Proteomics Data Normalization: How to Remove Batch Effects and Non-biological Bias
As mass spectrometry throughput and robustness continue to improve, proteomics is increasingly moving from small-scale discovery studies toward multi-center and large-cohort analyses. However, subtle differences in sample preparation, chromatographic separation, instrument performance, and computational processing can accumulate into systematic, non-biological bias (batch effects). These effects may mask true biology or create spurious signals. Normalization and batch correction aim to reduce technical variation while preserving genuine biological differences, thereby improving comparability and the reliability of downstream statistics.
1. What Are Batch Effects in Proteomics?
Batch effects are systematic, non-biological shifts in measured abundance that arise from technical factors, such as acquisition date and injection order, reagent lots, operators, LC columns, instrument platforms, and database-search or quantification parameters. Because the proteomics pipeline is long (sampling → lysis/digestion → cleanup → LC → MS → identification/quantification → protein inference), small drifts at multiple steps can compound into substantial bias in the final quantitative matrix.
Proteomics datasets are especially vulnerable because they typically exhibit (i) missing values, (ii) intensity-dependent variance (lower-abundance features are less stable), and (iii) long-run drift during extended acquisition. For these reasons, normalization and batch correction are generally considered essential quality-control steps in large-scale studies [1-2].
2. Common Sources of Batch Effects in LC–MS Proteomics
2.1 Variability in sample preparation
Sample preparation often contributes the largest share of technical variability. Differences in collection, storage, freeze-thaw history, extraction chemistry, digestion efficiency, cleanup, and enrichment can alter both identification rates and quantitative accuracy. In plasma proteomics, hemolysis or platelet contamination may introduce strong, non-biological signatures, while oxidation or processing-induced modifications can skew quantification.
2.2 Chromatography and LC separation
LC-MS/MS remains the backbone of proteomics. Yet chromatography introduces its own failure modes, including retention-time drift, column aging, carryover, and clogging. Nano-flow LC can offer higher sensitivity, but in long cohorts it is often less reproducible. In contrast, micro-flow LC-MS/MS has been reported to provide stable performance across thousands of injections with low retention-time variation and improved quantitative precision, making it attractive for large clinical-style studies [2].
2.3 Instrument performance fluctuations
Even on the same instrument model, performance can drift over time due to ion-source contamination, calibration shifts, changes in resolution, and temperature or vacuum fluctuations. These effects often produce nonlinear trends with injection order and disproportionately impact low-abundance features.
2.4 “Invisible” computational batches
Batch structure can also be introduced during computational processing. Differences in software versions, FDR settings, match-between-runs behavior, spectral-library updates, peak-extraction parameters, or protein inference rules may create systematic differences that mimic technical batches. Practical mitigation includes standardizing software versions and parameters, reprocessing all raw files in a single consistent run when possible, and capturing complete metadata for traceability.
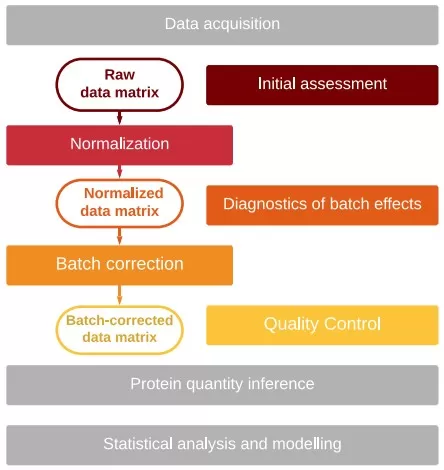
Batch effect processing workflow [1]
Image reproduced from Čuklina et al., 2021, Mol Syst Biol, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
3. Normalization vs Batch Correction: What’s the Difference?
Normalization typically refers to global adjustments that align sample-scale or distributional differences (e.g., loading amount, overall signal intensity). Batch correction refers to statistical removal of batch-associated structure, often via explicit batch variables or latent-factor models.
A commonly used workflow is:
• Filtering and QC (remove failed runs, extreme outliers, obvious contaminants).
