A Complete Guide to Spearman Rank Correlation in Multi-Omics Research
In multi-omics data analysis, researchers frequently need to quantify associations between molecular features—whether examining gene-gene co-expression, protein-metabolite interactions, or cross-omics regulatory relationships. The Pearson correlation coefficient has long been the default choice for such analyses, yet it carries strict assumptions: data must follow a normal distribution and relationships must be linear. However, high-throughput biological data rarely satisfies these conditions. Due to biological sample heterogeneity, technical artifacts, and the presence of extreme expression values, omics data typically exhibit non-normal distributions and non-linear relationships. This article provides a comprehensive guide to Spearman‘s rank correlation—a robust non-parametric alternative that addresses these limitations. We will explore why Spearman’s method has become indispensable for modern multi-omics research and how you can leverage it to uncover biologically meaningful associations in your data.
What is Spearman Correlation? Basic Concepts and Principles
First introduced by psychologist Charles Spearman in 1904, Spearman‘s rank correlation coefficient (denoted as ρ or rₛ) is a non-parametric measure of statistical dependence between two variables. Unlike Pearson correlation, which evaluates linear relationships based on raw data values, Spearman’s method assesses monotonic relationships—whether the variables tend to increase or decrease together, regardless of whether the relationship follows a straight line.
The core insight underlying Spearman correlation is elegantly simple: instead of working with the original measurements, we convert the data to ranks. For each variable, the smallest value receives rank 1, the next smallest rank 2, and so on. Spearman‘s correlation is then calculated as the Pearson correlation applied to these ranked values. The formula can be expressed as:
ρ = 1 - (6∑dᵢ²)/(n(n²-1))
where dᵢ represents the difference between the ranks of corresponding values, and n is the number of observations. This rank-based approach explains why Spearman correlation is described as a non-parametric method—it makes no assumptions about the underlying distribution of the data.
To understand what a “monotonic relationship” means, think of two variables that move in the same direction consistently: whenever one increases, the other always increases (positive monotonic) or always decreases (negative monotonic). A linear relationship (captured by Pearson) requires that this directional consistency follow a straight-line pattern. A monotonic relationship (captured by Spearman) is broader: the relationship could be curved, following an exponential or logarithmic pattern, yet Spearman will still detect it effectively, as long as the directional consistency holds. This flexibility makes Spearman particularly valuable for biological data, where molecular interactions rarely follow simple linear patterns. (Learn more at: Pearson vs Spearman Correlation)
Why Spearman Correlation Matters in Multi-Omics Analysis
Spearman‘s rank correlation offers distinct advantages that align remarkably well with the characteristics of multi-omics data. The suitability of this method for modern biological research arises from several key properties:
• Handling non-normal distributions
Gene expression data in conditions such as cancer frequently follow exponential or heavy-tailed distributions rather than normal distributions. Under these circumstances, Pearson correlation can lead to inflated Type I error rates, falsely identifying associations that aren't meaningful. Spearman’s rank-based approach remains robust regardless of the underlying distribution, providing reliable results where parametric methods falter.
• Resistance to outliers
Multi-omics experiments routinely encounter extreme values—a single sample with anomalously high expression due to biological variation or technical artifacts. Pearson correlation is notoriously sensitive to such outliers, which can dramatically inflate or deflate correlation estimates. Because Spearman operates on ranks, the impact of extreme values is limited: an outlier becomes simply the highest rank, exerting no more influence than any other ranked observation.
• Capturing non-linear monotonic relationships
Many biological regulatory relationships are not strictly linear. Consider enzyme-substrate interactions that exhibit saturation kinetics, or transcription factor binding effects that plateau at high concentrations. As long as these relationships are monotonic, Spearman correlation detects them effectively, whereas Pearson would underestimate the strength of association.
• Applicability to ordinal data
Clinical research often involves variables measured on ordinal scales—tumor staging (I, II, III, IV), pathological grading, or treatment response categories. When correlating such ordinal variables with continuous omics measurements, Spearman correlation is the statistically appropriate choice.
How to Calculate and Report Spearman Correlation in Omics Data Analysis
The calculation and reporting of Spearman correlation in multi-omics studies involves a systematic workflow spanning several key stages. A complete analysis begins with appropriate software tools and implementation, followed by necessary data preprocessing steps to ensure reliable results. Subsequent interpretation of correlation coefficients and statistical significance, together with effective data visualization, forms the foundation for drawing meaningful biological conclusions from the analysis.
3.1 Software Tools for Spearman Correlation
Implementing Spearman correlation in your analysis pipeline is straightforward with standard bioinformatics tools. In R, the base `stats` package provides comprehensive functionality:
- Basic correlation calculation:cor(x, y, method = “spearman”)
- Correlation with significance testing:cor.test(x, y, method = “spearman“)
For Python users, the SciPy library offers equivalent functionality:
rho, p_value = stats.spearmanr(x, y)
These functions handle the rank transformation internally and return both the correlation coefficient and associated p-value.
