Spatial Metabolomics (MSI) Metabolite Identification: From Accurate Mass to Evidence-Integrated Annotation
Spatial metabolomics moves beyond the traditional chromatography–mass spectrometry framework of LC–MS-based bulk metabolomics, enabling direct metabolite mapping in tissue while also creating distinct statistical and chemical challenges for compound identification. Guided by widely used metabolite annotation standards in the metabolomics community (MSI confidence levels) and leveraging the added dimension of spatial context, this article outlines a practical metabolite identification workflow and decision framework based on multi-level evidence integration. In addition, we highlight LC–MS/MS-assisted identification as a complementary strategy to improve MS/MS coverage and enhance isomer discrimination in mass spectrometry imaging (MSI) studies [1] [2]. (Learn more at: Metabolite Identification in LC-MS Metabolomics)
1. Why Metabolite Identification Is Harder in Spatial Metabolomics MSI
In conventional untargeted metabolomics using LC–MS/MS, compound identification is typically built on a sequential workflow. Samples are first separated by liquid chromatography to reduce chemical complexity, followed by high-resolution MS1 acquisition to measure accurate precursor masses. Next, data-dependent acquisition (DDA) is used to collect MS/MS fragmentation spectra, which are then matched against spectral libraries and, when available, authentic reference standards. Together, chromatographic separation, accurate-mass MS1, and MS/MS library/standard matching provide the basis for confident structural confirmation.
In contrast, spatial metabolomics platforms (e.g., MALDI-MSI, DESI-MSI, SIMS) deliberately sacrifice chromatographic separation to preserve spatial context and generate metabolite distribution maps directly from tissue sections [3]. This trade-off creates three major bottlenecks:
i. Ion suppression and signal overlap (ion suppression & co-localization): On the tissue surface, hundreds to thousands of metabolites are effectively co-detected within the same microenvironment, producing strong isobaric and background interference. As a result, the m/z signal within a single pixel is rarely chemically “pure,” making isomers and isobars difficult to resolve.
ii. Sensitivity limits (single-pixel sensitivity): Ion yield at the pixel level is extremely low (often in the fmol–amol range), which substantially reduces signal-to-noise (S/N). Consequently, acquiring informative, high-quality on-tissue MS/MS spectra can be challenging, and in some cases not feasible.
iii. Data sparsity (limited fragmentation coverage): Because MSI acquisition time is constrained, deep MS/MS collection across all pixels is impractical. The resulting dataset therefore combines high-dimensional spatial information with comparatively sparse fragmentation data.
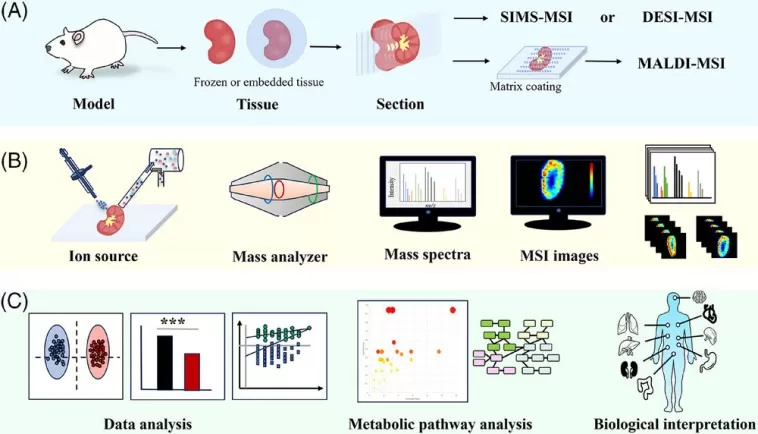
Typical workflow for spatially resolved metabolomics [2].
(A) Sample acquisition and preprocessing. (B) MSI data acquisition and visualisation are achieved through desorption and ionisation by an ion source, followed by the acquisition of MS information via a mass analyser, resulting in numerous MSI images. (C) Data processing, metabolic pathway analysis and biological interpretation.
Although spatial metabolomics does not include chromatographic separation, it adds a powerful orthogonal layer of information: spatial distribution. An ion image in mass spectrometry imaging (MSI) is more than a visual output—it encodes anatomical localization and region-specific enrichment that can support or challenge a proposed metabolite annotation. As a result, compound identification in spatial metabolomics is rarely a single-step “match,” but instead an iterative process that integrates multiple lines of evidence and converges on the most plausible assignment through evidence-weighted inference [3].
