Spatial Metabolomics in Liver Disease Research
Why Spatial Perspectives Matter in Liver Disease Research?
The liver is a vital organ, playing a central role in metabolism, detoxification, and immune regulation. Its intricate spatial architecture, particularly the division of function across hepatic lobules, dictates its ability to perform these multifaceted tasks. Hepatocytes, organized in distinct zones within the lobules, display diverse metabolic profiles and varying susceptibilities to damage. This zonal organization is influenced by blood flow gradients—oxygen, nutrients, and hormones exhibit significant variation across these zones, leading to functional diversity and distinct metabolic signatures [1].
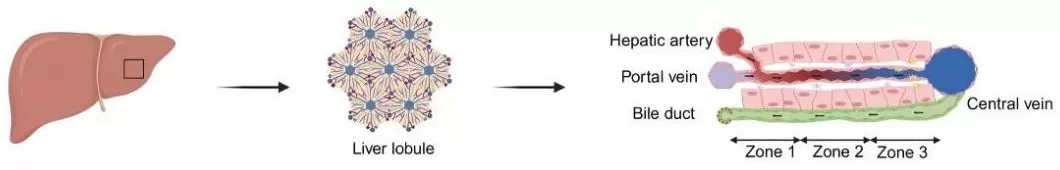
Liver anatomy and the hepatic lobule.
Image reproduced from Suo, Y., Thimme, R., & Bengsch, B, 2025, Gut, licensed under the Creative Commons Attribution License (CC BY 4.0).
Traditional omics approaches that pool tissue samples fail to capture this spatial complexity. Spatial location is paramount in liver function and disease progression, as certain diseases often display a “geographical discrimination” in their onset and development.
Spatial metabolomics, particularly technologies like matrix-assisted laser desorption/ionization mass spectrometry imaging (MALDI-MSI), enables the high-resolution visualization of metabolites within tissue sections. This innovative approach adds a spatial layer to our molecular understanding of liver pathophysiology, providing valuable insights into disease mechanisms. This blog will explore the application of spatial metabolomics through the framework of “physiological basis → injury patterns → disease progression → clinical translation”, illustrating how it is reshaping our understanding of liver diseases.
Part 1: Mapping the Liver Lobule’s Metabolic Atlas: The Physiological Foundation
Multimodal Imaging and Single-Cell Integration in Spatial Metabolomics
To understand liver disease, it’s essential to first define normal liver function. The liver’s metabolic processes vary across different zones of the hepatic lobule, with each zone performing distinct tasks. Traditional methods often overlook this spatial variability, but spatial metabolomics offers the tools to map these metabolic gradients in high detail, providing critical insights into liver physiology and disease.
1. Mass Spectrometry Imaging (MSI): Technologies like MALDI-MSI and SIMS enable the visualization of metabolites at subcellular resolution. These MSI-based approaches preserve spatial information, offering high-resolution data on how metabolites are distributed across the liver. This helps to reveal zone-specific metabolic activities that are crucial for maintaining liver function, such as gluconeogenesis in the periportal zone and detoxification in the pericentral region.
2. Integration with Single-Cell Transcriptomics: By combining spatial metabolomics with single-cell transcriptomics, metabolite distributions can be linked to specific cell types and gene expression patterns along the liver’s portal-central axis. This integration provides insights into how distinct metabolic pathways are regulated at the cellular level and how they vary across liver zones, deepening the understanding of both normal physiology and disease progression.
These advances allow for multimodal imaging, where metabolomics, lipidomics, and proteomics can be integrated into a single analysis, generating a comprehensive metabolic profile at the single-cell level.
Case Study: Metabolic Zonation in Liver Health and Disease
The liver’s functional zonation plays a key role in its metabolic activities. Periportal hepatocytes handle gluconeogenesis and β-oxidation, while pericentral hepatocytes focus on lipid synthesis and detoxification [1]. This zonal distribution is crucial for the liver’s overall function.
