Spatial Metabolomics Ion Mode Selection: Positive vs. Negative Polarity in MALDI-MSI and DESI-MSI
Spatial metabolomics leverages mass spectrometry imaging (MSI) to visualize where metabolites reside within tissue sections, linking molecular information to histological context. Because metabolite classes differ widely in ionization behavior, the choice of ionization polarity—positive mode, negative mode, or both—is a pivotal step that can markedly affect chemical coverage, signal strength, and confidence in biological interpretation. In well-designed spatial metabolomics workflows, polarity selection is typically driven by metabolite chemistry (acidic versus basic functional groups), the ionization platform and its matrix/solvent environment, and the specific biological question being addressed. This article aims to provide a practical framework for deciding when to prioritize positive mode, negative mode, or complementary dual-mode acquisition.
Fundamentals of Positive vs. Negative Ion Modes in Spatial Metabolomics
In spatial metabolomics, ionization refers to metabolites gaining or losing charge in the ion source to generate detectable ions. The source of charge differs between positive and negative modes.
- Positive Ion Mode: In positive mode, metabolites typically acquire a proton (H⁺) or form cation adducts (e.g., Na⁺, K⁺), generating ions such as [M+H]⁺ and [M+Na]⁺. Ionization efficiency is strongly influenced by basic sites in the molecule (e.g., amino, imino, or quaternary ammonium groups). In general, stronger basicity supports more efficient protonation and higher signal intensity.
- Negative Ion Mode: In negative mode, metabolites typically lose a proton (H⁺) or form anion adducts (e.g., Cl⁻), generating ions such as [M–H]⁻ and [M+Cl]⁻. Ionization efficiency depends on acidic functionalities (e.g., carboxyl, phosphate, sulfonate, or phenolic hydroxyl groups). In general, stronger acidity supports more efficient deprotonation and higher signal intensity.
Ion Mode Selection in MSI Platforms: MALDI-MSI vs. DESI-MSI
MALDI-MSI and DESI-MSI are two of the most commonly used mass spectrometry imaging (MSI) platforms for spatial metabolomics, and both can be run in positive or negative ion mode. In practice, polarity selection is not an isolated setting—it should be considered together with the matrix (for MALDI-MSI) or spray solvent system (for DESI-MSI), because these conditions strongly influence ion formation and metabolite coverage. At the same time, spatial-resolution requirements also matter: some modes and chemistries tend to produce cleaner signals and sharper images under high-resolution settings. For this reason, polarity choice is typically made alongside matrix/solvent selection and the resolution goal of the experiment.
MALDI-MSI: Matrix-Driven Polarity Selection
MALDI is a soft-ionization technique in which the matrix absorbs laser energy and transfers it to analytes, enabling desorption and ion formation from metabolite molecules. Because matrix chemistry strongly influences ion formation, MALDI mode selection is closely linked to matrix choice. MALDI is well suited to both biomacromolecules and small-molecule metabolites (Choi H., 2019).
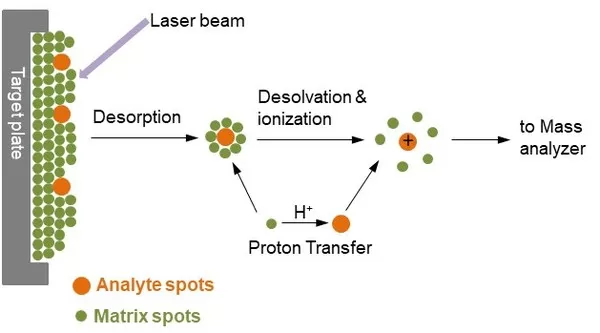
Mechanism of matrix assisted laser desorption ionization process (Veeravalli et al., 2019)
- Positive ion mode: Acidic matrices (e.g., 2,5-dihydroxybenzoic acid) are commonly used. These matrices facilitate protonation, making positive mode well suited for amino acids, nucleotides, bile acids, organic acids, hormones, and related compounds.
- Negative ion mode: Neutral or weakly basic matrices (e.g., 9-aminoacridine) are typically preferred. They promote deprotonation and are often effective for bile acids, amino acids, free fatty acids, and related compounds.
In practice, matrix diffusion effects are often smaller in positive ion mode, so the achievable spatial resolution in positive mode is frequently better than in negative mode. When high-resolution imaging at ≤10 μm is required, positive mode is commonly prioritized, with negative mode used as a complementary measurement.
