In spatial metabolomics experiments, spatial resolution is one of the most critical yet challenging parameters to set. Selecting too high a resolution may significantly increase acquisition time, data volume, and cost. Conversely, choosing too low a resolution risks missing essential microstructural information that could be vital for understanding tissue-specific metabolic heterogeneity. In this guide, we provide a comprehensive framework to help you determine the optimal spatial resolution for your spatial metabolomics study, integrating both practical considerations and biological insights.
1. What Is Spatial Resolution in Mass Spectrometry Imaging (MSI)?
In mass spectrometry imaging (MSI), spatial resolution refers to the physical size of an individual pixel in the imaging dataset. Smaller pixels provide finer spatial detail, allowing researchers to observe metabolite distributions at higher granularity.
A simple analogy illustrates the concept: consider a tissue section composed entirely of cells with a diameter of 10 μm. Imaging at a 50 μm resolution means each pixel encompasses a 50 μm × 50 μm area, representing the average metabolite content of roughly 25 cells. Reducing the pixel size (increasing resolution) decreases the number of cells per pixel, thereby revealing previously hidden heterogeneity within tissues. Higher spatial resolution is particularly critical for investigating microstructures or studying cell-type-specific metabolic patterns.
2. Impact of Spatial Resolution on Metabolite Imaging in Spatial Metabolomics
To illustrate how spatial resolution influences the interpretation of MSI data in spatial metabolomics, consider a detailed rapeseed imaging example. Different resolutions reveal distinct tissue features and metabolite distribution patterns, demonstrating that higher resolution does more than improve image clarity—it uncovers microstructural heterogeneity and localized metabolic hotspots, which are critical for accurate biological insights and downstream quantitative analyses. Understanding these differences guides researchers in designing efficient spatial metabolomics experiments.
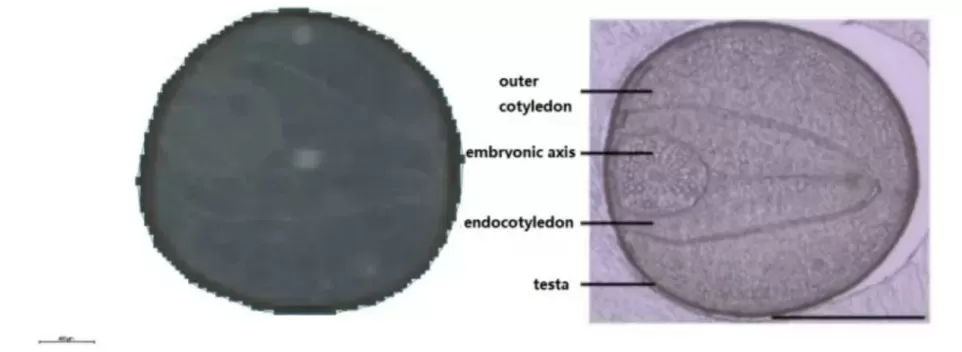
Figure 1. Scanned image and structural schematic of rapeseed.
100 μm resolution: Approximately 400 pixels; metabolite distribution appears homogeneous with no clear tissue specificity. Only the embryonic axis can be roughly distinguished.
 at 100 μm resolution._1775610249_WNo_963d381.webp)
Figure 2. Spatial distribution and segmentation mapping of metabolites (m/z 464.1908) at 100 μm resolution.