• Transformation (often log2; alternatively variance-stabilizing transformations).
• Global normalization (median, total-signal, quantile, or related approaches).
• Drift correction using QC samples (if available).
• Batch correction using known batch labels or latent-factor approaches.
• Post-correction validation (check that biology is preserved and technical structure is reduced).
For peptide-to-protein summarization workflows, the correction level matters: correcting at the peptide level can reduce bias introduced during protein inference, whereas correcting only after protein summarization may leave peptide-level drift partially unaddressed. The best choice depends on data completeness and the quantification strategy.
4. Practical Methods for Proteomics Normalization & Batch Correction
Batch-effect correction reduces differences attributable to technical factors (e.g., preparation batches or measurement batches) by adjusting feature abundances (genes, peptides, metabolites) across samples during data transformation. Many algorithms have been proposed, and within-batch mean/median-based normalization has been widely used in proteomics preprocessing. Below are several commonly used methods. Below is a practical overview of commonly used methods. In practice, methods are often combined, and the choice should be guided by experimental design, QC availability, missingness patterns, and the risk of confounding between batch and biological groups.
4.1 Global scaling normalization
Equalize medians (median normalization): robust and easy to interpret
TIC / sum scaling (total-signal normalization): sensitive to high-abundance signals/contamination
Shared-protein scaling: useful when missingness is substantial
In a multi-sample quantitative matrix, protein intensities can vary widely across samples, and high-intensity samples may dominate comparisons. Therefore, raw intensities are typically not directly comparable and require scaling to a common reference. Median normalization is a frequently used within-batch approach that reduces between-sample technical differences arising from factors such as processing, loading, fractionation, and instrument response.
Median normalization is simple and highly robust. Its key elements are:
1) Assumption: Across samples, most proteins remain relatively stable, so distribution centers should be similar. In proteomics, it is commonly assumed that only a subset of proteins changes substantially.
2) Procedure: For each sample, divide all protein abundance values by the sample median: X’ = X / Median(X), where X is the original value and X’ is the normalized value.
3) Effect: After normalization, sample medians are aligned (often to 1 or another constant), improving comparability across samples.
Because it aligns global levels while being resistant to outliers, median normalization is often recommended as a baseline preprocessing step before downstream analyses such as differential abundance testing.
Quantile normalization is based on the principle of forcing all samples to share the same abundance distribution. In many proteomics datasets, overall distributions differ substantially across samples; quantile normalization enforces comparability by aligning distributions across samples. The typical steps are:
1) Sorting: Sort each sample (column) in ascending order.
2) Compute mean quantiles: At each rank position, compute the mean value across samples to form an average quantile vector.
3) Replace: Replace each original value with the corresponding value from the average quantile vector based on rank (not magnitude).
Quantile normalization is a strong method that can effectively reduce technical differences in cross-batch comparisons and multi-strategy integration. However, it should be applied cautiously: if true biological effects shift global distributions, aggressive alignment may attenuate meaningful biological signals.
4.2 Intensity dependence bias correction: variance stabilization (VSN) and cyclic LOESS
Proteomics data commonly exhibit heteroscedasticity (variance increases with mean intensity). In label-free proteomics, technical noise is often intensity-dependent: low-abundance features show larger relative fluctuations, whereas high-abundance features are more stable. If only median or total-signal scaling is applied, several issues may persist: (i) variance remains larger at low intensity, biasing analyses toward high-abundance proteins; (ii) PCA or clustering may still reflect abundance-driven structure or run-to-run drift; and (iii) residuals may still depend on intensity, violating homoscedasticity assumptions in many linear models. For these reasons, VSN and (cyclic) LOESS are often applied (in place of, or alongside, log transforms depending on implementation) to make variability more consistent across the intensity range and improve the reliability of downstream modeling (e.g., linear models, ComBat, SVA) [3].