DESI-MSI: Solvent/Additive-Driven Polarity Selection
DESI is an ambient-pressure soft-ionization method in which charged droplets impact the sample surface, extracting, desorbing, and ionizing metabolites. As a result, spray-solvent composition is a primary determinant of ionization performance (Pól J., 2010).
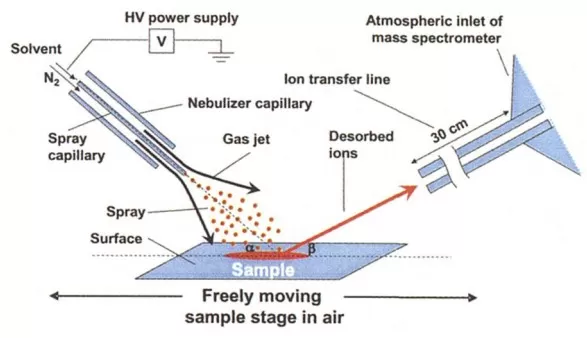
DESI-MSI principle (Takáts Z., 2004)
- Positive ion mode: A commonly used spray solvent is “methanol–water (7:3) + 0.1% formic acid.” Formic acid enhances protonation efficiency, supporting detection of amino acids, organic acids, hormones, and related compounds.
- Negative ion mode: A commonly used spray solvent is “methanol–water (7:3) + 0.1% ammonia.” Ammonia promotes deprotonation and is often suitable for fatty acids, phospholipids, sulfonated compounds, and related species.
MALDI-MSI Ionization Mechanisms
MALDI ion formation has been explained through several complementary frameworks. Commonly discussed models include photoionization, the “lucky survivor” (cluster ionization) model, and the thermal proton transfer model. Additional derivative and integrated concepts—such as exciton migration/excited-state coupling and energy-transfer induced disproportionation—help interpret the complexity of matrix-assisted laser desorption/ionization (Knochenmuss R., 2010).
Photoionization Model (Electron Excitation–Dominated)
This model proposes that matrix molecules absorb laser photons and enter an electronically excited state (S₁). Through excited-state interactions—such as pooling (S₁+S₁→S₀+Sₙ) and ion-pair formation (S₁+Sₙ→M⁺+M⁻)—matrix ions are generated. Analytes are then ionized via gas-phase proton or electron transfer. The model highlights electronic excitation and non-thermal pathways, is often discussed in the context of UV-MALDI matrices (e.g., DHB), and can explain efficient ionization under relatively low laser energy. It is also conceptually linked to exciton migration and excited-state pooling mechanisms.
Lucky Survivor Model (Cluster Ionization Mechanism)
The lucky survivor model suggests that analytes may exist as preformed ions within the matrix crystal. Laser ablation produces charged clusters containing both matrix and analyte. During cluster fragmentation, most ions undergo recombination and neutralization, while a small subset avoids recombination and is detected as the surviving ion signal. This model emphasizes the roles of preformed ions, charge separation during cluster breakup, and signal enhancement effects observed with salt additives. Mechanistically, it shares similarities with ion evaporation concepts discussed for ESI.
Thermal Proton Transfer Model
This model emphasizes thermal processes, proposing that laser energy induces localized matrix melting and that elevated temperature promotes thermally driven proton transfer between matrix and analyte, producing ions such as [M+H]⁺. Within this framework, ion yield depends on matrix melting behavior and proton-donor capacity. The model is often considered particularly relevant to IR-MALDI, where energy absorption is largely converted to heat, and it provides a rationale for how melting point and proton affinity influence ionization efficiency.
Derivative and Integrated Theories
i. Exciton migration/excited-state pooling mechanism: Excitons migrate through matrix crystals and can generate higher-energy states via biexciton pooling, facilitating ion-pair formation. This mechanism strengthens the plausibility of photoionization, especially in matrices with strong intermolecular interactions.
ii. Energy-transfer induced disproportionation (ETID) model: Laser energy can drive disproportionation reactions in matrix–analyte complexes, generating paired positive and negative ions. This model has been used to rationalize symmetric positive/negative ion signals observed in MALDI of biomacromolecules.
iii. Surface/interface ionization model: This concept emphasizes interfacial electron-transfer processes at the matrix–analyte boundary, such as electron injection from an excited matrix into the analyte. It helps explain efficient ionization of surface-associated analytes and complements photoionization by focusing on surface processes.
Overall, these models are not mutually exclusive. Instead, they describe MALDI ion formation from different angles—photochemical pathways, cluster fragmentation behavior, and thermally driven proton exchange—and together provide a more complete mechanistic context for matrix selection, parameter optimization, and signal enhancement.