3.2 Data Preprocessing for Reliable Correlation
Before calculating Spearman correlations, data preprocessing typically involves the following steps:
- Handling missing values: Most implementations automatically exclude cases with missing data (pairwise or listwise deletion). However, for large-scale omics analyses, consider more sophisticated approaches such as imputation using k-nearest neighbors or matrix factorization methods, particularly when missingness exceeds 5-10%.
- Ties in the data: When multiple observations share the same value, they receive identical ranks (the average of the ranks they would have received). This adjustment, built into standard software implementations, ensures accurate correlation estimates even with tied measurements.
- Normalization considerations: Unlike Pearson correlation, Spearman does not assume normality, so data transformation purely for distributional reasons is unnecessary. However, normalization to correct for technical variation (e.g., library size normalization in RNA-seq, batch effect correction) should still be performed prior to correlation analysis to ensure biological rather than technical associations are captured.
3.3 Interpreting Spearman Correlation Coefficients
Spearman‘s correlation coefficient ρ ranges from -1 to +1, with the sign indicating direction:
- Positive correlation (ρ > 0): As one variable increases, the other tends to increase
- Negative correlation (ρ < 0): As one variable increases, the other tends to decrease
- Zero correlation (ρ ≈ 0): No monotonic association detected
While context-specific thresholds vary, general guidelines for interpreting absolute ρ values in biological data:
- |ρ| < 0.3: Weak correlation
- 0.3 ≤ |ρ| < 0.6: Moderate correlation
- |ρ| ≥ 0.6: Strong correlation
Statistical significance is equally important. A high correlation must be accompanied by a significant p-value (typically < 0.05 after multiple testing correction). The p-value tests whether the observed correlation differs significantly from zero.
With very small samples (n ≤ 10) and extremely non-normal distributions, permutation tests provide more reliable p-values than asymptotic approximations. Research has demonstrated that Spearman correlation maintains good performance even with limited sample sizes, though caution is warranted in extreme cases (Rosa et al., 2022).
3.4 Visualizing Spearman Correlations
Effective visualization enhances interpretation of Spearman correlation results:
- Heatmaps display large correlation matrices efficiently, with color intensity representing correlation strength. These are particularly useful for visualizing co-expression patterns across hundreds or thousands of features.
- Correlation networks transform correlation matrices into graph structures, where nodes represent molecular features and edges connect significantly correlated pairs. This approach reveals modular structure and hub genes within biological systems.
- Scatter plots with ranked axes provide intuitive validation: plotting the ranked values of two variables should reveal a monotonic pattern. Adding a LOESS (locally estimated scatterplot smoothing) curve helps visualize the relationship‘s shape and confirm monotonicity.
Pitfalls and Considerations in Spearman Correlation Analysis
While Spearman correlation is robust and widely applicable, several important considerations warrant attention:
-
Loss of information with rank transformation
Converting continuous measurements to ranks does discard some information about the magnitude of differences between values. For data that genuinely satisfy normality and linearity assumptions, Pearson correlation offers greater statistical power. The key is appropriate method selection: when assumptions are met, Pearson is preferred; when they are violated, Spearman provides more reliable results.
-
Correlation does not imply causation
This fundamental principle bears repeating regardless of the correlation method used. A high Spearman correlation between a transcription factor and its predicted target gene may reflect direct regulation, indirect effects through shared pathways, or confounding by a third variable. Experimental validation remains essential for establishing causal relationships.
-
Neglecting multiple testing correction
In multi-omics analyses examining thousands or tens of thousands of feature pairs, the probability of false-positive findings becomes substantial. When performing genome-wide correlation analyses, multiple testing correction methods such as False Discovery Rate (FDR) control must be applied to ensure reported associations are statistically meaningful rather than chance occurrences.
Spearman Correlation Applications in Omics and Bioinformatics
Spearman correlation has found extensive application across diverse areas of omics research, where its robustness to data distribution assumptions proves particularly valuable. These applications span the major omics layers—genomics, proteomics, and metabolomics—as well as integrative analyses that combine multiple molecular modalities. Examining these use cases reveals how Spearman correlation enables researchers to uncover biologically meaningful associations, from gene co-expression networks to cross-omics regulatory relationships, underscoring its versatility as an analytical tool in modern bioinformatics.
