Tissue Cryosectioning Guide for Spatial Metabolomics: Best Practices and Troubleshooting
The primary goal of spatial metabolomics is to preserve and measure the in situ distribution and relative abundance of metabolites within intact tissue architecture, making sample preparation a decisive factor for data quality. Section integrity directly influences downstream mass spectrometry imaging (MSI) performance, including detection sensitivity, achievable spatial resolution, and overall reproducibility. Because tissue types differ markedly in microstructure, water content, and biochemical composition, tissue cryosectioning parameters and handling precautions must be adapted accordingly to ensure reliable spatial metabolite maps.
Cryosectioning Principles for Spatial Metabolomics Sample Preparation
Across tissue types, cryosections for spatial metabolomics should follow standardized practices built around three essentials: preserved morphology, metabolite stability, and minimal exogenous contamination. First, samples should remain fresh, and the workflow from collection to sectioning must be completed rapidly to reduce metabolite degradation associated with prolonged handling. Second, strict temperature control is critical; maintaining a consistently low-temperature environment throughout processing helps minimize temperature-driven metabolic perturbation and preserves chemical integrity. Finally, sections should be cut to a uniform thickness and remain structurally intact—free of folds, cracks, or tears—so that the measured spatial distributions reflect true biology rather than preparation artifacts.

Figure 1: Examples of abnormal tissue outcomes
The table below summarizes common issues encountered during spatial metabolomics sectioning and practical solutions.
Table 1. Common Tissue Sectioning Issues in Spatial Metabolomics (MSI) and Recommended Solutions
|
Common issue |
Representative tissues |
Solution |
|
Ice-crystal voids in tissue |
Brain, muscle, intestine |
Blot surface moisture before embedding; section promptly after embedding; ensure storage at −80°C |
|
Calcification/necrosis leads to hard, brittle, fragmented tissue |
Atherosclerotic plaque, human tumor tissue, bone tissue |
Perform chemical decalcification (e.g., EDTA-based reagents); remove or avoid calcified regions during trimming |
|
Multilayer tissue with uneven hardness between layers |
Skin, eyeball |
Adjust the cutting (blade-entry) angle to reduce the effective contact area between tissue and blade |
|
Collagen fiber recoil causes wrinkled sections |
Cartilage |
Pre-soak using an appropriate ratio to improve tissue toughness |
|
Internal voids make intact sectioning difficult |
Lung tissue |
Use appropriate filling/infusion handling before embedding |
|
Large cavity structures within tissue |
Heart, aorta, eyeball, auricular concha |
Use appropriate filling/infusion handling before embedding |
|
Fat at tissue edges |
Testis, epididymis, kidney, some human tissues |
Remove fat before embedding; adjust block orientation to prevent fat from covering regions of interest |
|
High intrinsic fat content |
High-fat model liver, pancreas |
Use a more supportive embedding medium; increase section thickness |
|
Very small samples or high positional precision required |
Blood vessels, organoids, spheroids |
Use tissue staining or a contrasting embedding medium for localization; section immediately after embedding |
|
Tissue separates from embedding medium |
Skin, intestine |
Pre-soak tissue in embedding medium; add embedding medium stepwise to reduce bubbles |
|
Tissue deformation |
Intestine, brain |
Control embedded block size; add embedding medium in multiple steps |
Tissue-Specific Sectioning Guidance for Spatial Metabolomics
Although the core workflow of spatial metabolomics is broadly consistent, optimal outcomes depend heavily on tailoring tissue cryosectioning conditions to the structural and biochemical properties of each sample type. Differences in water content, lipid abundance, extracellular matrix density, and internal cavities can all reshape section integrity and introduce artifacts that compromise mass spectrometry imaging (MSI) readouts, including MALDI-MSI and DESI-MSI. The guidance below summarizes practical, tissue-adapted parameters—embedding media, cryostat temperature, blade control, and section thickness—to help generate reproducible, morphology-preserving sections that support reliable spatial metabolite mapping.
Animal parenchymal organs (liver, kidney, heart, spleen, etc.)
These organs are relatively dense and typically have moderate water content, so the main objective is to maintain both structural integrity and metabolite stability. After collection, gently blot surface fluids with sterile filter paper, embed the tissue in 2%–4% sodium carboxymethyl cellulose (CMC), and rapidly freeze the embedded block on dry ice until fully solidified. For longer-term storage, keep blocks at −80°C. Set the cryostat to −15 to −20°C, equilibrate the tissue block for approximately 30 minutes, and cut sections at an appropriate thickness of 12–20 μm. Use a cold-mounting approach to transfer sections onto conductive ITO slides, then return slides to −80°C immediately after mounting to preserve spatial metabolite profiles for MSI workflows such as MALDI-MSI or DESI-MSI.
