A Practical Guide to OPLS-DA: Principles, Workflow, and Result Interpretation in Omics Data Analysis
High-dimensional omics datasets rarely speak in a simple way. In metabolomics, proteomics, and transcriptomics, researchers often need to go beyond asking whether samples differ and start asking which variables drive that separation. Orthogonal partial least squares discriminant analysis, or OPLS-DA, is widely used for exactly that purpose because it combines supervised classification with a model structure that is easier to interpret than standard PLS-DA. At the same time, OPLS-DA is also one of the most frequently misused multivariate statistical methods in omics data analysis, especially when preprocessing, validation, or result interpretation is handled loosely. This guide provides a systematic introduction to the principles, functions, workflow, result interpretation, applications, and key precautions of OPLS-DA, aiming to support researchers with practical, accurate, and informative guidance for omics data analysis.
1. What Is OPLS-DA?
OPLS-DA is a supervised multivariate analysis method derived from partial least squares. In practical terms, it models the relationship between an input matrix of measured variables and a predefined class label, such as control versus treatment or responder versus non-responder. What makes OPLS-DA distinctive is that it separates the variation in X into two parts: predictive variation related to the class labels and orthogonal variation unrelated to those labels. That separation does not magically make the model more truthful, but it often makes the model easier to interpret.
This is the core difference between OPLS-DA and PLS-DA. In PLS-DA, class-related and unrelated sources of variance may remain mixed within the same latent structures. In OPLS-DA, orthogonal structure is filtered out from the predictive component, which can make score plots, loadings, and variable importance patterns more straightforward to read. That interpretability is one reason OPLS-DA became especially popular in metabolomics, where datasets are often high-dimensional, collinear, noisy, and biologically heterogeneous. (Learn more at: PCA vs PLS-DA vs OPLS-DA)
2. What OPLS-DA Can Do in Omics Data Analysis?
OPLS-DA is useful because it does more than generate a visually attractive separation plot. When applied appropriately, it can help organize complex omics data into a model that supports class discrimination, variable prioritization, and more interpretable visualization. Its real value lies in linking sample groups to the subset of structured variation that most strongly relates to the study question.
Class discrimination and sample pattern recognition
The most common use of OPLS-DA is to distinguish predefined groups in a supervised setting. In metabolomics, that may mean healthy versus diseased samples, before versus after treatment, or different environmental exposures. Because the class labels are incorporated during modeling, OPLS-DA often reveals group-related structure more clearly than PCA, which is unsupervised and does not know which samples belong together.
Variable prioritization and biomarker screening
OPLS-DA is also frequently used to rank variables that contribute most strongly to class separation. In practice, researchers often combine loadings, S-plots, and VIP scores to identify candidate metabolites, proteins, or transcripts that may deserve further statistical and biological evaluation. This makes OPLS-DA attractive in biomarker discovery workflows, although the model should be treated as a prioritization tool rather than a standalone proof generator.
Noise separation and model interpretability
A major practical advantage of OPLS-DA is its ability to isolate variation unrelated to the target class in orthogonal components. In real datasets, much of the measured variance can come from individual heterogeneity, technical drift, matrix effects, or unrelated biology. By separating that structure from predictive variation, OPLS-DA can simplify interpretation and make the class-associated signal easier to inspect.
3. Step-by-Step OPLS-DA Workflow
The workflow below provides a practical path for performing OPLS-DA in software such as SIMCA or R packages like ropls, while following the same general analytical logic across platforms. Although the interface and parameter settings may vary slightly between tools, the core steps remain largely consistent. A clear and well-structured workflow is essential for building a reliable model and avoiding misleading interpretations.
1) Prepare the data matrix and sample metadata
Start with a clean feature table in which rows represent samples and columns represent variables. The sample annotation file should contain the correct class labels together with any relevant covariates, such as batch, sex, time point, or treatment group. Before building the model, it is essential to confirm that sample IDs, feature names, and metadata are fully aligned, since even minor mismatches at this stage can compromise the entire analysis.