2. MSI Confidence Levels: A Four-Layer Evidence Framework for Annotation
Following the Metabolomics Standards Initiative (MSI) framework and incorporating spatial MSI characteristics, identification evidence can be organized into four layers that should be accumulated progressively [3]:
Level 4: Accurate-mass MS1 for candidate generation
Principle: High-resolution MS (e.g., Orbitrap/FT-ICR) provides accurate precursor mass (often < 3 ppm), enabling elemental composition constraints (C, H, N, O, P, S, etc.) [1].
Role: Database-assisted filtering (HMDB, METLIN, LipidMaps) removes the majority of implausible structures and generates a tractable candidate list [3].
Limitation: Accurate mass alone cannot distinguish isomers (e.g., glucose vs fructose; PC(16:0/18:1) vs PC(18:1/16:0)).
LC–MS/MS-assisted identification: To reduce isotopic ambiguity and interference in spatial MS1 features, LC–MS/MS can be performed on adjacent tissue sections or matched samples from the same batch. Chromatographic separation improves S/N and provides orthogonal chemical evidence to support candidate selection [2].
Community consensus: Annotation based solely on MS1 is typically considered MSI Level 3 or Level 4 (putative annotation) and should not be used to support detailed mechanistic claims [1].
Level 2/3: MS/MS fragmentation as the primary chemical evidence
Principle: MS/MS fragmentation provides diagnostic ions that constrain molecular class and structural motifs, forming the core chemical evidence for metabolite annotation [3].
Common strategies:
a) On-tissue MS/MS: Acquire MS/MS within a region of interest (ROI) and aggregate signals to improve S/N [3].
b) Library matching: Search public libraries (GNPS, mzCloud) and/or curated in-house MS/MS libraries [3].
c) In-silico prediction: Apply tools such as CFM-ID and MetFrag when authentic standards are unavailable [3].
LC–MS/MS-assisted validation: When single-pixel MS/MS is too weak, acquire high-quality LC–MS/MS spectra from adjacent sections or tissue extracts. These spectra can validate on-tissue assignments and improve confirmation of molecular backbones and lipid acyl-chain composition [2].
Community consensus: In practice, informative MS/MS remains the most defensible criterion for confirming a molecular class (e.g., lipid class) and moving beyond purely putative identification [3].
Spatial co-localization as supportive validation
Principle: Endogenous biomolecules often display coherent spatial distributions and anatomical specificity (e.g., restricted to cortex, localized at tumor margins) [3].
Common strategies:
a) Use spatial correlation metrics to suppress random single-pixel noise and strengthen signal robustness [3].
b) Co-register MSI ion images with histology (e.g., H&E) to test whether a metabolite aligns with tissue microanatomy [3].
LC–MS/MS assistance: Cross-reference LC–MS-annotated candidates with MSI m/z features and evaluate ROI-level consistency across spatial patterns and chemical evidence, improving overall annotation confidence through complementary validation [2].
Warning: Spatial patterns support, but cannot replace, chemical structure evidence. Contaminants can also exhibit spatial structure. Spatial evidence should be used to increase confidence and reduce false positives, not as a stand-alone identification criterion [3].
Biological plausibility as logical reinforcement
Principle: Putative annotations should be consistent with known tissue physiology and biochemical pathways [4].
Applications:
a) If a highly abundant “plant-specific alkaloid” is annotated in brain tissue, the result warrants strong skepticism even if spectral scores appear high, as contamination or ion-suppression artifacts may be responsible.
b) Pathway enrichment analysis can help prioritize biologically plausible candidates and deprioritize chemically possible but physiologically unlikely matches.
3. Practical MSI Annotation Workflow: From MS1 Features to Publishable IDs
The workflow below reflects a spatial metabolomics compound identification strategy aligned with common reporting expectations:
|
Step |
Operation |
Evidence level |
Output |
|
Step 1 |
Peak extraction and denoising |
Technical preprocessing |
A curated list of m/z features |
|
Step 2 |
MS1 matching |
Level 4 |
Candidate list (including isomers); LC–MS/MS assistance can improve candidate reliability |
|
Step 3 |
MS/MS validation |
Level 2/3 |
Supported molecular scaffold/class; LC–MS/MS adds high-coverage fragment evidence |
|
Step 4 |
Spatial pattern analysis |
Supportive evidence |
Noise suppression and biological relevance assessment |
|
Step 5 |
Pathway mapping |
Supportive evidence |
Biochemical interpretation and plausibility checks |
|
Step 6 |
Integrated rating |
MSI Level |
Publishable annotation set with stated confidence |
In practice, the workflow summarized in the table is implemented through an evidence-building process that starts with MS1 accurate-mass feature detection and database-driven candidate generation, then moves toward stronger chemical support through MS/MS-based metabolite annotation. Spatial metabolomics adds a critical layer at this stage: ion images and ROI-level consistency checks help validate whether a candidate metabolite shows anatomically meaningful localization rather than random noise. The figure below mirrors this progression, illustrating how MSI software tools and scoring systems connect MS1 screening, MS/MS confirmation, spatial pattern evaluation, and finally MSI confidence-level reporting for transparent, publishable compound identification in mass spectrometry imaging (MSI).
