Plant Hormone Signaling Through Proteomics and Phosphoproteomics
Why “activity” matters more than “expression” in fast hormone signaling
Plant hormones reprogram growth and stress responses on a timescale that often outpaces transcription. This mismatch is one reason why researchers frequently encounter a familiar frustration: RNA‑seq shows clear differential expression, yet the phenotypic readout is driven by events that unfold within minutes. In hormone pathways, those early decisions are commonly encoded by post‑translational regulation—especially phosphorylation—together with protein turnover. Total proteomics answers a necessary but incomplete question (“which proteins change in abundance?”), whereas phosphoproteomics addresses the question that is usually rate‑limiting for mechanism (“which proteins are switched on or off, and how does the signal propagate from receptors to transcription factors?”).
This article is written for plant biologists who want to design publishable (phospho)proteomics studies and convert large tables into mechanistic stories. You will learn which hormone pathways are particularly suitable for phosphoproteomics, how to structure sampling in time‑course and perturbation experiments, how to interpret site‑level changes as kinase–substrate relationships, and how to avoid common pitfalls that cause elegant but incorrect conclusions. We close with a practical checklist for initiating a hormone signaling phosphoproteomics project, including enrichment considerations and deliverables that help move from discovery to validation.
1. Why phosphorylation is so central to plant hormone signaling
Phosphorylation is central to hormone signaling because it delivers speed, reversibility, and specificity in a single chemical event. A phosphorylation change can alter catalytic activity, create or erase docking surfaces, reshape protein–protein interactions, or redirect subcellular localization—often without any change in transcript level or protein abundance. This is particularly important in plants, where signaling must remain plastic under fluctuating environments. A clear example is drought‑associated remodeling of auxin outputs: phosphorylation of the auxin signaling repressor IAA15 by MAP kinases stabilizes the protein and suppresses root development, demonstrating how a single phosphorylation‑dependent stability shift can reweight a growth–stress trade‑off without requiring a new transcriptional program to take effect first [1].
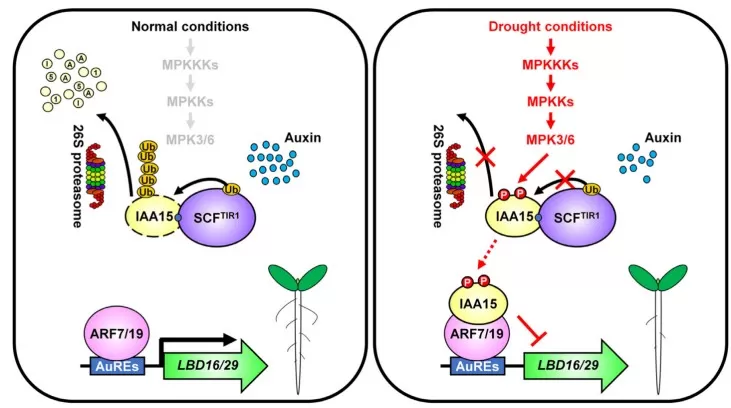
Suppression of lateral root development by the phosphorylation of IAA15
Image reproduced from Kim et al., 2022, Nucleic Acids Res, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
At a systems level, phosphorylation behaves like a molecular “code” that can integrate multiple cues. Distinct combinations of phosphosites on the same protein can encode different inputs and route signals toward different outcomes, explaining why hormone pathways show context‑dependent behavior rather than rigid linearity [2]. Stomatal closure in abscisic acid (ABA) signaling illustrates why this matters for physiology: ABA responses require rapid tuning of ion transport and guard cell behavior, and these processes are governed by phosphorylation cascades rather than delayed changes in gene expression. MAP4K1 and MAP4K2 exemplify this logic by functioning in ABA‑induced and calcium‑mediated stomatal closure, tying site‑level phosphorylation events to a well‑defined, fast phenotypic readout [3].