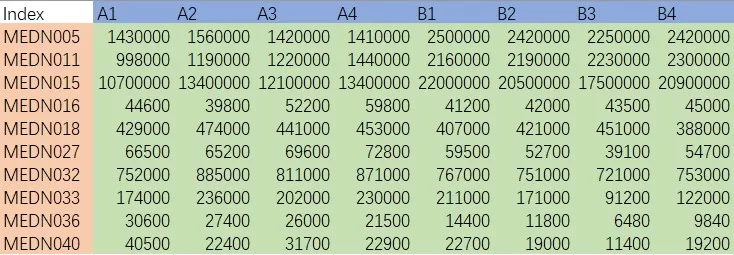
Example of Quantitative Data Matrix of Metabolomics Analysis
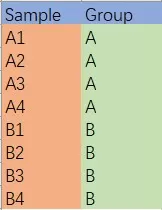
Example of Sample Grouping Information
2) Handle missing values and low-quality features
Variables with excessive missing values, poor reproducibility, or weak analytical reliability should be removed before modeling. Missing data should then be addressed using an imputation strategy that is appropriate for the data type and missingness pattern, rather than by arbitrary replacement. Features dominated by noise are best filtered out before supervised analysis so that the model reflects meaningful biological structure rather than technical instability.
3) Normalize, transform, and scale the data
Data preprocessing is a critical part of OPLS-DA because the model is highly sensitive to differences in variable magnitude and distribution. Normalization helps reduce unwanted intensity variation across samples, while log transformation is often used to stabilize variance and reduce right-skewed distributions. Scaling further adjusts the relative influence of variables, and common choices such as autoscaling or Pareto scaling should be reported clearly because they can substantially affect score patterns, loadings, and VIP values.
4) Build the OPLS-DA model
Once the dataset has been properly preprocessed, the OPLS-DA model can be fitted using the processed X matrix and the class vector Y. At this stage, the software extracts predictive and orthogonal components that describe the relationship between the measured variables and the predefined sample groups. The resulting model provides the foundation for subsequent visualization, discrimination analysis, and variable interpretation.
5) Select predictive and orthogonal components
Component selection should be guided by model performance and stability rather than by the desire to produce a more visually separated score plot. Adding too many components can improve the apparent fit of the training data while reducing the model’s robustness and generalizability. For that reason, the number of predictive and orthogonal components should be chosen carefully based on cross-validation results and the overall interpretability of the model.
6) Perform cross-validation and permutation testing
Model validation is one of the most important stages in OPLS-DA analysis. Cross-validation estimates how well the model performs on held-out data, while permutation testing evaluates whether the observed class separation is stronger than what could be achieved by random label assignment. Together, these procedures help distinguish a biologically meaningful model from one that is merely overfitted, and they are essential for establishing confidence in the results.
7) Export score plots, loading plots, and VIP results
After the model has been adequately validated, the main outputs can be exported for interpretation and downstream analysis. These typically include the score plot, loading plot, S-plot when applicable, VIP table, and key quality metrics. Taken together, these results support sample discrimination assessment, candidate feature prioritization, and subsequent integration with biological annotation, pathway analysis, and mechanism-focused interpretation.
4. How to Interpret OPLS-DA Results Correctly
OPLS-DA does not simply produce a picture of truth; it produces a model-based summary of labeled data. A careful interpretation requires understanding what each output represents, what each metric measures, and what none of them can prove on their own.
i. How to read the OPLS-DA score plot
The score plot displays how samples are distributed along the predictive and orthogonal components of the OPLS-DA model. Each point represents an individual sample, while different colors indicate different sample groups. The ellipse typically represents the 95% confidence interval, providing a visual summary of the dispersion pattern within each group. The predictive component scores reflect inter-group variation, and the percentage shown indicates the proportion of the total dataset variance explained by that predictive component. In contrast, the orthogonal component scores reflect intra-group variation that is unrelated to the predefined class separation, with the corresponding percentage indicating its explanatory contribution to the overall dataset. Clear separation in the score plot may suggest meaningful group-related structure, but it should not be considered conclusive evidence on its own, since an apparently strong separation can also result from overfitting, confounding factors, or unstable data preprocessing.