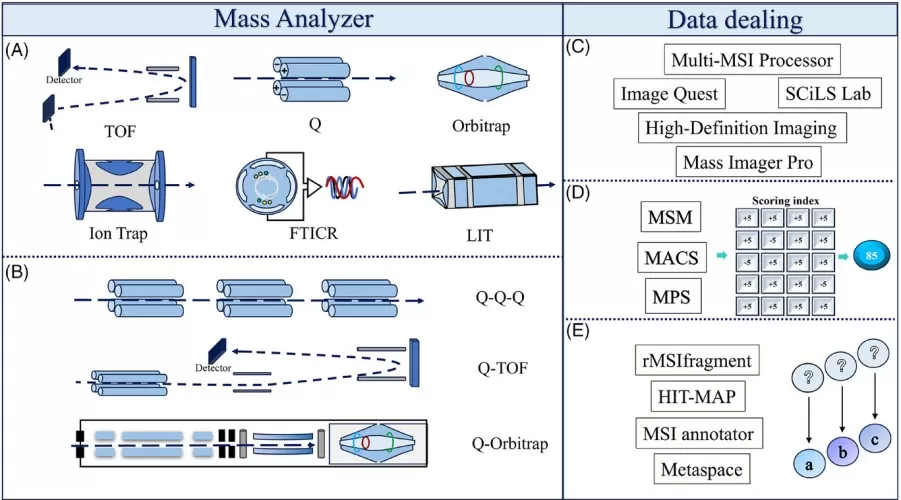
Schematic of spatial metabolomics analysis tools and data processing workflow [2].
(A) Single MS instrument types (TOF, Q, Orbitrap, Ion Trap, FTICR, LIT) are used for MS1 feature detection; (B) Tandem MS instruments (QqQ, Q-TOF, Q-Orbitrap) are used for MS/MS validation; (C) Data visualization software (Multi-MSI Processor, Image Quest, SCiLS Lab, High-Definition Imaging, Mass Imager Pro) is used to generate metabolite spatial distribution maps; (D) Metabolite annotation scoring systems (MSM, MACS, MPS) evaluate annotation credibility; (E) Automated annotation tools (rMSIfragment, HIT-MAP, MSI annotator, METASPACE) rapidly generate candidate annotations.
To show how this spatial metabolomics metabolite annotation workflow works in practice, we use the ion feature at m/z 760.5857 as an example. By combining accurate-mass MS1 screening, ROI-based MS/MS fragmentation, spatial localization patterns in MSI, and biological plausibility from tissue context, the evidence converges on a defensible putative assignment and an MSI confidence-level reportable result.
1) Goal: Annotate the ion signal at m/z 760.5857.
2) MS1 (accurate mass): The feature matches C₄₂H₇₈NO₈P, generating candidates that include multiple isomers of phosphatidylcholine (PC) or phosphatidylethanolamine (PE).
3) MS/MS (ROI-based fragmentation): ROI-aggregated MS/MS reveals the diagnostic fragment at m/z 184.07 (phosphocholine headgroup), supporting assignment to the phosphatidylcholine lipid class.
4) Spatial validation (ion image + histology): The ion image shows a coherent distribution enriched at the boundary of the tumor necrotic zone and overlapping with cell-dense regions on H&E staining, making random noise less likely.
5) Biological plausibility: This microenvironment is consistent with active phospholipid remodeling, reinforcing the plausibility of the annotation.
6) Final annotation (MSI confidence level): Putatively annotated as PC(34:1) (MSI Level 2 based on MS/MS plus spatial context). Species-level confirmation typically requires authentic standards and/or CCS evidence.
4. Common MSI Identification Bottlenecks and Emerging Technical Solutions
Even with a structured, evidence-based annotation strategy, spatial metabolomics and mass spectrometry imaging (MSI) workflows still face recurring technical bottlenecks that can limit annotation confidence and biological interpretability. In practice, the most common issues cluster around isomer discrimination, insufficient on-tissue MS/MS coverage, false positives driven by matrix effects and noise, and incomplete spectral databases for diverse metabolite classes. The table below summarizes these pain points and highlights representative frontier solutions that are increasingly used to strengthen compound identification and metabolite annotation in MSI studies [4].