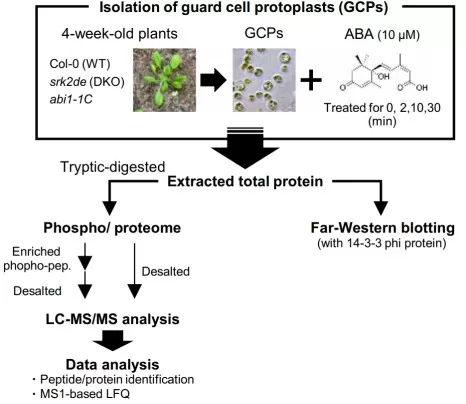
Phosphoproteomic analysis using Arabidopsis guard cells
Image reproduced from Yamashita et al., 2025, Sci Adv, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
2. Proteomics vs. phosphoproteomics: what each layer can and cannot answer
Quantitative proteomics is the right tool when the mechanism depends on protein abundance, complex composition, or large‑scale remodeling of cellular machinery. Hormones frequently modulate translation, trafficking, and degradation; therefore, changes in transporter abundance, chaperone systems, or proteostasis components can be mechanistically informative, particularly in long treatments or during developmental transitions. However, abundance is often a lagging indicator in signaling: many core regulators remain present at near‑constant levels while their functional state changes rapidly through phosphorylation or other post‑translational modifications.
Phosphoproteomics complements proteomics by quantifying site‑specific phosphorylation (typically Ser/Thr/Tyr) to reveal activity states and infer the directionality of signal flow. This distinction becomes obvious in large atlases. A tissue‑resolved phosphoproteomic landscape of rice reported extensive phosphorylation variation across organs, capturing signaling differences that cannot be inferred reliably from protein abundance alone [4]. When the biological question is “which nodes are activated, and when,” phosphoproteomics is therefore the primary readout, with proteomics serving as an essential control layer to distinguish true activation from abundance‑driven confounding. Finally, the practical reason these layers must be analyzed together is analytical: phosphorylation often occurs at low stoichiometry, so phosphopeptides represent a small fraction of the peptide pool. Reviews of end‑to‑end workflows emphasize that enrichment, quality control, and consistent processing are not optional details; they are the determinants of whether signaling events are observed at all and whether quantitative comparisons across conditions are interpretable [5].
3. Four experimental designs that reliably capture hormone phosphosignaling
Because hormone phosphosignaling is fast and transient, experimental design is not a downstream convenience—it is the difference between seeing the signal and missing it. Time‑course sampling is usually the most information‑dense design, because it separates early signaling events from later adaptation. The key is to include very early points (for example 0/5/15/30/60 minutes) and to treat biological replication as a biological requirement, not merely a statistical one. A universal plant phosphoproteomics workflow applied to tomato cold stress demonstrates why: the authors could resolve dynamic signaling phases across time, revealing phosphorylation patterns that would have collapsed into noise if measured only at a single late time point [6].
Dose‑response designs are most valuable when you suspect threshold behavior, such as a switch‑like activation of a pathway that only appears above a critical hormone concentration. Genetic perturbation designs—wild type versus receptor or kinase mutants, or overexpression lines—are the most persuasive route to causality, because the disappearance of a phosphosite response in a mutant background strongly suggests it is downstream rather than merely correlated. Auxin perception through ABP1–TMK signaling provides an example where phosphorylation profiling was used to connect perception modules to downstream phosphorylation programs and to auxin canalization outcomes [7]. Finally, stress × hormone “crosstalk” designs are essential when signals are integrated in real environments. In infection‑related development of the rice blast fungus, phosphorylation landscapes captured state transitions and pathway rewiring, illustrating how phosphoproteomics can reveal integrated decision points that are invisible to single‑factor designs [8].
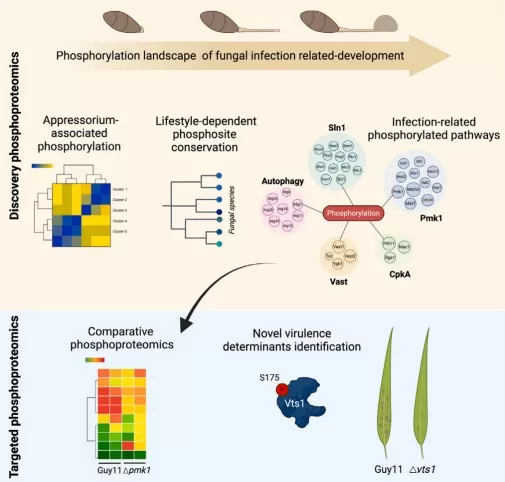
The phosphorylation landscape of infection-relateddevelopment
Image reproduced from Cruz-Mireles et al., 2024, Cell, licensed under the Creative Commons CC-BY license (CC BY).