Genomics and Proteomics Correlation Analysis
In transcriptomics, Spearman correlation enables construction of gene co-expression networks that reveal functional relationships and regulatory modules. A 2022 study in hepatocellular carcinoma demonstrated the power of this approach, examining correlations between copy number alterations (CNA) and mRNA expression across the genome (Ng et al., 2022). The researchers calculated per-gene Spearman correlation coefficients between CNA and mRNA levels, identifying genes whose expression is particularly sensitive to copy number changes. Pathway enrichment analysis of these genes revealed biological processes significantly associated with CNA-driven gene expression alterations, providing insights into cancer driver mechanisms.
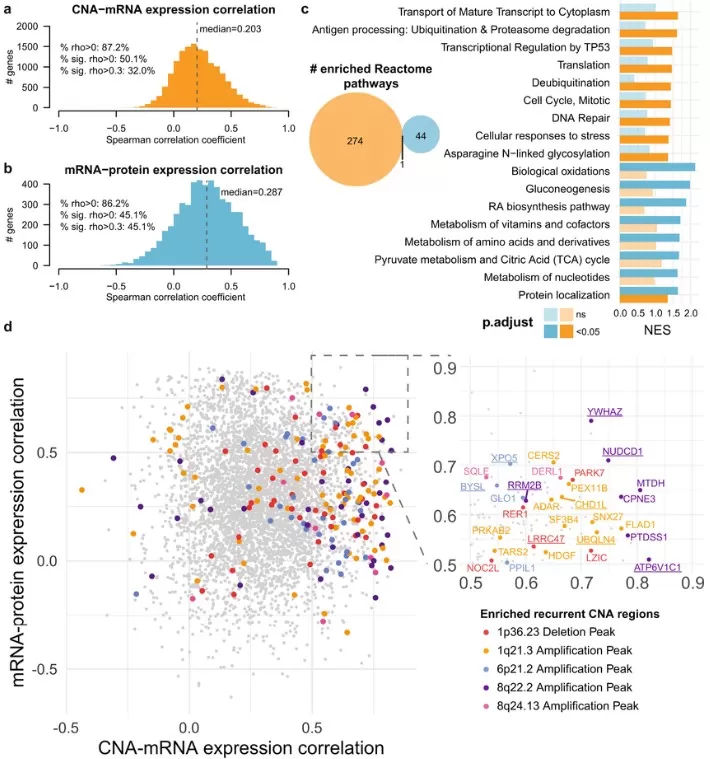
CNA-mRNA-protein correlation.
Image reproduced from Ng et al., 2022, Nature Communications, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
Metabolomics and Metagenomics Integration Analysis
Metabolomic data frequently exhibit complex, non-linear relationships due to pathway interconnectivity and feedback regulation. A metabologenomic study investigating Alzheimer‘s disease mouse models employed Spearman correlation to integrate metabolomic and metagenomic datasets (Favero et al., 2022). The researchers constructed Circos plots and hierarchical heatmaps visualizing correlations between bacterial genera and metabolite classes, revealing associations between gut microbiota composition and metabolic alterations during disease progression. This approach enabled identification of specific microbe-metabolite relationships potentially involved in Alzheimer’s pathogenesis.
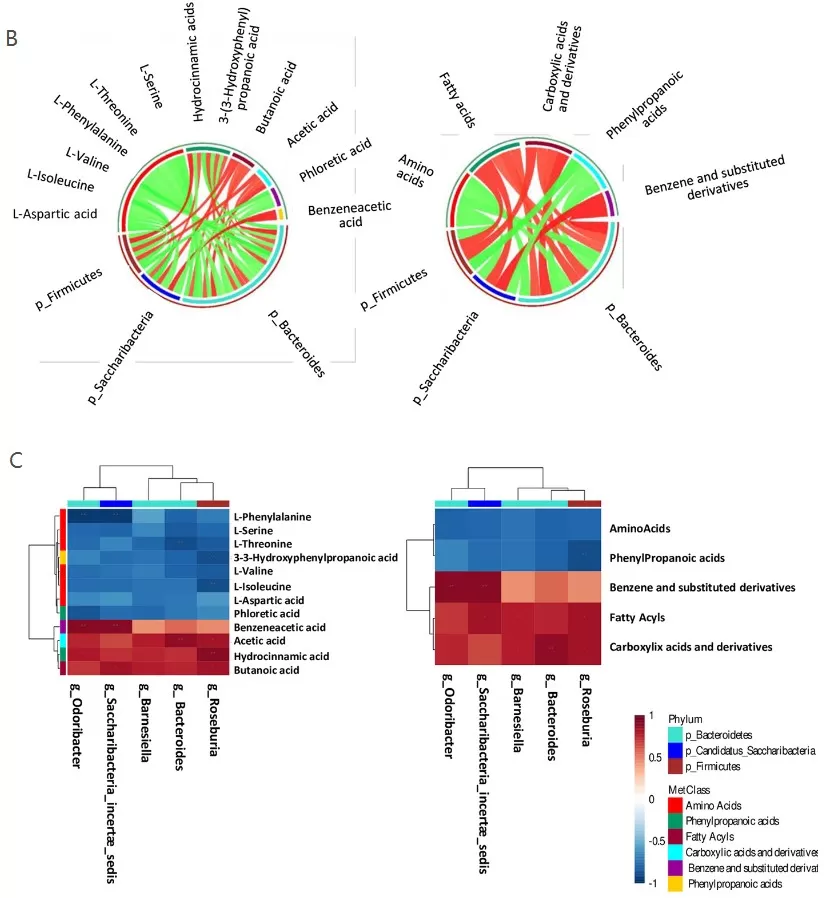
Microbiota and Metabolome Correlation Analysis.
