Spatial Metabolomics Sample Preparation: Cryosectioning vs. Pressing vs. Imprinting for MSI
Spatial metabolomics has opened a revolutionary window into biology, allowing scientists to map the molecular landscape of tissues directly within their native architectural context. The power of techniques like MALDI-MSI and DESI-MSI to visualize metabolite distributions is undeniable. However, the fidelity of these stunning molecular images is fundamentally determined at the very first step: sample preparation. The critical challenge lies in preserving two equally important aspects—the precise spatial integrity of the tissue and the chemical integrity of the metabolome. This guide provides a detailed, comparative exploration of the three cornerstone sample preparation methods in spatial metabolomics: cryo-sectioning, tissue pressing, and imprinting. By understanding the principles, strengths, and optimal applications of each, you can make an informed decision that ensures the success and reliability of your imaging experiment.
1. Core Sample Preparation Methods for Spatial Metabolomics
The journey to a successful spatial metabolomics image begins with transforming a complex, three-dimensional tissue into a format amenable to mass spectrometry imaging. Each preparation method follows a distinct physical or chemical principle to achieve this goal, catering to different sample types and research questions. Mastering these foundational techniques is paramount for obtaining data that accurately reflects the in situ metabolic state of your sample.
1.1. Cryosectioning for Spatial Metabolomics: The High-Resolution Benchmark
Cryo-sectioning is the most widely established and versatile method, often considered the gold standard for high-resolution studies. Its core principle relies on rapid physical stabilization through freezing, followed by precise mechanical slicing.
Workflow: Fresh tissue samples are promptly snap-frozen, typically using liquid nitrogen or isopentane cooled by liquid nitrogen, to vitrify water and halt enzymatic degradation instantaneously. The frozen tissue is then mounted on a cryostat and sectioned into thin slices (usually 5-20 µm thick) at temperatures between -15°C to -25°C. These thin sections are carefully thaw-mounted onto conductive glass slides or specialized targets compatible with MALDI-MSI. For DESI-MSI, mounting onto standard glass slides is often sufficient.
Key Advantages: This method excels in preserving fine histological details, enabling spatial resolutions down to cellular dimensions (~5-10 µm with modern systems). It maintains excellent metabolite localization and is compatible with a vast array of tissues, from animal organs (brain, liver, kidney) to dense plant structures (roots, seeds, fruits). A major strategic benefit is its compatibility with consecutive sectioning, allowing adjacent slices to be used for correlative histology (H&E, IHC) or other omics analyses, enabling powerful multimodal integration.
Considerations: The process requires access to a cryostat and technical expertise to avoid artifacts like folding, cracking, or thawing. Optimal cutting temperature must be empirically determined for different tissues to balance brittleness and plasticity.
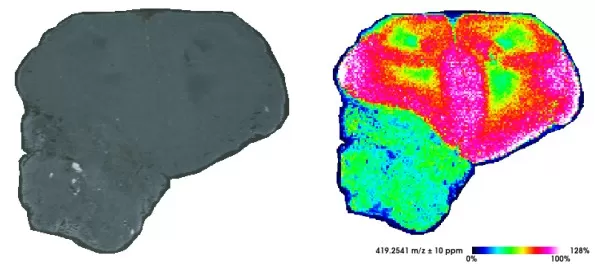
Figure 1. MALDI-MSI imaging of mouse brain tissue using cryo-sectioning.
1.2. Tissue Pressing in Spatial Metabolomics: Simplicity for Delicate Structures
Tissue pressing offers an elegantly simple, equipment-light alternative for samples that are challenging to section using a cryostat. It is a mechanical flattening process that avoids the need for freezing and complex microtomy.
Workflow: A fresh or briefly frozen sample is placed between two flat, clean surfaces—often glass slides or metal plates. Controlled physical pressure is applied manually or via a simple press, flattening the tissue into a thin, extended layer. The pressure and duration are adjusted based on the sample's texture and water content. The resulting "pressed mount" can be analyzed directly or after gentle drying.
Key Advantages: Its foremost advantage is simplicity and minimal sample handling, which is crucial for fragile, leafy, or petal-like structures (e.g., plant leaves, flower petals, thin animal membranes) that would disintegrate during cryo-sectioning. It requires no specialized freezing or slicing equipment, reducing cost and preparation time. It is well-suited for DESI-MSI and can be adapted for MALDI-MSI with careful application of matrix.