VSN (Variance Stabilizing Normalization) is not a simple scaling operation. Instead, it applies a parameterized transformation designed to flatten the mean–variance relationship, making variance more uniform across intensities [4]. (Cyclic) LOESS can be used to correct nonlinear, intensity-dependent bias and can be viewed as a combined transformation and normalization approach that may better match proteomics noise structure than a plain log2 transform. These methods are most suitable when sample size is moderate to large, downstream analyses rely on linear modeling, and diagnostic plots (e.g., MA/RLE) indicate clear intensity-dependent variability. However, if VSN is applied to extremely sparse matrices with severe missingness, missingness patterns may be fitted as signal, yielding stabilized variance but potentially dampening biological differences [4-5]. In addition, applying log2 and then VSN can amount to applying transformations twice; in many workflows, VSN is used as an alternative to log2, or the VSN implementation performs its own transformation.
4.3 QC-based drift correction: QC-LOESS / QC-RLSC
When QC injections are sufficiently frequent and stable, a LOESS model of feature intensity versus injection order can be fitted and used for correction (the QC-RLSC concept). This approach is particularly effective for long-term drift, provided the QC material is stable and representative of the overall workflow.
During continuous MS acquisition, instrument response may drift nonlinearly due to subtle loading differences, ion-source contamination, and environmental fluctuations. Such drift can introduce systematic differences in measured intensities for the same underlying abundance, often affecting low-abundance proteins more strongly. LOESS-based QC correction (QC-LOESS) is widely used to reduce this instrument-driven signal drift.
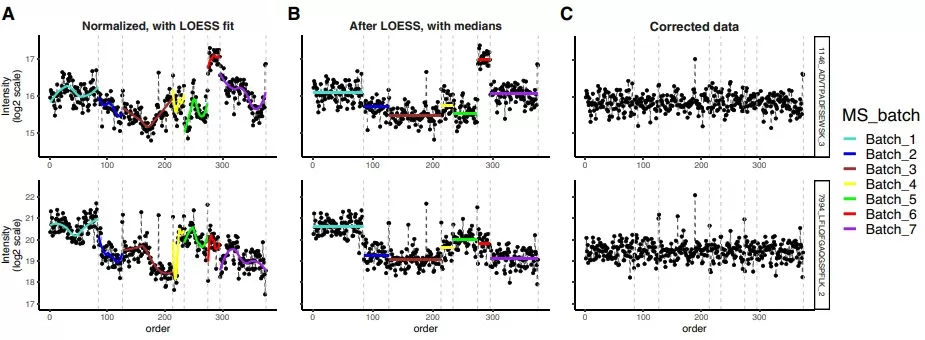
LOESS correction effectively reduces batch effects
Image reproduced from Čuklina et al., 2021, Mol Syst Biol, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
The core assumption is that most proteins in QC samples should remain stable over time; therefore, systematic deviations are attributed to drift. A typical workflow includes:
1) Select QC samples: In many designs, a pooled QC is injected at regular intervals (e.g., every ~30 injections).
2) Construct an M–A plot: For adjacent QC injections, compute M (log ratio) and A (mean intensity).
3) LOESS fitting: Fit a smooth LOESS curve on the M–A relationship to model drift across intensity ranges.
4) Apply correction: Use the fitted curve to shift or scale intermediate sample intensities and reduce drift effects.
For very large studies requiring multi-day continuous acquisition, QC-LOESS can be highly effective. However, QC injections must be stable; otherwise, the fitted curve may be unreliable. In practice, QC-LOESS is often combined with other approaches—for example, applying QC-LOESS to correct time drift first [5], followed by ComBat or RUV to address discrete batch differences (e.g., different instruments or acquisition dates).
4.4 Known batch variables: ComBat and regression-based correction
ComBat (empirical Bayes): adjusts batch-specific mean and variance; supports covariates
limma::removeBatchEffect: removes batch effects using a linear model; often used for visualization after correction
MSstats: a linear mixed-model framework with multiple normalization options for DDA/DIA and label-based workflows
Among these methods, ComBat (Combining Batches) is a widely used empirical Bayes approach originally developed for microarray batch correction and subsequently applied to proteomics, metabolomics, and other high-throughput omics datasets when batch labels are known.