Spatial metabolomics plays a vital role in visualizing these metabolic zones. For instance, in alcoholic liver disease (ALD), the following observations are made:
- Lipid accumulation is notably high in the central vein region, where pericentral hepatocytes are most affected by alcohol-induced damage [2].
- Antioxidant depletion (e.g., glutathione) is observed in the periportal region, which reduces the liver's ability to combat oxidative stress, contributing to ALD’s progression [2].
By mapping these spatial differences, spatial metabolomics uncovers how zonal metabolic changes drive liver disease, offering valuable insights into disease mechanisms and potential therapeutic targets.
Spatial metabolomics reveals that the centrilobular region of the liver is especially vulnerable in ALD due to its combination of high lipid synthesis and low antioxidant capacity. This insight is crucial for understanding how diseases like NAFLD and cirrhosis progress, as similar zonal disruptions are common across various liver conditions.
In conclusion, spatial metabolomics allows for a highly detailed, spatially resolved view of liver metabolism, transforming the understanding of liver diseases by capturing how spatial variability impacts disease initiation and progression.
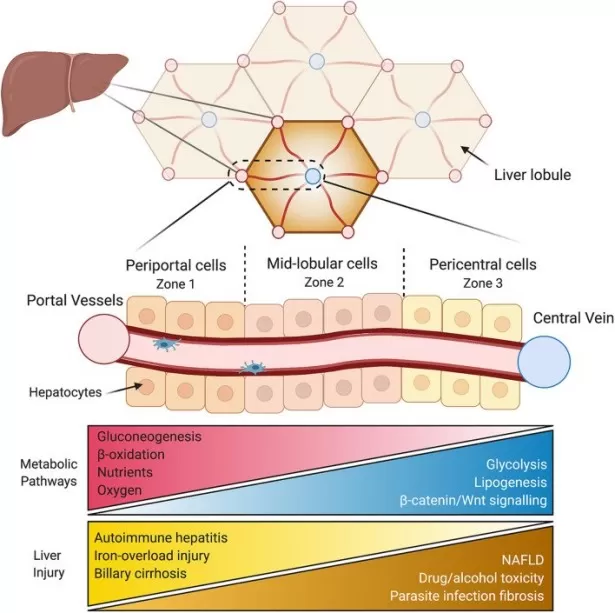
Spatial Zonation and Metabolic Pathways in the Liver Lobule.
Image reproduced from Cunningham, R. P., Porat-Shliom, N., 2021, Frontiers in physiology, licensed under the Creative Commons Attribution License (CC BY 4.0).
Part 2: Decoding Zonal Sensitivity in Liver Injury
Spatial Metabolomics for Profiling Liver Injury
Spatial metabolomics provides a powerful method for studying how toxins target specific liver regions. By comparing metabolic profiles across different injury models, it identifies shared molecular signatures and injury-specific biomarkers, highlighting zonal sensitivity in liver damage.
1. Cross-Model Comparison: Injury models (e.g., ethanol, acetaminophen, thioacetamide) are analyzed using mass spectrometry imaging (MSI) to map metabolite distributions across liver zones. This helps reveal zonal metabolic alterations and the specific effects of different toxins on liver regions, deepening the understanding of regional susceptibility.
2. Spatial lipidomics and Phospholipid Profiling: Lipidomics identifies regional lipid remodeling, focusing on phospholipids like lysophosphatidylcholines (LysoPC) and lysophosphatidylethanolamines (LysoPE). Their spatial distributions help distinguish pathological mechanisms between alcoholic liver disease (ALD) and non-alcoholic fatty liver disease (NAFLD), offering insights into universal injury mechanisms across these conditions.