Considerations: Spatial resolution is inherently lower and less uniform than sectioning, as cellular architecture is compressed. It provides a composite view through the thickness of the pressed layer rather than a defined single-plane section. Quantitative interpretation can be complicated by this compression and potential uneven metabolite spreading.

Figure 2. MALDI-MSI imaging of walnut petals using tissue pressing.
1.3. Imprinting (Blotting) for Spatial Metabolomics: Capturing the Surface Metabolome
Imprinting, or blotting, is a unique, indirect sampling technique designed to selectively analyze metabolites present on a tissue's surface or those that readily transfer from it.
Workflow: A fresh, minimally processed tissue sample is brought into firm, uniform contact with a porous, chemically inert substrate. Common substrates include polytetrafluoroethylene (PTFE) membranes or specialized blotting papers. Through a combination of adsorption and mild pressure, a fraction of the metabolites from the tissue surface is transferred onto the substrate. The tissue is then removed, and the substrate bearing the "metabolite imprint" is subjected to MSI analysis.

Figure 3. Principle of the imprinting method (Wu L., 2022)
Key Advantages: This method effectively decouples metabolites from the complex biological matrix of the tissue, which can reduce ion suppression effects during MS analysis, particularly benefiting DESI-MSI. It is excellent for focused studies on surface lipids, waxes, excreted compounds, or metabolites in highly hydrated tissues. The imprints can be stable for some time, allowing for batch analysis or archiving.
Considerations: It provides no direct information on the tissue's internal structure or depth-resolved metabolite gradients. Transfer efficiency can vary for different metabolite classes based on their polarity and adhesion properties, making absolute quantification challenging. The method is primarily suitable for flat, moist tissue surfaces.

Figure 4. DESI-MSI imaging of tea leaves using imprinting (Wu L., 2022)
2. A Comparative Framework: Selecting the Optimal Method for Spatial Metabolomics
With a clear understanding of each method's individual profile, a side-by-side comparison reveals their complementary roles within the spatial metabolomics workflow. The choice is rarely about finding the "best" method in absolute terms, but rather about identifying the "most appropriate" one based on a matrix of experimental requirements and sample constraints. The following analysis and synthesis are designed to guide this critical decision.
2.1. Technical Comparison: Cryosectioning vs. Pressing vs. Imprinting for MSI
The table below summarizes the critical performance metrics of the three methods, providing a clear, at-a-glance comparison based on key parameters crucial for experimental design in MALDI-MSI and DESI-MSI.
Table 1. Comparative Analysis of Technical Performance in Sample Preparation Methods
|
Parameter |
Cryo-Sectioning |
Tissue Pressing |
Imprinting |
|
Spatial Resolution |
High (5-40 µm). Enables cellular/subcellular mapping. |
Low to Moderate. Resolution is limited by tissue compression and spreading. |
Low. Reflects transfer pattern, not original cellular architecture. |
|
Metabolite Coverage |
Broad and Representative. Captures metabolites throughout the section thickness. |
Broad but Compressed. Captures metabolites from the entire pressed volume. |
Selective (Surface-biased). Primarily captures readily transferable surface metabolites. |
|
Sample Integrity |
Excellent structural preservation. Maintains original tissue morphology. |
Altered structure. Flattens and compresses 3D architecture. |
No tissue structure analyzed. Only the metabolite transfer is imaged. |
|
Technical Complexity |
High. Requires cryostat, expertise in freezing/sectioning. |
Very Low. Minimal equipment and training needed. |
Low. Simple procedure but requires optimization of contact. |
|
Multi-omics Correlation |
Excellent. Enables direct, adjacent-section correlation with histology/transcriptomics. |
Poor. No corresponding morphology for precise alignment. |
Not applicable. No tissue structure remains for correlation. |
|
Typical Sample Types |
Animal organs, tumors, plant roots/seeds/fruits, biofilms. |
Delicate plant leaves & petals, soft animal tissues, thin films. |
Plant leaf surfaces, skin, microbial mats, moist tissue surfaces. |
2.2. How to Choose: A Decision Guide for Spatial Metabolomics Sample Preparation
The final choice should be driven by a hierarchy of considerations: 1) Sample Type & Physical Properties, 2) Primary Research Question, 3) Available Technology (MSI Platform).