Metabolite Class Preferences for Positive and Negative Ion Modes
Because metabolite classes differ in dominant functional groups, their preferred ionization modes and adduct types often differ accordingly. Representative detection forms are summarized below.
|
Substance class |
Functional group |
Representative compounds |
Mode |
Adduct ion |
|
Quaternary ammonium |
N⁺-R4 |
Phosphatidylcholine (PC) / acylcarnitines (CAR) |
Positive |
M⁺ / M+H |
|
Benzopyranone class |
Benzopyranone core |
Positive |
M⁺ |
|
|
Fatty amines |
R-NH, R-NH-R, N-R3 |
Polyamines / alkaloids |
Positive |
M+H |
|
Aldehyde/ketone/lipid class |
R-CHO, R=O-R, R-COOR |
Small aldehyde/ketone/ester metabolites |
Positive |
M+H |
|
Polyhydroxyl |
Glucose / glycosides |
Positive |
M+Na |
|
|
Phenols |
Phenolic hydroxyl |
Polyphenols / phenolic acids |
Negative |
M–H |
|
Carboxylic acids |
R-COOH |
Negative |
M–H |
|
|
Sulfonic acids |
R-SO3H |
Sulfatides (ST) |
Negative |
M–H |
|
Phosphates |
R-PO4³⁻ |
Negative |
M–H |
Practical Ion Mode Decisions in Spatial Metabolomics Studies
Polarity selection in spatial metabolomics becomes most intuitive when it is tied to a concrete biological question, because different pathways and metabolite classes show clear, chemistry-driven preferences for positive or negative mode. The scenarios below translate the general ionization principles into practical choices that help maximize metabolite coverage while keeping images interpretable and reproducible.
1. Spatial Metabolomics Focused on Lipid Metabolism
Lipid targets in spatial metabolomics commonly include phospholipids, acylcarnitines, fatty acids, sphingolipids, and related subgroups. From a functional-group perspective, they can be grouped into five categories: (1) quaternary ammonium lipids (e.g., phosphatidylcholine, PC, and acylcarnitines, CAR, containing the N⁺-R₄ group); (2) phosphate lipids (phospholipids other than PC such as PE/PG, containing the R-PO₄³⁻ group); (3) carboxylic-acid lipids (free fatty acids and related acyl species, containing R-COOH); (4) sulfonic-acid lipids (e.g., sulfatides, ST, and related sphingolipids, containing R-SO₃H); and (5) ester lipids (e.g., triacylglycerols and wax esters, containing R-COOR).
Quaternary ammonium lipids and ester lipids (such as PC and triacylglycerols) are commonly measured in positive ion mode using DHB as the matrix. In positive mode, quaternary ammonium compounds readily form stable M⁺ or M+H⁺ ions, while ester lipids typically yield M+H⁺ adducts, which are often the most informative detection forms for these classes.
By contrast, phosphate lipids, carboxylic-acid lipids, and sulfonic-acid lipids are often best targeted in negative ion mode, where 9-AA can be used as the matrix. These classes tend to form stable M–H⁻ ions through phosphate, carboxyl, or sulfonate groups, and negative mode often yields fewer interfering peaks and higher sensitivity than positive mode for these analytes.
2. Spatial Metabolomics Focused on Energy Metabolism
Energy metabolism pathways include glycolysis, the tricarboxylic acid (TCA) cycle, and the pentose phosphate pathway. Spatial metabolomics can be applied to investigate, for example, differential glycolysis in tumors versus normal tissue, metabolic remodeling in cardiac tissue, or energy metabolism in plant photosynthetic and respiratory tissues.
Key intermediates and substrates in energy metabolism can be grouped into four functional categories: (1) sugars (glucose, fructose, glycogen, and glycosides, enriched in hydroxyl groups); (2) carboxylic acids (TCA intermediates such as citrate, lactate, α-ketoglutarate, and succinate, containing R-COOH); (3) aldehydes/ketones (glycolytic intermediates such as pyruvate and acetaldehyde, containing carbonyl functionality); and (4) phosphates (phosphorylated intermediates such as ATP/ADP/AMP and sugar phosphates, containing R-PO₄³⁻).
Sugars and aldehyde/ketone compounds are often well suited to positive ion mode with DHB. Sugars frequently form stable M+Na⁺ adducts in positive mode, whereas efficient adduct formation can be challenging in negative mode. Aldehyde/ketone compounds commonly yield M+H⁺ ions in positive mode, supporting reliable detection.