Case Study: Zonal Lipid Remodeling in Liver Injury Models
A zebrafish model study examining three liver injury types (ethanol, acetaminophen, thioacetamide) provides a clear example of how spatial metabolomics can uncover common injury patterns [2]:
- Universal Injury Markers: Across all injury models, there was significant downregulation of LysoPC 16:0 and LysoPE 18:0. This consistent reduction across injuries suggests that these lipids could serve as universal biomarkers of hepatocyte damage, regardless of the injury type.
- Zonal Lipid Remodeling: Phosphatidylcholine (PC 34:1) was notably upregulated specifically in the central vein region of the liver. This finding indicates that, while the liver may experience global stress due to injury, the central vein zone is particularly involved in lipid remodeling as part of an adaptive response to injury.
- Distinct Signatures for ALD and NAFLD: Further analysis revealed that specific metabolites, such as fatty acids, glutamine, and malic acid, exhibited contrasting changes between ALD and NAFLD models, providing spatial signatures that could distinguish between these two types of liver diseases.
These findings underscore the importance of zonal sensitivity in liver injury, with the centrilobular region (central vein) being particularly vulnerable due to its role in lipid metabolism and its heightened sensitivity to oxidative stress. Spatial metabolomics reveals how this region functions as a focal point for both injury and repair mechanisms, highlighting the need for targeted therapeutic strategies.
Part 3: Spatial Insights Into Major Liver Diseases and Disease Progression
Precision Subtyping and Inflammation in Fatty Liver Disease Progression
Metabolic dysfunction-associated steatotic liver disease (MASLD) and metabolic dysfunction-associated steatohepatitis (MASH) are increasingly recognized as major liver diseases that are characterized by complex spatial interactions between hepatocytes, immune cells, and other cellular components. These diseases involve both metabolic alterations in hepatocytes and immune responses that are spatially organized within the liver. Understanding these interactions at a spatial level is crucial for identifying specific disease mechanisms and targeting the metabolic-inflammatory microenvironment for therapeutic interventions.
Recent studies using spatial multimodal analysis (SMA) have combined spatial metabolomics with spatial transcriptomics to capture the intricate spatial heterogeneity of these liver diseases. Spatial metabolomics, through techniques like mass spectrometry imaging (MSI), allows researchers to map the distribution of metabolites within liver tissue at high resolution. This provides crucial insights into how different metabolic zones and immune cell interactions contribute to disease progression, highlighting the need for precision subtyping in fatty liver diseases.
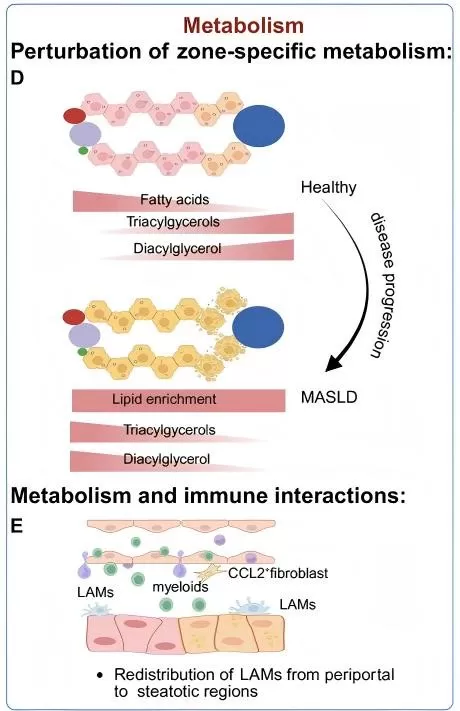
Disruption of Zone-Specific Metabolism and Immune Interactions in Liver Disease Progression.
Image reproduced from Suo, Y., Thimme, R., & Bengsch, B, 2025, Gut, licensed under the Creative Commons Attribution License (CC BY 4.0).