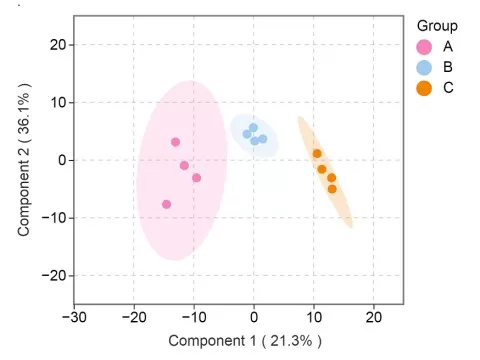
OPLS-DA Score Plot
ii. What predictive and orthogonal components mean
The predictive component captures variance associated with the class labels, while orthogonal components capture systematic variance that is unrelated to the labels. This decomposition is what gives OPLS-DA its interpretive advantage over standard PLS-DA, but it does not remove the need for biological and statistical scrutiny.
iii. How to Interpret the OPLS-DA S-Plot
OPLS-DA S-plots are often used to visualize covariance and correlation patterns for variable selection. Each point represents one variable, such as a metabolite, protein, or transcript. The x-axis reflects the covariance of each variable with the predictive component, indicating the magnitude of its contribution to class discrimination, while the y-axis reflects the correlation of that variable with the predictive component, indicating the stability and consistency of its contribution across samples. Variables located farther away from the origin generally have a stronger influence on group separation. In particular, variables distributed in the upper right and lower left regions are often considered the main contributors to discrimination between groups, because they show both high contribution and strong correlation. By contrast, variables clustered near the center usually contribute little to class separation and are less informative for feature screening.
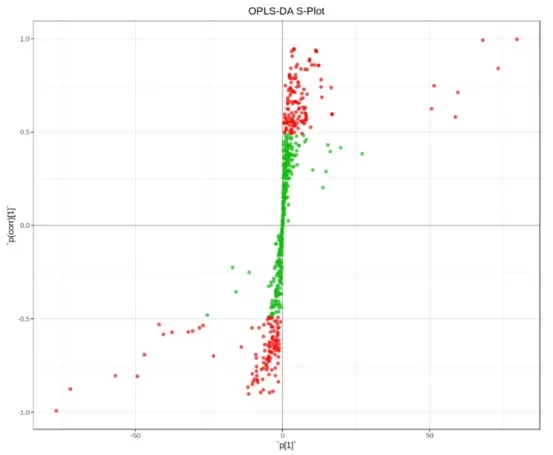
OPLS-DA S-Plot
iv. What VIP scores mean in OPLS-DA
VIP scores summarize the contribution of variables to explaining Y in projection-based models. A common heuristic is that variables with VIP greater than 1 are influential, but VIP does not equal statistical significance, effect size, or biological importance. A variable can have a high VIP and still fail independent statistical testing or biological validation.
v. What R2X, R2Y, and Q2 mean in OPLS-DA
R2X, R2Y, and Q2 are three of the most commonly reported parameters in OPLS-DA, and together they provide a basic view of model quality from different angles. R2X indicates how much variation in the input data matrix X is explained by the model, while R2Y reflects how much of the class structure in Y is captured. Q2, estimated by cross-validation, measures predictive ability on unseen data. In general, R2X and R2Y describe model fit, whereas Q2 is more informative for model robustness. A high R2Y combined with a low Q2 often suggests overfitting.
vi. How to interpret a Permutation Test Plot
Permutation testing assesses whether an OPLS-DA model captures real class-related structure rather than random variation. In this plot, the original model is compared with multiple models built from randomly permuted class labels. The main focus is whether the original R²Y and especially Q² values are clearly separated from the permuted distributions. Clear separation supports model reliability, whereas substantial overlap suggests that the observed discrimination may not be robust.