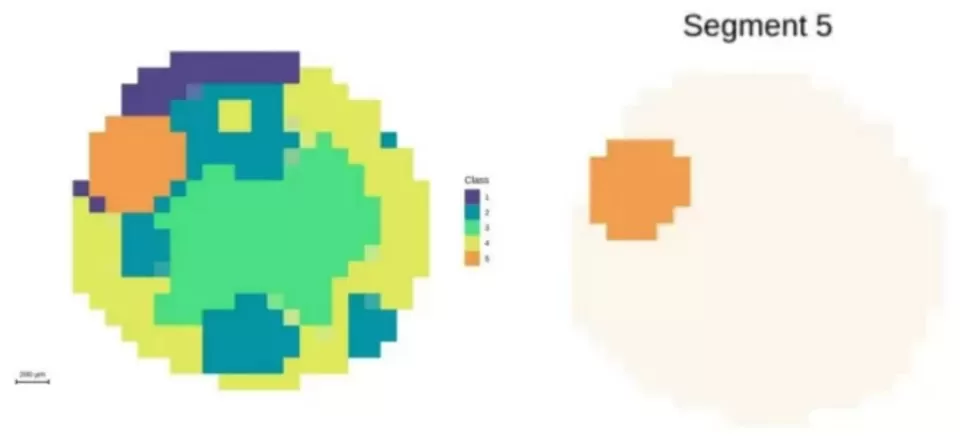
Figure 3. Segmentation analysis highlighting the embryonic axis at 100 μm resolution.
50 μm resolution: Approximately 1,600 pixels; metabolic enrichment in the embryonic axis begins to emerge.
 at 50 μm resolution._1775610316_WNo_961d357.webp)
Figure 4. Spatial distribution and segmentation of metabolites (m/z 464.1908) at 50 μm resolution.
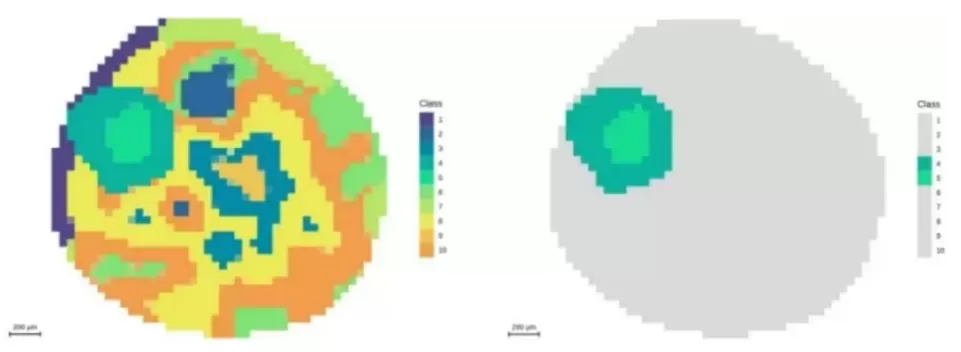
Figure 5. Segmentation analysis highlighting the embryonic axis at 50 μm resolution.
20 μm resolution: Approximately 10,000 pixels; fine metabolite distribution patterns within the seed coat become visible.
 at 20 μm resolution_1775610423_WNo_962d349.webp)
Figure 6. Spatial distribution and segmentation mapping of metabolites (m/z 464.1908) at 20 μm resolution.
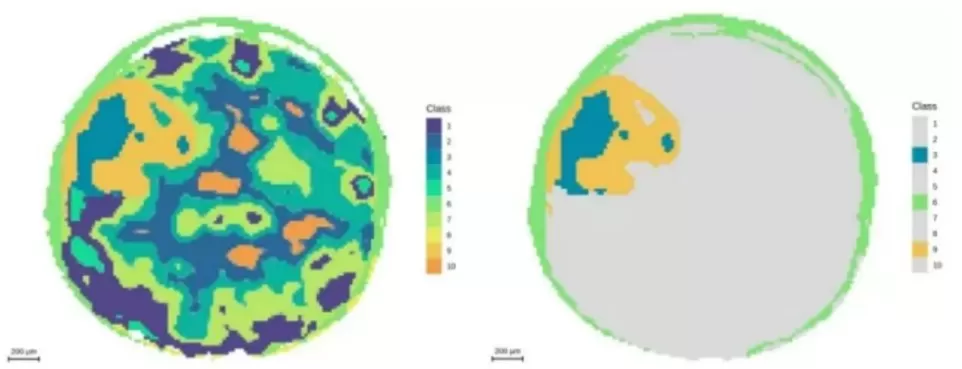
Figure 7. Segmentation analysis highlighting the embryonic axis at 20 μm resolution.