Case Study: Lipid-Associated Macrophages as Key Players in MASH Progression
In MASH, the spatial dynamics between hepatocytes and immune cells, particularly lipid-associated macrophages (LAMs), play a central role in driving inflammation and liver damage. Spatial metabolomics has been used to map these interactions in detail, uncovering key insights into the disease mechanisms [3]:
- Phospholipid Accumulation in “Hotspots”:
One of the major findings from spatial metabolomics in MASH is the accumulation of specific phospholipids, such as sphingomyelins, in “hotspots” where lipid-associated macrophages (LAMs) are concentrated. These phospholipids are crucial components of cell membranes and are involved in lipid remodeling during liver injury. The spatial mapping of these lipids within the liver lesions revealed that their accumulation is closely associated with the presence of LAMs, suggesting a direct link between lipid metabolism and immune cell activity in liver disease. The ability to localize these phospholipids to specific areas of the liver tissue underscores the power of spatial metabolomics in identifying region-specific metabolic shifts that are central to disease progression.
- Hepatoprotective Functions of LAMs:
In addition to their role in inflammation, LAMs have been found to secrete hepatocyte growth factor (HGF), which promotes hepatocyte proliferation and survival. This suggests that LAMs play a protective role in fatty liver disease by promoting tissue repair and reducing hepatocyte apoptosis. Spatial metabolomics has allowed researchers to visualize the spatial distribution of HGF and observe its localized effects on hepatocytes. The precise mapping of HGF distribution in relation to lipid accumulation and immune cell infiltration provides a more nuanced understanding of how spatial metabolic signals influence both disease progression and tissue repair.
Spatial metabolomics in MASH reveals the importance of the liver’s spatial organization in fatty liver diseases, identifying hotspots of lipid accumulation and immune cell interactions. It uncovers therapeutic targets, such as modulating LAMs to enhance hepatoprotective functions and targeting lipid metabolism in specific liver zones to prevent disease progression. These insights enable personalized therapies, advancing targeted interventions tailored to each patient’s metabolic-inflammatory microenvironment.
Mapping Metabolic-Immune Crosstalk in the Tumor Microenvironment of Hepatocellular Carcinoma
In hepatocellular carcinoma (HCC), spatial metabolomics plays a key role in understanding the metabolic-immune interactions within the tumor microenvironment (TME). By combining mass spectrometry imaging (MSI) with transcriptomics, it reveals how metabolic reprogramming and immune modulation support tumor progression.
- Tumor-Immune-Fibroblast Interface: Spatial metabolomics maps lactate accumulation in the TME, primarily produced by cancer-associated fibroblasts (CAFs). This lactate gradient influences tumor growth and immune cell infiltration, highlighting how CAFs reshape the local metabolic landscape to support the tumor [4].
- Immunosuppressive Effects of Lactate: The visualization of lactate-driven immune suppression shows how macrophages are polarized to an M2 phenotype, promoting immune evasion. Spatial metabolomics tracks this process in relation to immune cell locations, revealing how lactate alters immune responses within the tumor [4].
- Therapeutic Implications: By targeting lactate production or its effects on immune cells, spatial metabolomics opens new avenues for combining metabolic and immune therapies. It identifies metabolic vulnerabilities in the TME, guiding more precise, personalized treatment strategies [4].
Part 4: Clinical Translation of Spatial Metabolomics in Liver Disease Therapy
4.1 Spatial Metabolomics in Drug Toxicity and Liver Injury Mechanisms
Spatial metabolomics provides high-resolution, spatially-resolved insights into drug metabolism and toxicity. Using techniques like mass spectrometry imaging (MSI), researchers can map drug distribution and metabolite accumulation within liver tissue, revealing zonal variations in drug-induced damage. This approach helps identify early signs of toxicity and provides a deeper understanding of drug-induced liver injury (DILI) mechanisms, enabling safer drug development and more precise adverse reaction predictions.