B) Circos plot of Spearman correlation between metabolite-phyla (left panel) and class-metabolites-phyla (right panel). A positive correlation is distinguished by red lines, while a negative correlation by green lines. C) Hierarchical heat maps with Spearman correlation between bacterial genera and metabolites (left panel), and with bacterial genera and metabolites-classes (right panel).
Image reproduced from Favero et al., 2022, PloS one, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
Conclusion and Future of Spearman Correlation in Multi-Omics Research
Spearman‘s rank correlation has established itself as an indispensable tool in the multi-omics researcher’s analytical toolkit. Its ability to handle non-normal distributions, resist outlier influence, and capture monotonic relationships aligns perfectly with the characteristics of modern biological data. When comparing Spearman with Pearson correlation, Spearman offers robustness at the cost of some statistical power under ideal conditions—a trade-off that increasingly favors Spearman as datasets grow larger and more complex. Kendall‘s τ provides an alternative non-parametric approach with similar robustness but different mathematical properties, though Spearman remains more widely used and computationally efficient for large-scale analyses.
Looking toward the future, several trends will shape the application of correlation methods in multi-omics research. First, the emergence of single-cell multi-omics technologies presents new challenges, including technical noise and data sparsity that can obscure true biological associations. Recent methodological developments, such as the SCRaPL framework, demonstrate that careful noise modeling can substantially improve correlation detection sensitivity compared to standard Spearman or Pearson implementations (Maniatis et al., 2022). Second, integration of increasingly diverse omics layers—from epigenomics to metabolomics to imaging data—will require correlation methods that can handle heterogeneous data types and missingness patterns. Finally, the growing scale of public databases and consortia projects will enable more powerful correlation analyses, but also demand rigorous multiple testing correction and validation strategies.
As multi-omics research continues to advance our understanding of complex biological systems, Spearman correlation will remain a fundamental tool—valued for its simplicity, interpretability, and robust performance across the diverse data types that characterize modern molecular profiling.
References
1. Rosa, J. C. D., Aleman, J. O., Mohabir, J., Liang, Y., Breslow, J. L., & Holt, P. R. (2022). The Application of Spearman Partial Correlation for Screening Predictors of Weight Loss in a Multiomics Dataset. Omics: a journal of integrative biology, 26(12), 660–670. https://doi.org/10.1089/omi.2022.0135
2. Ng, C. K. Y., Dazert, E., Boldanova, T., Coto-Llerena, M., Nuciforo, S., Ercan, C., Suslov, A., Meier, M. A., Bock, T., Schmidt, A., Ketterer, S., Wang, X., Wieland, S., Matter, M. S., Colombi, M., Piscuoglio, S., Terracciano, L. M., Hall, M. N., & Heim, M. H. (2022). Integrative proteogenomic characterization of hepatocellular carcinoma across etiologies and stages. Nature communications, 13(1), 2436. https://doi.org/10.1038/s41467-022-29960-8
3. Favero, F., Barberis, E., Gagliardi, M., Espinoza, S., Contu, L., Gustincich, S., Boccafoschi, F., Borsotti, C., Lim, D., Rubino, V., Mignone, F., Pasolli, E., Manfredi, M., Zucchelli, S., Corà, D., & Corazzari, M. (2022). A Metabologenomic approach reveals alterations in the gut microbiota of a mouse model of Alzheimer's disease. PloS one, 17(8), e0273036. https://doi.org/10.1371/journal.pone.0273036
4. Maniatis, C., Vallejos, C. A., & Sanguinetti, G. (2022). SCRaPL: A Bayesian hierarchical framework for detecting technical associates in single cell multiomics data. PLoS computational biology, 18(6), e1010163. https://doi.org/10.1371/journal.pcbi.1010163
Next-Generation Omics Solutions:
Proteomics & Metabolomics
Ready to get started? Submit your inquiry or contact us at support-global@metwarebio.com.