ComBat estimates batch-specific shifts in mean and variance via statistical modeling. It is commonly used in proteomics for large-scale integration across batches or platforms and often performs better than simple median-based adjustments. The main steps include:
1) Model specification: Model protein abundance as a function of batch effects, biological covariates (e.g., age, sex, disease status), and residual error.
2) Parameter estimation: Use empirical Bayes shrinkage across features to estimate batch-specific mean and variance shifts, which can improve robustness—especially when per-batch sample sizes are limited.
3) Correction: Apply a linear adjustment based on estimated parameters to reduce between-batch differences and make batches statistically comparable. In practice, careful attention is needed to (i) confounding between batch and biology and (ii) handling missing values.
A critical prerequisite is avoiding (or explicitly modeling) confounding: if a biological group is processed only within a single batch, batch correction may inadvertently remove true biology.
4.5 Hidden Batch Factors: SVA and RUV
In many projects, batch structure is not fully known or fully recorded. Multiple technical factors (operator, column, temperature, software version, etc.) may overlap and manifest as latent variables; even when batch labels exist, they may not capture all systematic variation. In such settings, SVA/RUV methods estimate latent factors to absorb unobserved technical variation.
SVA (Surrogate Variable Analysis) specifies a biological design matrix (e.g., groups and covariates), estimates surrogate variables from residual structure, and incorporates them into the model to reduce the impact of unobserved technical variation on differential analysis. The RUV family typically relies on auxiliary information such as negative controls or pseudo-replicates. In proteomics benchmarking studies, RUV-III-C has been included in comparisons and has shown competitive performance in certain scenarios.
4.6 Built-in normalization in quantification algorithms (MaxLFQ and DIA software)
Many quantification pipelines include internal normalization steps. MaxLFQ improves label-free protein quantification by using pairwise peptide ratios and delayed normalization. DIA pipelines (e.g., DIA-NN) often apply cross-run normalization and interference correction. Before applying additional normalization, confirm whether the exported intensity matrix is raw, partially normalized, or fully normalized to avoid double correction.
Comparison of commonly used data-processing methods
|
Method |
Data pattern it targets (what you see) |
Best used when |
Pros |
Cons / caveats |
|
Median scaling |
Samples show global intensity shifts; distributions aligned but shifted |
Default first-pass normalization for most proteomics matrices |
Simple, robust, easy to explain |
Can dampen true global biology (e.g., widespread up/down shifts) |
|
Total ion / sum scaling (TIC) |
Total signal differs across runs; a few high signals may dominate |
Quick adjustment when no heavy contamination/outliers |
Very fast, intuitive |
Sensitive to outliers and high-abundance features; can mis-scale |
|
Quantile normalization |
Same shape expected across samples but observed distributions differ |
When you believe overall distribution should be comparable |
Strong distribution alignment |
High risk of over-normalization; may erase real global changes |
|
LOESS drift correction (run-order based) |
Intensity drifts with injection order (up/down/curved trend) |
Long sequences with temporal drift; ideally with frequent QC |
Handles non-linear drift well |
Without reliable QC, can “correct away” true biology |
|
QC-based LOESS / QC-RLSC |
QC signals show smooth drift over time; samples track QC trend |
Large cohorts with pooled QC inserted regularly |
Drift correction anchored to QC (safer) |
Requires stable, representative QC and sufficient QC frequency |
|
VSN (variance-stabilizing normalization) |
Heteroscedasticity: low-intensity features are noisier; mean–variance trend |
When low-abundance noise dominates; for downstream linear modeling |
Stabilizes variance; improves comparability across intensity range |
Can be unstable with extreme sparsity; avoid stacking with aggressive distribution forcing |
|
Cyclic LOESS (intensity-dependent) |
MA plots show curved bias; bias differs by intensity range |
When intensity-dependent non-linear bias is evident |
Corrects non-linear, intensity-dependent bias |
If biology drives MA curvature, may remove true effects; validate carefully |
|
ComBat (Empirical Bayes) |
Clear batch clusters (date/instrument/lab); shifts in mean/variance by batch |
Known batch labels and (ideally) balanced design across groups |
Strong batch removal; widely used |
Confounded design (group≈batch) risks signal loss or artifacts |
|
limma removeBatchEffect |
Batch drives PCA/heatmaps; need quick de-batching for visualization |
Primarily for exploratory plots / PCA cleanup |
Simple, fast, practical |
Not a full statistical framework for final inference by itself |
|
SVA (surrogate variable analysis) |
Unexplained structure remains after known covariates; hidden technical factors |
Batch unknown/complex; you have a defined biological design to preserve |
Captures latent unwanted variation |
Choosing too many factors can over-correct; confounding is dangerous |
|
RUV / RUV-III-C |
Hidden variation evident; you also have controls/replicates/QC to anchor factors |
When negative controls, spike-ins, QC repeats, or technical replicates exist |
More grounded than SVA when controls are good |
Success depends on control quality; poor controls → wrong correction |
5. How to Validate Batch Correction (Post-correction QC Checklist)
Regardless of the method, validation is essential. A practical checklist includes:
QC metrics: peptide/protein IDs, total signal, RT stability, iRT performance, and missing-value rates.