Case Study: APAP Toxicity and Centrilobular Necrosis in Liver Injury
In acetaminophen (APAP) overdose models, spatial metabolomics has been instrumental in uncovering the zonal nature of APAP-induced liver damage. Specifically, NAPQI, the toxic metabolite of APAP, accumulates predominantly in the central vein region of the liver lobule. Through high-resolution MSI, researchers have demonstrated that NAPQI accumulation leads to glutathione depletion and results in centrilobular necrosis, a hallmark of APAP hepatotoxicity [2].
- Spatial Localization of Toxicity: By visualizing where NAPQI accumulates, spatial metabolomics allows researchers to directly link the central vein as the site of injury, confirming the previously observed zonal selectivity. This insight explains why the centrilobular region is particularly vulnerable in APAP overdose, offering potential biomarkers and strategies for early detection and intervention.
- Improved Risk Assessment: The ability to visualize the spatial distribution of toxic metabolites offers critical insights into dose-response relationships and allows for the identification of high-risk regions within the liver. This can guide better dosing regimens and liver-protective therapies for patients at risk of DILI.
4.2 Utilizing Spatial Metabolomics for Real-Time Surgical Guidance in Liver Disease
Spatial metabolomics is increasingly being integrated into real-time surgical guidance, particularly for complex procedures like liver cancer resection. MALDI-assisted imaging and other intraoperative spatial metabolomics techniques are helping surgeons navigate metabolic boundaries within liver tumors, allowing for more precise tumor removal and reducing the likelihood of recurrence [5,6].
- Mapping Metabolic Boundaries: In liver cancer surgeries, spatial metabolomics enables the mapping of metabolic signatures at the tumor margins. By visualizing distinct metabolic profiles at the edges of the tumor and surrounding healthy tissue, surgeons can clearly identify the tumor boundary, ensuring that all cancerous tissue is removed while preserving as much healthy tissue as possible.
- Real-Time Decision Making: The ability to visualize metabolic differences between tumor and normal tissue during surgery offers immediate feedback, allowing surgeons to make more informed decisions. This real-time guidance can lead to more complete resections, reducing the risk of leaving behind microscopic tumor foci that could lead to recurrence.
- Enhanced Surgical Outcomes: By using spatial metabolomics to guide surgery, the precision of tumor removal is improved, which may lead to lower recurrence rates, faster recovery times, and better overall outcomes for patients. This real-time approach also opens the door for personalized surgical strategies, where the unique metabolic profile of the patient's tumor can directly influence treatment decisions.
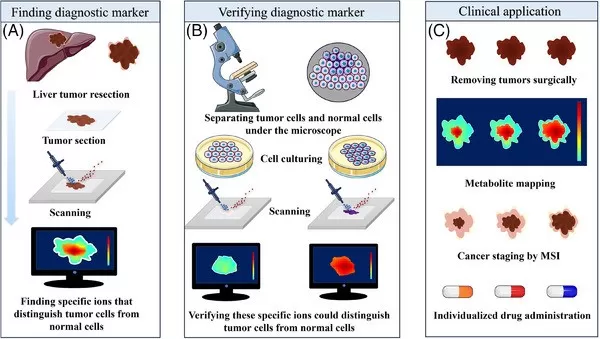
Utilizing Spatial Metabolomics for Real-Time Surgical Guidance and Tumor Diagnosis.
Image reproduced from Min, X., Zhao, Y., Yu, M. et al., 2024, Clinical and translational medicine, licensed under the Creative Commons Attribution License (CC BY 4.0).
Conclusion and Future Outlook
Spatial metabolomics is transforming liver research by moving beyond traditional molecular lists to create metabolic maps that reveal how metabolites are distributed within liver tissue. This approach enhances our understanding of zonal physiology and how metabolic gradients influence disease progression. Key breakthroughs, including subcellular resolution imaging, are allowing researchers to explore organelle-level compartments, providing deeper insights into how metabolic processes at the cellular level contribute to disease. This advancement is especially crucial in understanding liver diseases like NAFLD and hepatocellular carcinoma (HCC), where cellular metabolic heterogeneity impacts disease outcomes.
