have improved this substantially, but it remains a key differentiator versus MALDI/SIMS.
- Matrix effects / ionization variability: tissue microenvironments can change extraction and ion formation, complicating quantitation and cross-sample comparisons.
- Method optimization burden: solvent system tuning (polarity, additives), spray angle, and stage conditions are often sample- and target-dependent.
When DESI-MSI is the best choice
- High-throughput screening across many tissues
- Clinical workflows where simplicity and turnaround matter
- Rapid drug distribution checks and metabolite co-localization studies
3. SIMS: Subcellular Spatial Metabolomics and Isotope Imaging
Secondary ion mass spectrometry (SIMS) bombards the sample surface with a focused primary ion beam, sputtering secondary ions from the topmost layers. SIMS can reach extremely high spatial resolution and provides strong elemental and isotope sensitivity, but it often fragments larger biomolecules, complicating intact metabolite identification.
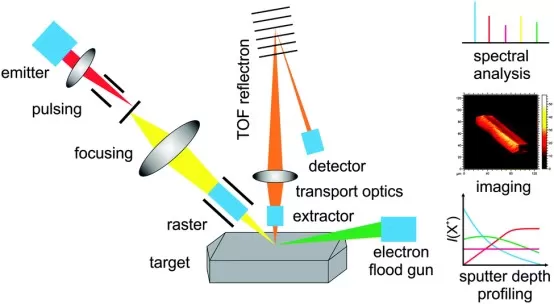
SIMS Workflow Overview.
Image reproduced from Hofmann, J. P., Rohnke, M., & Weckhuysen, B. M., 2014, Physical chemistry chemical physics, licensed under the Creative Commons Attribution License (CC BY 3.0).
Core instrument configurations
- Ion gun / primary beam: the main determinant of spatial resolution and sputter behavior
- Mass analyzer: ToF analyzers are common; NanoSIMS focuses on isotope imaging with high precision.
Examples of current platforms:
CAMECA describes NanoSIMS instruments as enabling ultra-high-resolution chemical imaging, with lateral resolution on the order of ~100 nm in some configurations.
What SIMS is best at
- Extreme spatial resolution: pushing from sub-micron down toward nanoscale, enabling subcellular mapping in the right context.
- Surface sensitivity: information-rich top-layer chemistry (but note: that also means sample prep and surface contamination matter a lot).
- Isotope tracing: NanoSIMS is widely used for isotope ratio imaging and metabolic tracing experiments where isotopic enrichment is the readout.
Where SIMS struggles
- Fragmentation is a feature and a bug: great for elements and certain fragments; challenging for intact small-molecule IDs.
- Limited molecular breadth relative to MALDI for intact biomolecules (application- and instrument-dependent).
- Vacuum and prep complexity: sample handling and compatibility can be more demanding.
When SIMS is the best choice
- Subcellular localization questions where resolution is the primary constraint
- Isotope-label tracing (e.g., 13C, 15N) mapped into microanatomy
- Elemental or small-fragment chemical imaging needs
4. How to Choose: A Practical Decision Framework
A useful way to choose is to start from the biological question and map it to the dominant constraint:
- If you care most about “seeing more molecules”: Choose MALDI-MSI. It is the most established compromise of coverage, resolution, and community tooling, with strong literature support for broad spatial omics use cases.
- If you care most about “moving fast”: Choose DESI-MSI. Ambient operation and reduced prep make it attractive for high-throughput or clinical translation paths.
- If you care most about “seeing smaller structures”: Choose SIMS. Subcellular resolution and isotope imaging capabilities make it uniquely powerful—if you accept limited intact-molecule coverage and higher operational complexity.
A high-spatial-resolution MSI review focused on spatial metabolomics makes the broader point: cellular and subcellular imaging is achievable, but the practical workflow is always a negotiation among resolution, sensitivity, and identification confidence.
Comparison of Key Features in MALDI-MSI, DESI-MSI, and SIMS for Spatial Metabolomics
|
Dimension |
MALDI-MSI |
DESI-MSI |
SIMS |
|
Probe Type |
Laser (requires matrix) |
Charged spray (no matrix required) |
Ion beam |
|
Spatial Resolution |
5-50 μm |
50-200 μm |
50 nm - 1 μm |
|
Molecular Coverage |
★★★★★ |
★★★★☆ (Moderate) |
★★☆☆☆ (Narrow, small molecules/elements) |
|
Molecular Identification |
★★★★☆ (MS/MS possible) |
★★★★☆ (MS/MS possible) |
★☆☆☆☆ (Heavy fragmentation, difficult identification) |
|
Sample Preparation Complexity |
Medium (requires matrix spraying) |
Low (minimal preparation required) |
High (requires ultra-high vacuum compatibility) |
|
Imaging Speed |
Fast (high-frequency laser) |
Moderate (spray scanning) |
Slow (point-by-point bombardment) |
|
Main Applications |
Lipids/peptides/drugs imaging |
Clinical screening/drug distribution |
Subcellular localization/isotope tracing |
5. Beyond the Instrument: Sample Prep, Acquisition, Software, and IDs
Sample prep and automation: where images are won or lost
For MALDI especially, sample prep is not a footnote—it’s the main event. Tissue collection, freezing, sectioning thickness, humidity control, and matrix coating uniformity can dominate reproducibility and spatial fidelity. Reviews of MALDI MSI for small molecules explicitly point to sample handling and matrix coating as major determinants of data quality.