Carboxylic acids and phosphates are often better captured in negative ion mode with 9-AA. TCA organic acids readily form M–H⁻ ions via carboxyl groups, while phosphorylated metabolites such as ATP/ADP/AMP commonly generate stable M–H⁻ ions via phosphate groups. These metabolites are central markers of energy metabolism and are often most efficiently detected in negative mode.
3. Spatial Metabolomics Focused on Plant Hormones
Spatial metabolomics of plant hormones can be used to characterize spatial accumulation, transport, and response patterns across tissues—for example, the polar distribution of auxin in root and shoot apices, regional variation in abscisic acid and gibberellins during fruit development, and hormone-mediated stress responses.
Although plant hormones span diverse chemistries, many key hormones are carboxylic acids, including auxin (IAA), abscisic acid (ABA), jasmonic acid (JA), and salicylic acid (SA), all of which contain the R-COOH group. Other hormones include subclasses such as cytokinins (CTK, containing amino functionality) and gibberellins (GA, containing ketone and hydroxyl groups). In practice, no single matrix enables uniformly efficient detection across all plant hormone classes. However, DMPI (4-(N,N-dimethylamino)phenyl isocyanate) is a highly specific isocyanate derivatization reagent that can react with active functional groups such as hydroxyl and amino groups, introducing a basic conjugated structure with an N-dimethylamino group. This derivatization strategy can broaden coverage across major hormone classes and is particularly compatible with MALDI-MSI, often improving ionization efficiency and sensitivity for hormone-related targets.
Key Guidelines for Choosing Ion Modes in Spatial Metabolomics
Ion-mode selection is a central element of spatial metabolomics workflows, with direct impact on metabolite coverage, detection sensitivity, and data reliability. Practical decisions should integrate ionization technology, target metabolite classes, and research objectives. In many studies, positive and negative modes are used in a complementary manner to increase chemical coverage and improve interpretability.
For mainstream platforms, MALDI-MSI mode selection is strongly matrix dependent: positive mode typically pairs with acidic matrices and often supports higher spatial resolution, whereas negative mode commonly pairs with neutral or weakly basic matrices. DESI-MSI is more directly regulated by spray-solvent composition, where formic acid and ammonia can enhance positive- and negative-mode ionization, respectively. Scenario-specific selection is often the most effective approach: lipid studies typically benefit from polarity choice matched to lipid subclasses; energy metabolism studies frequently require cross-mode coverage across sugars, carbonyl compounds, organic acids, and phosphates; and plant hormone studies can expand detectable coverage through derivatization strategies.
In general, positive ion mode tends to favor metabolites with basic sites (e.g., amino and quaternary ammonium groups), supporting ions such as [M+H]⁺ and [M+Na]⁺. Negative ion mode tends to favor metabolites with acidic sites (e.g., carboxyl and phosphate groups), where [M–H]⁻ is often dominant and signals may show less interference and higher sensitivity.
Reference
1. Choi, H., Lee, D., Kim, Y., Nguyen, H. Q., Han, S., & Kim, J. (2019). Effects of Matrices and Additives on Multiple Charge Formation of Proteins in MALDI-MS Analysis. Journal of the American Society for Mass Spectrometry, 30(7), 1174–1178. https://doi.org/10.1007/s13361-019-02213-7
2. Pól, J., Strohalm, M., Havlíček, V., & Volný, M. (2010). Molecular mass spectrometry imaging in biomedical and life science research. Histochemistry and cell biology, 134(5), 423–443. https://doi.org/10.1007/s00418-010-0753-3
3. Takáts, Z., Wiseman, J. M., Gologan, B., & Cooks, R. G. (2004). Mass spectrometry sampling under ambient conditions with desorption electrospray ionization. Science (New York, N.Y.), 306(5695), 471–473. https://doi.org/10.1126/science.1104404
4. Knochenmuss, R. (2010). MALDI Ionization Mechanisms: An Overview. In Electrospray and MALDI Mass Spectrometry, R.B. Cole (Ed.). https://doi.org/10.1002/9780470588901.ch5
5. Veeravalli, Vijayabhaskar & Madgula, Vamsi & Srivastava, Pratima. (2019). Mass Spectrometry as a Workhorse for Preclinical Drug Discovery: Special Emphasis on Drug Metabolism and Pharmacokinetics. 10.5772/intechopen.88385.
Next-Generation Omics Solutions:
Proteomics & Metabolomics
Ready to get started? Submit your inquiry or contact us at support-global@metwarebio.com.