|
Pain point |
Traditional challenge |
State-of-the-art solutions |
|
Isomer discrimination |
MS1 cannot resolve isomers |
Ion mobility (TIMS/TWIMS), diagnostic MS/MS fragment rules |
|
Missing MS/MS |
Single-pixel MS/MS is too weak |
ROI-aggregated acquisition; LC–MS/MS-assisted validation |
|
False positives |
Matrix effects and noise |
METASPACE FDR control, NMF (non-negative matrix factorization), deep learning–based denoising |
|
Database limitations |
Incomplete spectral coverage |
In-silico libraries (e.g., virtual metabolite libraries) and community resources such as GNPS |
5. Best Reporting Practices for Spatial Metabolomics MSI Identification
Robust compound identification in spatial metabolomics depends as much on transparent reporting as it does on instrumentation and algorithms. Because mass spectrometry imaging (MSI) often operates with limited MS/MS coverage and strong matrix effects, overstating annotation certainty can quickly undermine biological conclusions and reproducibility. The practices below reflect widely accepted best-reporting principles for metabolite annotation, helping ensure that MSI results are interpretable, comparable across studies, and suitable for publication [4].
- Do not equate image quality with structural certainty:
High-contrast or anatomically “beautiful” ion images do not prove correct identification. Figure legends and methods should explicitly state the MSI confidence level (Level 1–4) for each reported metabolite, and the evidence supporting that level.
- Treat MS1-only assignments as provisional annotations:
Accurate-mass matching alone (often MSI Level 4) is useful for candidate generation, but it is not sufficient for claiming specific structures, enzyme mechanisms, or definitive biomarkers. When only MS1 evidence is available, the text should clearly describe the result as putative and avoid over-interpretation.
- Report parameters that determine annotation outcomes:
To make MSI metabolite identification reproducible, report the database/library names and versions, precursor and fragment mass tolerances (ppm/Da), adduct assumptions, scoring thresholds, and key preprocessing steps such as denoising, normalization, and FDR control. MS/MS acquisition settings should also be explicit (DDA, DIA, or targeted; ROI-based strategies; collision energies).
- Use authentic standards for Level 1 claims whenever possible:
If a metabolite is reported as “identified” (MSI Level 1), it should be supported by matching MS/MS spectra and at least one orthogonal property under comparable conditions (e.g., retention time in LC–MS, mobility time/CCS in ion mobility). In spatial metabolomics, this commonly means validating MALDI-MSI or DESI-MSI annotations with LC–MS/MS from adjacent tissue sections or matched extracts to strengthen structural confidence.
This reporting discipline does not reduce the value of spatial metabolomics; instead, it makes MSI-based findings more credible by aligning claims with evidence, which is essential for downstream biological interpretation and translational use [4].
6. Key Takeaways: Evidence Integration for Robust MSI Metabolite Annotation
Compound identification in spatial metabolomics can be viewed as a Bayesian evidence-integration process: MS/MS provides chemical constraints, spatial distribution provides anatomical consistency, and biochemical plausibility provides pathway-level reinforcement. Together, these dimensions progressively increase the probability that a reported metabolite truly exists in the tissue context.
Rather than relying on a single dimension, robust MSI metabolite annotation requires complementary evidence. Accurate-mass MS1 provides direction, MS/MS provides structural anchors, spatial distribution provides contextual mapping, and biological logic provides interpretive control. When these elements converge, spatial metabolomics can move beyond images toward credible, mechanistically meaningful metabolite identification in complex tissues.
Reference
[1] Reisdorph N, Walmsley S, Reisdorph R. A Perspective and Framework for Developing Sample Type Specific Databases for LC/MS-Based Clinical Metabolomics. Metabolites. 2019;10(1):8. doi:10.3390/metabo10010008.
[2] Min X, Zhao Y, Yu M, Zhang W, Jiang X, Guo K, Wang X, Huang J, Li T, Sun L, He J. Spatially resolved metabolomics: From metabolite mapping to function visualising. Clin Transl Med. 2024;14(11):e70031. doi:10.1002/ctm2.70031.
[3] Ma S, Leng Y, Liu X, Meng Y, Yin Z, Hang W. High spatial resolution mass spectrometry imaging for spatial metabolomics: Advances, challenges, and future perspectives. Trends Anal Chem. 2023;159:116902. doi:10.1016/j.trac.2022.116902.
[4] Alseekh, S., Aharoni, A., Brotman, Y. et al. Mass spectrometry-based metabolomics: a guide for annotation, quantification and best reporting practices. Nat Methods 18, 747–756 (2021). https://doi.org/10.1038/s41592-021-01197-1
Next-Generation Omics Solutions:
Proteomics & Metabolomics
Ready to get started? Submit your inquiry or contact us at support-global@metwarebio.com.