Practical implications:
- Inconsistent coating → streaking, “hot spots”, missing regions, biased intensities
- Warm handling → metabolite delocalization risk
- Contamination control → cleaner low-mass spectra and fewer false features
DESI and SIMS have different prep pain points (solvent spreading and surface cleanliness for DESI; vacuum compatibility and surface integrity for SIMS), but the meta-rule is consistent: your chemistry is only as good as your surface.
Data acquisition and software ecosystem: from pixels to biology
Spatial metabolomics data is not just a list of peaks—it’s a 3D object (x, y, m/z), sometimes with MS/MS layers. Mature workflows typically include:
- Ion image reconstruction and normalization
- Peak picking and feature grouping across pixels
- Spatial statistics (segmentation, clustering, spatial autocorrelation)
- ROI-based comparisons (tumor vs stroma, cortex layers, etc.)
- Batch correction and QC metrics across runs
Vendors often provide streamlined imaging pipelines (e.g., queue-based acquisition and processing in DESI workflows), but many groups rely on standardized data formats like imzML plus downstream open-source or academic tools. The key is to plan analysis before data acquisition: pixel size, scan strategy, MS/MS triggers, and QC samples should all serve the analysis endpoints.
Identification and annotation: the Achilles’ heel of spatial metabolomics
The most common critique of spatial metabolomics is: “How sure are you that peak is what you say it is”? This is why robust workflows emphasize layered evidence:
- Putative: accurate mass + isotope pattern + plausible adducts
- More confident: MS/MS spectral match to a library or in-house standards
- Confirmed: on-tissue or tissue-matched standard confirmation under matched conditions
High mass resolving power helps reduce ambiguity for isobaric species, but it does not fully solve isomeric complexity—especially for lipids and many small molecules—so annotation strategy must be deliberate. Reviews on high-resolution spatial metabolomics discuss these identification challenges as a central limitation and a key area of method development.
Quantification and validation: turning “images” into “comparable measurements”
Pixel intensities are not automatically quantitative. They depend on extraction/ionization efficiency, local salt content, matrix crystallization (MALDI), spray stability (DESI), and surface effects (SIMS). That said, spatial metabolomics quantification is advancing quickly, especially with internal standard strategies.
A recent Nature Metabolism study reports a quantitative MSI workflow using isotopically labeled yeast extract as internal standards to improve quantification across tissues and enable quantification of hundreds of metabolic features.
Practical quantification concepts to know:
- Relative quantification: compare within-sample patterns (good for localization, weaker for cross-sample claims)
- Normalization strategies: TIC, median, internal standards, or region-aware approaches (each has assumptions)
- Absolute/semi-quant: requires internal standards and careful calibration logic (often tissue-matched)
And for high-stakes claims (biomarkers, mechanism, drug exposure), a common best practice is to validate key findings with orthogonal methods (e.g., LC-MS/MS targeted quantification). That gives the image a quantitative backbone.
6. Multimodal Spatial Platforms: The “Combination Punch” Strategy
Modern spatial biology is trending toward multimodal integration—because molecules alone rarely answer “why here”?
Typical combinations include:
- MSI + histology (H&E / IHC): align chemical features with morphology and cell types
- MSI + spatial transcriptomics / spatial proteomics: connect metabolite patterns to pathway activity and microenvironmental states
- Near-single-cell strategies: rather than promising true single-cell metabolomics everywhere, many practical workflows aim for “cellular neighborhood” resolution and interpret results with multimodal context
SIMS and MALDI are also increasingly used in correlative imaging frameworks (SIMS for isotopes and nanoscale structure; MALDI for broader molecular coverage), consistent with the SIMS tutorial review’s discussion of correlative approaches in biology.
References:
1. Zhu, X., Xu, T., Peng, C., & Wu, S. (2022). Advances in MALDI Mass Spectrometry Imaging Single Cell and Tissues. Frontiers in chemistry, 9, 782432. https://doi.org/10.3389/fchem.2021.782432
2. Unlocking spatial metabolomics with isotopically labelled internal standards. (2025). Nature metabolism, 7(9), 1730–1731. https://doi.org/10.1038/s42255-025-01341-7
3. Cousineau, S. L., Sarikahya, M. H., Jurcic, K., Laviolette, S. R., Hardy, D. B., & Yeung, K. K. (2025). Monitoring the degree of in-source phospholipid fragmentation during MALDI mass spectrometry imaging. Analytica chimica acta, 1367, 344297. https://doi.org/10.1016/j.aca.2025.344297
4. Bottomley, H., Phillips, J., & Hart, P. (2024). Improved Detection of Tryptic Peptides from Tissue Sections Using Desorption Electrospray Ionization Mass Spectrometry Imaging. Journal of the American Society for Mass Spectrometry, 35(5), 922–934. https://doi.org/10.1021/jasms.4c00006
5. Hofmann, J. P., Rohnke, M., & Weckhuysen, B. M. (2014). Recent advances in secondary ion mass spectrometry of solid acid catalysts: large zeolite crystals under bombardment. Physical chemistry chemical physics : PCCP, 16(12), 5465–5474. https://doi.org/10.1039/c3cp54337d
Next-Generation Omics Solutions:
Proteomics & Metabolomics
Ready to get started? Submit your inquiry or contact us at support-global@metwarebio.com.