In clinical settings, spatial metabolic biomarkers have the potential to guide more accurate biopsy targeting and personalized drug regimens, driving the move toward precision medicine in hepatology. By identifying unique metabolic signatures in individual patients, spatial metabolomics can enable more tailored treatments, improving therapeutic efficacy. As the technology continues to evolve, it holds great promise for enhancing early diagnosis, disease classification, and more effective, personalized care for liver disease patients.
References:
1. Santos, A. A., Delgado, T. C., Marques, V., Ramirez-Moncayo, C., Alonso, C., Vidal-Puig, A., Hall, Z., Martínez-Chantar, M. L., & Rodrigues, C. M. P. (2024). Spatial metabolomics and its application in the liver. Hepatology (Baltimore, Md.), 79(5), 1158–1179. https://doi.org/10.1097/HEP.0000000000000341
2. Zhu, Z., Zhang, Y., Li, J., Han, Y., Wang, L., Zhang, Y., Geng, H., Zheng, Y., Wang, X., Sun, C., Li, B., & Chen, P. (2024). Mass spectrometry imaging-based metabolomics highlights spatial metabolic alterations in three types of liver injuries. Journal of pharmaceutical and biomedical analysis, 242, 116030. https://doi.org/10.1016/j.jpba.2024.116030
3. Li, Z., Luo, G., Gan, C., Zhang, H., Li, L., Zhang, X., Xing, X., Hu, S., Tan, X., Ding, J., Zhang, L., Peng, Y., Xu, Z., Pan, Q., Byrne, C. D., Targher, G., Jin, X. Z., Xie, W., Ouyang, X., Zheng, M. H., … Chai, J. (2025). Spatially resolved multi-omics of human metabolic dysfunction-associated steatotic liver disease. Nature genetics, 57(12), 3112–3125. https://doi.org/10.1038/s41588-025-02407-8
4. Chen, P., Geng, H., Ma, B., Zhang, Y., Zhu, Z., Li, M., Chen, S., Wang, X., & Sun, C. (2025). Integrating spatial omics and single-cell mass spectrometry imaging reveals tumor-host metabolic interplay in hepatocellular carcinoma. Proceedings of the National Academy of Sciences of the United States of America, 122(31), e2505789122. https://doi.org/10.1073/pnas.2505789122
5. Planque, M., Igelmann, S., Ferreira Campos, A. M., & Fendt, S. M. (2023). Spatial metabolomics principles and application to cancer research. Current opinion in chemical biology, 76, 102362. https://doi.org/10.1016/j.cbpa.2023.102362
6. Berrell, N., Sadeghirad, H., Blick, T., Bidgood, C., Leggatt, G. R., O'Byrne, K., & Kulasinghe, A. (2024). Metabolomics at the tumor microenvironment interface: Decoding cellular conversations. Medicinal research reviews, 44(3), 1121–1146. https://doi.org/10.1002/med.22010
7. Suo, Y., Thimme, R., & Bengsch, B. (2025). Spatial single-cell omics: new insights into liver diseases. Gut, gutjnl-2024-332105. Advance online publication. https://doi.org/10.1136/gutjnl-2024-332105
8. Cunningham, R. P., & Porat-Shliom, N. (2021). Liver Zonation - Revisiting Old Questions With New Technologies. Frontiers in physiology, 12, 732929. https://doi.org/10.3389/fphys.2021.732929
9. Min, X., Zhao, Y., Yu, M., Zhang, W., Jiang, X., Guo, K., Wang, X., Huang, J., Li, T., Sun, L., & He, J. (2024). Spatially resolved metabolomics: From metabolite mapping to function visualising. Clinical and translational medicine, 14(11), e70031. https://doi.org/10.1002/ctm2.70031
Next-Generation Omics Solutions:
Proteomics & Metabolomics
Ready to get started? Submit your inquiry or contact us at support-global@metwarebio.com.