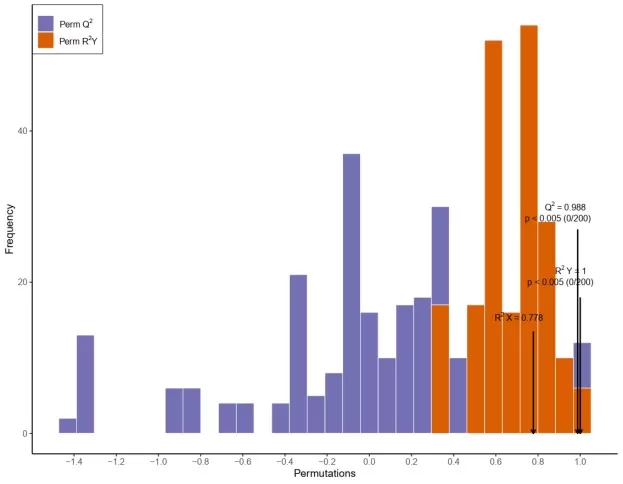
OPLS-DA Permutation Test Plot
5. How to Identify Differential Features After OPLS-DA
OPLS-DA can help prioritize candidate features, but it should not be the only basis for declaring differential metabolites, proteins, or transcripts. In practice, stronger workflows combine multivariate importance with univariate evidence such as fold change, p values, and multiple-testing correction. This reduces the chance of selecting variables that look influential in the projection model but are not robust across other criteria.
A practical strategy is to treat OPLS-DA as one layer of evidence. Variables with strong VIP scores can be cross-checked against fold change, confidence intervals, FDR-adjusted significance, analytical quality, and annotation confidence. For metabolomics in particular, downstream pathway analysis and biological context are essential. A prioritized list is only useful if it can be linked back to chemistry, pathways, phenotype, and mechanism.
Key Metrics for Differential Metabolite Selection After OPLS-DA
|
Metric |
FC |
P value |
FDR |
Q value |
VIP |
|
Full name |
Fold Change |
Probability value |
False Discovery Rate |
Adjusted significance measure based on FDR |
Variable Importance in Projection |
|
What it reflects |
Magnitude of abundance difference between groups |
Statistical significance under the null hypothesis |
Expected proportion of false positives after multiple testing correction |
Minimum FDR at which a feature is considered significant |
Contribution of a variable to class discrimination in the OPLS-DA model |
|
Main strength |
Intuitive and biologically easy to understand |
Widely used and easy to interpret |
More reliable than raw p values in high-dimensional omics data |
Directly reflects multiple-testing-adjusted significance |
Captures multivariate importance and group-separating power |
|
Main limitation |
Does not account for variance or statistical significance |
Sensitive to sample size and multiple testing |
Can be conservative in small datasets |
Can be confused with Q2 in OPLS-DA if terminology is unclear |
Does not indicate statistical significance or analytical reliability |
|
Recommended use |
Evaluate effect size and direction of change |
Perform initial significance assessment |
Control false positives in large-scale screening |
Use as an adjusted significance metric for robust screening |
Prioritize features together with FC and adjusted significance |
6. Applications and Appropriate Use of OPLS-DA
Although OPLS-DA is a powerful supervised method for omics data analysis, its performance and interpretability depend heavily on whether it is applied in an appropriate analytical context. Understanding where OPLS-DA works well and where it should be used more cautiously is essential for drawing reliable biological conclusions.
6.1 Best Applications of OPLS-DA in Metabolomics
OPLS-DA is most effective when the research question is clearly supervised and the sample classes are biologically meaningful, well defined, and supported by a reasonably balanced study design. Under these conditions, the method can be highly useful for separating class-related variation from unrelated background structure and for identifying variables that contribute most strongly to group discrimination. This is one of the main reasons why OPLS-DA is especially common in metabolomics, where researchers often work with predefined experimental groups and high-dimensional datasets that benefit from interpretable supervised modeling. Typical applications include case-control comparisons, intervention studies, toxicology, nutrition research, pharmacometabolomics, and mechanistic experiments in which the biological contrast is already clearly established.