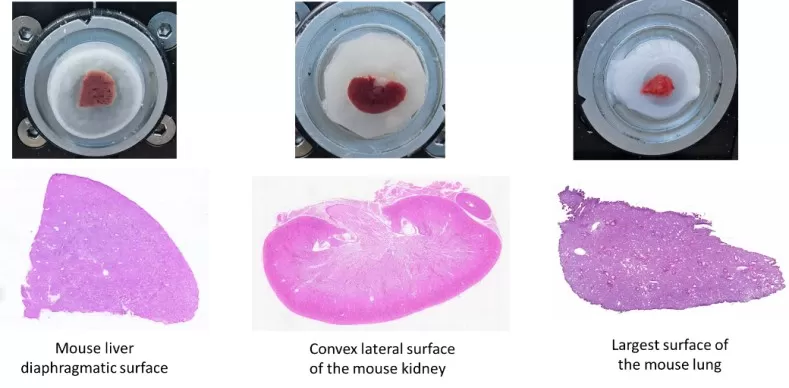
Figure 2: Mouse liver, kidney, and lung sections and H&E staining results
Neural and soft tissues (brain, spinal cord, muscle, adipose tissue)
Neural and soft tissues often have high water content and looser structure, making them more susceptible to deformation and metabolite loss during sectioning. For these tissues, embedding selection and freezing conditions are especially important. FSC22 is generally recommended as the embedding medium; for adipose-rich samples, 4% CMC can be used to improve mechanical support. Keep the blade temperature 5–10°C lower than the tissue block to reduce compression and smearing, and cut sections at 15–20 μm. Adipose tissue may require a low-temperature defatting pretreatment. After mounting, apply appropriate sealing to reduce oxidation and help stabilize lipids and other oxidation-sensitive metabolites commonly profiled in spatial metabolomics.
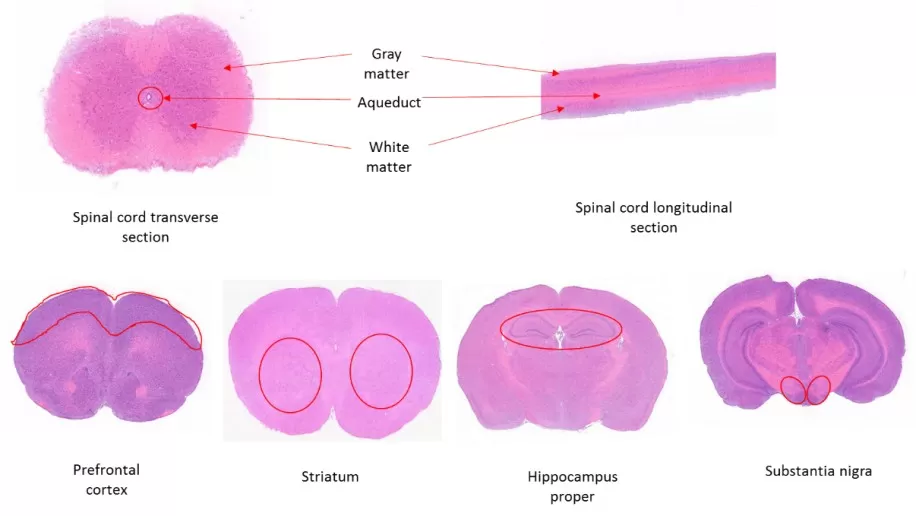
Figure 3: H&E staining of a typical mouse brain tissue plane
Clinical tumor tissues (tumor lesions and matched adjacent tissues)
Clinical tumor tissues are highly heterogeneous and often limited in quantity, so preparation should prioritize lesion preservation, strict contamination control, and efficient use of material. Ideally, snap-freeze tissues in liquid nitrogen within 30 minutes of excision; precisely mark the lesion area and embed with clear orientation to support downstream spatial interpretation. Handle tumor and adjacent tissues separately to avoid cross-contamination. Cut serial sections at 10–15 μm, and distribute them into sets for mass spectrometry imaging, H&E pathological validation, and reserve storage. This organization supports accurate spatial alignment between molecular ion images and histopathological morphology—an important requirement for robust tumor spatial metabolomics.
Tissues with special structures (blood vessels, cartilage, lung, skin, eyeball, etc.)
Tissues with complex structures—such as cavities, internal voids, strong layer-to-layer heterogeneity, or collagen-rich matrices—often require customized handling to improve section integrity. For cavity or void-containing tissues, fill the space with embedding medium to enhance mechanical support. For multilayer tissues, embed with deliberate orientation so that key interfaces are perpendicular to the cutting direction, reducing tearing at boundaries. For cartilage, minimize ex vivo time and use cold-mount pressing to reduce wrinkling driven by collagen recoil. For very small tissues (e.g., blood vessels or organoids), miniature molds can improve positional accuracy and fixation. For calcified or necrotic regions that are hard and brittle, reduce cutting temperature, increase section thickness, and trim away highly necrotic areas when possible to improve overall section quality for MSI-based spatial metabolomics.