1) For Animal Tissues, Clinical Specimens, and Dense Plant Organs: CHOOSE CRYO-SECTIONING. These samples often have complex, heterogeneous anatomy where correlating metabolism with specific histological regions (e.g., tumor core vs. stroma, brain nuclei) is the research goal. Cryo-sectioning is the only method that provides the necessary structural fidelity and high resolution. It is the unequivocal first choice for integrative pathology-spatial metabolomics studies.
2) For Fragile, Sheet-like Plant Samples (Leaves, Petals): CHOOSE TISSUE PRESSING or IMPRINTING.
- Choose Pressing if the goal is to obtain a general metabolic profile of the entire leaf/petal structure with simple preparation. It is robust and effective for DESI-MSI analysis of such samples.
- Choose Imprinting if the research question is specifically focused on surface metabolites (e.g., cuticular lipids, pheromones, environmental interactions). It minimizes interference from internal tissue matrix.
3) General Best Practices:
- Pilot Studies: When working with a novel sample type, running a small pilot comparing feasible methods can be highly informative.
- Quality Control: Always include a method to assess preparation quality. For sectioning, this is H&E staining of a consecutive section. For pressing/imprinting, microscopic examination of the tissue before/after or the imprint itself can reveal issues.
- MSI Platform Synergy: Remember that DESI-MSI is generally more tolerant of sample topography and humidity than MALDI-MSI. Pressed and imprinted samples often perform better with DESI.
Table 2. Strategic Selection Guide for Common Sample Types
|
Sample Type |
Recommended Method |
Compatible MSI Technology |
Rationale for Selection |
|
Animal Organs / Clinical Tumors / Plant Roots & Fruits |
Cryo-Sectioning |
MALDI-MSI, DESI-MSI |
These samples have complex, dense structures. Sectioning provides uniform thin slices that best preserve spatial architecture and internal metabolite distribution for high-resolution mapping and correlation with histopathology. |
|
Plant Leaves / Petals / Delicate Herbal Specimens |
Tissue Pressing |
DESI-MSI (partially MALDI-MSI) |
These samples are large, soft, and prone to shredding during cryo-sectioning. Pressing quickly creates a coherent, flattened layer suitable for analysis with minimal damage, making it ideal for DESI-MSI workflows. |
|
Fresh Plant Leaves / Petals (Surface Metabolite Focus) |
Imprinting |
DESI-MSI |
For studies targeting surface metabolites (e.g., cuticular lipids), imprinting effectively enriches and stabilizes these compounds from moist surfaces, separating them from the bulk tissue matrix—a perfect match for DESI-MSI analysis. |
3. Future Perspectives: Standardizing Spatial and Chemical Integrity in MSI
In the dynamic field of spatial metabolomics, sample preparation is the critical bridge between the biological specimen and the insightful molecular image. Cryo-sectioning, tissue pressing, and imprinting are not competing techniques but specialized tools within a comprehensive toolkit. The "correct" method is the one that most faithfully translates your specific biological question into an analytical format compatible with MALDI-MSI or DESI-MSI, while respecting the physical limits of your sample.
As technologies advance, these fundamental methods will also evolve, potentially through automation, improved standardization, or the development of novel embedding matrices. However, the core principles of preserving spatial and chemical integrity will remain paramount. By making a strategic, informed choice at this foundational stage, you ensure that the powerful downstream capabilities of mass spectrometry imaging are applied to data of the highest possible quality and biological relevance, paving the way for robust and groundbreaking discoveries.
Reference
Wu L, Qi K, Liu C, Hu Y, Xu M, Pan Y. Enhanced Coverage and Sensitivity of Imprint DESI Mass Spectrometry Imaging for Plant Leaf Metabolites by Post-photoionization. Anal Chem. 2022 Nov 1;94(43):15108-15116. doi: 10.1021/acs.analchem.2c03329. Epub 2022 Oct 6. PMID: 36201321.
Read more
-
Spatial Metabolomics: Transforming Biomedical and Agricultural Research
-
Spatial Metabolomics Explained: How It Works and Its Role in Cancer Research
-
MALDI, DESI, or SIMS? How to Choose the Best MSI Techniques for Spatial Metabolomics
-
How to Prepare Samples for Spatial Metabolomics: The Essential Guide You Need
-
Unlocking Precision in Spatial Metabolomics: Essential Detection Parameters for Cutting-Edge Research
Next-Generation Omics Solutions:
Proteomics & Metabolomics
Ready to get started? Submit your inquiry or contact us at support-global@metwarebio.com.