10 μm resolution: Approximately 40,000 pixels; tissue boundaries become sharp, and the seed coat can be subdivided into four distinct subregions.
 at 10 μm resolution_1775610475_WNo_964d373.webp)
Figure 8. Spatial distribution and segmentation mapping of metabolites (m/z 464.1908) at 10 μm resolution.
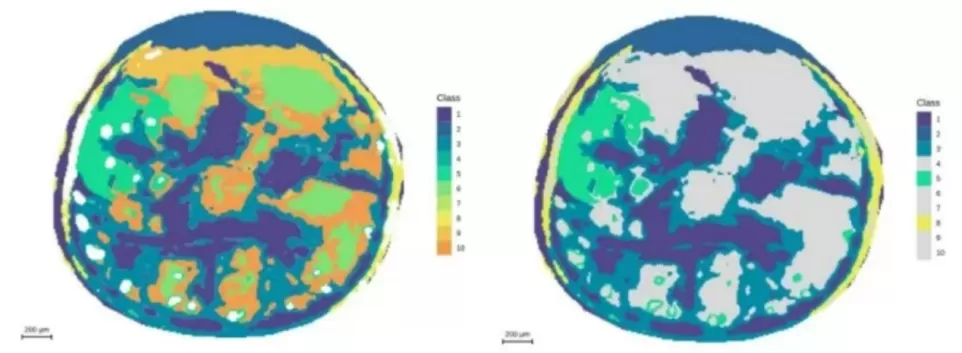
Figure 9. Segmentation analysis highlighting the embryonic axis at 10 μm resolution.
5 μm resolution: Approximately 160,000 pixels; ultra-fine layered structures of the seed coat are clearly resolved.
 at 5 μm resolution_1775610517_WNo_939d369.webp)
Figure 10. Spatial distribution and segmentation mapping of metabolites (m/z 464.1908) at 5 μm resolution.
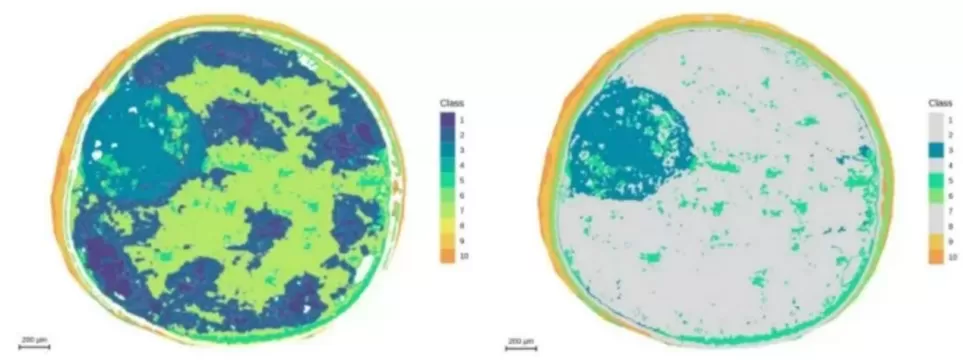
Figure 11. Segmentation analysis highlighting the embryonic axis at 5 μm resolution.
3. Stepwise Framework for Selecting the Right Spatial Resolution in MSI
Choosing the optimal spatial resolution in MSI-based spatial metabolomics requires careful consideration of three interconnected factors: the size of the target structure, the overall sample dimensions, and the specific research objectives. Proper selection ensures meaningful metabolite mapping while avoiding unnecessary data volume or loss of biological detail. This framework helps researchers align experimental design with scientific questions, maximizing the biological insights obtainable from high-quality spatial metabolomics datasets.
Step 1: Assess the Target Structure Size
Core principle: Resolution should be smaller than the smallest structure of interest. To accurately capture microstructural details, it is recommended to follow the "4-pixel rule", meaning that the width of the target structure should span at least four pixels.
Calculation formula:
Required resolution ≤ (Smallest target structure size) ÷ 4
Examples:
- For a mouse renal tubule with a diameter of approximately 30 μm, the required resolution is 7.5 μm or smaller; therefore, a resolution in the range of 5–10 μm is recommended.