Exploratory plots: PCA/UMAP should reduce batch-driven clustering while preserving biological separation.
RLE (relative log expression) or MA plots: assess whether residual intensity-dependent bias remains.
Replicates: technical/biological replicates should become more similar after correction (lower CV) without losing expected contrasts.
Negative controls: proteins expected not to change should remain stable; known markers should retain expected trends.
6. Summary: A Robust Workflow for Batch-Effect-Resistant Proteomics
In the era of high-throughput proteomics, normalization is not a routine preprocessing formality but a core quality gate that strongly influences biological conclusions. Different approaches target different artifacts: global scaling aligns overall levels, variance stabilization addresses intensity-dependent noise, QC-based methods correct time drift, and statistical models remove batch structure using known or latent factors. A well-chosen and well-validated strategy minimizes technical noise while preserving biological signal, enabling more reliable discovery and interpretation.
References
1. Čuklina J, Lee CH, Williams EG, Sajic T, Collins BC, Rodríguez Martínez M, Sharma VS, Wendt F, Goetze S, Keele GR, Wollscheid B, Aebersold R, Pedrioli PGA. Diagnostics and correction of batch effects in large-scale proteomic studies: a tutorial. Mol Syst Biol. 2021 Aug;17(8):e10240. doi: 10.15252/msb.202110240.
2. Bian Y, Zheng R, Bayer FP, Wong C, Chang YC, Meng C, Zolg DP, Reinecke M, Zecha J, Wiechmann S, Heinzlmeir S, Scherr J, Hemmer B, Baynham M, Gingras AC, Boychenko O, Kuster B. Robust, reproducible and quantitative analysis of thousands of proteomes by micro-flow LC-MS/MS. Nat Commun. 2020 Jan 9;11(1):157. doi: 10.1038/s41467-019-13973-x.
3. Čuklina J, Pedrioli PGA, Aebersold R. Review of Batch Effects Prevention, Diagnostics, and Correction Approaches. Methods Mol Biol. 2020;2051:373-387. doi: 10.1007/978-1-4939-9744-2_16.
4. Huber W, von Heydebreck A, Sültmann H, Poustka A, Vingron M. Variance stabilization applied to microarray data calibration and to the quantification of differential expression. Bioinformatics. 2002;18(Suppl 1):S96–S104.
5. Chen Q, Cao Z, Liu Y, Zhang N, Xie Y, Chen H, Mai Y, Duan S, Li J, Yu Y, Zhao Y, Shi L, Zheng Y. Protein-level batch-effect correction enhances robustness in MS-based proteomics. Nat Commun. 2025 Nov 4;16(1):9735. doi: 10.1038/s41467-025-64718-y.
Read more:
Next-Generation Omics Solutions:
Proteomics & Metabolomics
Ready to get started? Submit your inquiry or contact us at support-global@metwarebio.com.