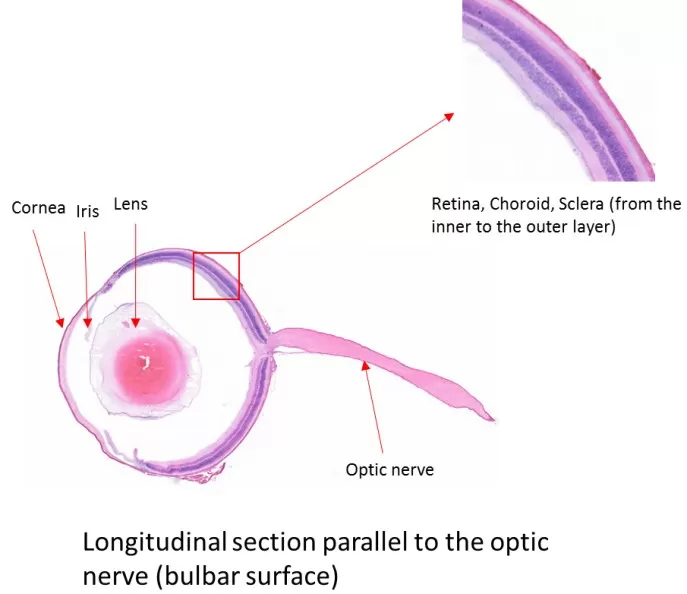
Figure 4: H&E staining of an eyeball tissue plane
Plant tissues (leaves, roots, stems, fruits, etc.)
Plant tissues contain cellulose-rich cell walls and often exhibit substantial structural heterogeneity, which increases the risk of cracking and fragmentation during cryosectioning. The primary goals are to maintain section integrity and minimize pigment-related interference. After cleaning and blotting, snap-freeze in liquid nitrogen and embed in 2% CMC. For harder tissues, a controlled pre-freeze can improve cutting behavior, and equilibration may be extended to ~40 minutes. Adjust section thickness according to tissue texture (8–30 μm), set the cryostat to −20 to −25°C, and use a sharp blade to reduce mechanical stress. A brief low-temperature dehydration step can help reduce pigment interference, supporting cleaner ion images for plant spatial metabolomics analyses.
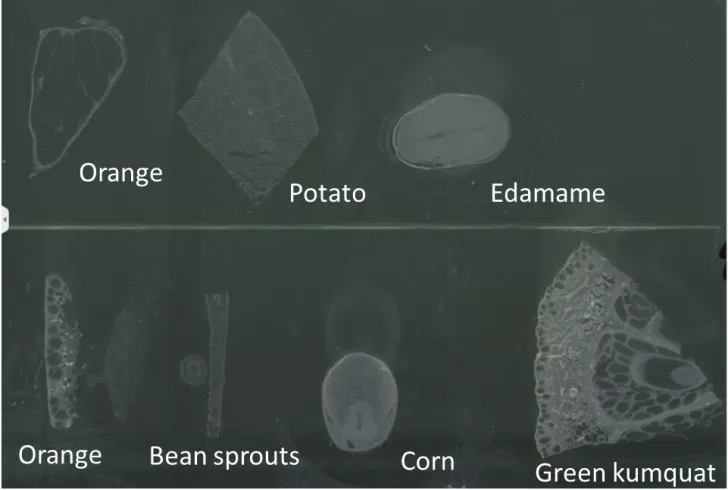

Figure 5: Representative Cryosections of Diverse Plant Tissues for Spatial Metabolomics and Mass Spectrometry Imaging (MSI)
Post-Section Handling and Quality Control for Spatial Metabolomics (MSI)
For sections that are not analyzed immediately, store slides at −80°C for long-term preservation. Minimize each room-temperature exposure to ≤5 minutes, and avoid repeated freeze–thaw cycles to reduce chemical degradation and preserve spatial fidelity. Morphology quality control should be performed by optical microscopy, ideally alongside stained images, to confirm that sections are free of damage, contamination, or folds and that tissue boundaries remain clear. This QC step ensures that sections meet the practical requirements for reliable mass spectrometry imaging and interpretable spatial metabolomics results.
Overall, tissue cryosection quality is a primary determinant of spatial resolution, signal robustness, and biological interpretability in spatial metabolomics and related MSI workflows (including MALDI-MSI and DESI-MSI). Because tissues differ in structure, composition, and physical properties, tissue-specific sectioning challenges are common. Optimizing key parameters—particularly embedding medium, freezing conditions, and section thickness—based on tissue characteristics is therefore fundamental to stable sectioning performance and successful downstream experiments.
References
1. Sheerin K, Malin A, Jennifer L H, Todd A G, Richard M C, et al. Direct Molecular Analysis Of Whole-Body Animal Tissue Sections By Imaging Maldi Mass Spectrometry[J], Analytical Chemistry, 2006, 78(18): 6448-6456.
2. Erin H S, Stacey R O, Deming M, Pierre C, Richard M C, et al. Enhancement of Protein Sensitivity for MALDI Imaging Mass Spectrometry after Chemical Treatment of Tissue Sections[J], Journal of the American Society for Mass Spectrometry, 2008, 19(8): 1069-1077.
Next-Generation Omics Solutions:
Proteomics & Metabolomics
Ready to get started? Submit your inquiry or contact us at support-global@metwarebio.com.