6.2 OPLS-DA Applications in Proteomics and Transcriptomics
Although OPLS-DA is most frequently discussed in the context of metabolomics, its application is not limited to that field. It can also be used in proteomics and transcriptomics for supervised classification, sample pattern recognition, and variable prioritization, particularly when researchers want a latent-variable model that offers both discrimination and interpretability. In these settings, OPLS-DA can provide a useful complement to standard differential analysis by helping reveal structured group-related variation across complex omics datasets. At the same time, its role may vary depending on the field and study objective, since proteomics and transcriptomics often rely more heavily on linear models, clustering approaches, or machine learning methods as primary analytical frameworks.
6.3 When OPLS-DA Should Be Used with Caution
OPLS-DA should not be treated as a universal solution for all omics datasets. It should be used with particular caution in purely exploratory studies, very small cohorts, heavily imbalanced group designs, and datasets dominated by batch effects or other uncontrolled confounding factors. In these situations, the supervised nature of the method can increase the risk of overfitting or produce visually convincing but biologically unreliable separation. When the study design is weak or the source of variation is still unclear, unsupervised exploration with PCA, more conventional statistical modeling, or carefully benchmarked classification methods may provide a more appropriate starting point.
7. Common Pitfalls and Misinterpretations in OPLS-DA
A reliable OPLS-DA analysis is defined as much by what it avoids as by what it reports. Many weak studies do not fail because OPLS-DA itself is flawed, but because the model is treated as a shortcut to certainty rather than a tool that requires careful validation and interpretation. Several recurring problems are especially important to recognize in practical omics data analysis.
- Overfitting in small-sample, high-dimensional datasets: Omics datasets typically contain far more variables than samples, which makes supervised models particularly vulnerable to overfitting. When validation is weak or too many components are included, the model may capture random structure in the training data rather than reproducible biological differences, resulting in an apparently strong but unreliable separation.
- Overinterpreting visual group separation: A clearly separated score plot can be informative, but it is not definitive evidence of biological truth or predictive robustness. Visual discrimination alone does not prove that the model is statistically valid, reproducible, or generalizable, and it should always be interpreted together with validation metrics and study design context.
- Using VIP values alone to identify biomarkers: VIP is useful for ranking variables according to their contribution to class discrimination, but it is not a substitute for statistical significance, analytical reliability, annotation quality, or biological relevance. A feature with a high VIP score may still be unstable, poorly identified, or unsupported by independent statistical and biological evidence.
- Neglecting preprocessing, batch effects, and confounding factors: Supervised models can easily absorb hidden technical variation if the input data are not carefully prepared. When normalization, transformation, scaling, or covariate control are inadequate, the model may separate technical artifacts such as batch structure rather than the biological effect of real interest.
- Reporting the model without sufficient validation details: An OPLS-DA result cannot be properly evaluated if key methodological details are missing. Without clear reporting of cross-validation strategy, permutation testing results, preprocessing steps, and component selection criteria, readers cannot determine whether the model is robust, reproducible, or simply overfit.
8. One-Click OPLS-DA Analysis with the Metware Cloud Platform
For researchers who need a faster and more accessible way to perform OPLS-DA, the Metware Cloud Platform provides a practical one-click cloud tool for streamlined analysis. Backed by a professional expert team in multi-omics bioinformatics analysis, the Metware Cloud Platform offers free registration, supports user-uploaded data in the required input format, and includes more than 50 omics data analysis tools with text instructions and video tutorials.
In the OPLS-DA analysis panel, users only need to upload a sample quantitative matrix and a sample grouping file, then select the required processing parameters, such as log transformation, zero-value replacement, normalization method, and VIP threshold, while also customizing visualization settings including shape, color, and size. Once the job is submitted, the platform generates a downloadable compressed result package containing the completed OPLS-DA outputs. With its user-friendly interface, straightforward workflow, and free unlimited access, this cloud-based tool is especially suitable for researchers who want reliable OPLS-DA results without relying on programming.
Next-Generation Omics Solutions:
Proteomics & Metabolomics
Ready to get started? Submit your inquiry or contact us at support-global@metwarebio.com.