- For a glomerular minor axis measuring around 20 μm, the required resolution is 5 μm or smaller; thus, a resolution of 5 μm is recommended.
- For a seed coat with a thickness between 26 and 28 μm, the required resolution is approximately 6.5–7 μm; a resolution of 5–10 μm is suitable.
This approach ensures that critical structures are represented by multiple pixels, minimizing the risk of losing spatial information.
Step 2: Determine Base Resolution According to Sample Size
The total pixel count in a tissue section should exceed 10,000 for statistically meaningful segmentation. The required base resolution varies depending on sample dimensions:
- Millimeter-scale samples (small tissues, insects, embryos): 5–10 μm
- 5 mm–1 cm samples (mouse organs, plant seeds): 20–40 μm
- Centimeter-scale samples (rat organs, large tissue sections): 50–100 μm
Choosing an appropriate base resolution balances image coverage, acquisition time, and computational feasibility, ensuring sufficient data density for analysis without unnecessary resource expenditure.
Step 3: Adjust Resolution According to Research Objectives
- Exploratory studies: Start with 50 μm to obtain a global view of metabolite distributions. Once regions of interest are identified, increase the resolution to capture microstructural details.
- Validation or targeted studies: Directly select a resolution sufficient to resolve the structure of interest (e.g., 20 μm or higher). This targeted approach avoids oversampling and optimizes resource use.
4. Recommended Spatial Resolutions for Common Spatial Metabolomics Applications
To provide practical guidance, this table summarizes recommended MSI spatial resolutions for typical spatial metabolomics applications. By aligning resolution choices with sample type, tissue heterogeneity, and research goals, researchers can efficiently balance coverage, data acquisition time, and the ability to resolve microstructures. These recommendations serve as a starting point for optimizing experimental design and ensuring that metabolite imaging produces robust, reproducible, and biologically meaningful results.
| Research Scenario | Recommended Resolution | Key Consideration |
|---|---|---|
| Brain tissue | 20–50 μm | Suitable for most regional analyses; single-neuron studies require 10 μm or higher |
| Tumor microenvironment | 50 μm | Adequate for tumor vs. normal tissue; boundaries or infiltration require 20 μm or higher |
| Kidney research | 50 μm | Overall partitioning sufficient; glomeruli/tubule analysis requires 5–20 μm |
| Plant tissues | 20–50 μm | Standard analysis; seed coat or pollen microstructures require 5–10 μm |
| Small-volume samples | ≤20 μm | Ensures sufficient pixel representation for meaningful segmentation |
5. Best Practices for Spatial Resolution in Metabolite Imaging
Selecting spatial resolution in spatial metabolomics experiments involves a strategic trade-off between detail depth, detection breadth, and cost-efficiency. There is no universally ideal resolution; the best choice depends on the biological question, sample characteristics, and imaging objectives. The following best practices in MSI experiment design can help ensure accurate mapping of metabolite distributions, reveal tissue-specific heterogeneity, and maximize the utility of spatial metabolomics data for both exploratory and targeted studies.
| Research Stage | Recommended Resolution | Key Consideration |
|---|---|---|
| Exploratory studies, differential molecule screening | 50 μm | Broad coverage to identify "what is present" |
| Targeting specific structures/cell populations | 20 μm | Prevents averaging; determine "where it is localized" |
| Precise microstructure analysis | 5–10 μm | Pixel-level accuracy to reveal "how it is distributed" |
Once your biological question is clearly defined, choosing the appropriate spatial resolution becomes much more straightforward. Lower resolutions are efficient for broad tissue-level screening, while higher resolutions are better suited for resolving fine structures and localized metabolic features. In practice, the optimal setting is the one that best matches your target structure, sample size, and study objective.
MetwareBio Spatial Metabolomics Service
Need help choosing the right spatial resolution for your MSI study? MetwareBio offers spatial metabolomics services with flexible resolution options from 5 μm to 100 μm, helping researchers capture both global metabolic patterns and fine microstructural details.
Our team can recommend a suitable imaging strategy based on your sample type and research goals. Contact us to discuss your project and find the best resolution for your study.