4. Turning phosphosites into mechanisms: a layered interpretation strategy
A phosphoproteomics dataset becomes mechanistic when it is organized into layers that progressively increase explanatory power. The global layer asks what changes overall: which phosphosites and proteins are significantly regulated, which pathways are enriched, and which motif patterns suggest kinase families that are engaged. This layer is where proteome and phosphoproteome should be interpreted together; if a protein’s abundance is stable but its phosphorylation changes sharply, you have a strong candidate for activity‑based regulation rather than expression‑based regulation.
The second layer is modular: map regulated phosphosites onto canonical signaling chains (receptor → kinase/phosphatase → transcription factor) and ask where the response is entering, where it branches, and which steps appear rate‑limiting. The third layer is network‑based: integrate regulated sites with known protein interactions and kinase–substrate knowledge to prioritize directional hypotheses that can be tested. Databases that connect plant phosphorylation sites to protein networks, such as P³DB 3.0, are valuable at this stage because they support practical tasks—annotating known sites, finding conserved motifs, and projecting candidates into interaction neighborhoods that suggest functional experiments [9].
Across all layers, the central interpretive principle is to avoid treating every regulated site as equally meaningful. Mechanism‑level claims require high localization confidence, consistent regulation across replicates, and biological plausibility (for example, sites in regulatory regions or near interaction motifs). This discipline is what converts a “list of sites” into a coherent receptor‑to‑transcription factor narrative.
5. What phosphoproteomics enables in practice: three representative application scenarios
The first practical value of phosphoproteomics is discovering regulatory nodes that connect signaling to physiology. For drought tolerance in tomato, phosphorylation of the sugar transporter TST2 by the protein kinase CPK27 was shown to enhance drought tolerance, linking a specific kinase–substrate phosphorylation event to metabolite partitioning and stress performance [10]. This kind of result is difficult to obtain from transcriptomics alone because the decisive layer is protein function and transport activity, not gene expression.
A second value is translating molecular regulation into spatial and tissue logic. Hormone signaling is often localized—gradients, tissue boundaries, or infection fronts matter as much as bulk averages. Imaging approaches that map the spatial distribution of structurally diverse plant hormones illustrate why integrating signaling with spatial context can change interpretation: a phosphorylation‑defined “activated pathway” becomes far more actionable when you know where the hormone accumulates and where downstream responses must be executed [11].
A third value is decoding immune and stress pathway rewiring into actionable intervention points. In rice immunity, release of a “ubiquitin brake” activates OsCERK1‑triggered responses, highlighting how post‑translational regulation can gate receptor‑proximal immunity and rapidly shift the signaling landscape [12]. Together, these examples show the unique contribution of phosphoproteomics: it identifies switching events and regulatory bottlenecks that are closest to causal control, providing sharper hypotheses for validation and engineering.
6. Why enrichment and sample handling determine whether you “see” signaling at all
Most failures in phosphoproteomics are not caused by the mass spectrometer—they are caused upstream, during sample collection, processing, and phosphopeptide enrichment. The reason is stoichiometry: phosphopeptides are typically far less abundant than their non‑phosphorylated counterparts, and they are easily masked in complex plant samples that contain highly abundant proteins and interfering secondary metabolites. If you skip enrichment, you usually obtain a dataset biased toward the most abundant proteins rather than the most informative signaling events.
Modern phosphopeptide enrichment strategies—commonly based on metal oxide affinity (such as TiO₂) or immobilized metal affinity chromatography (IMAC)—increase the relative abundance of phosphopeptides and therefore the effective depth of signaling coverage. Importantly, enrichment is not a generic plug‑in step; it shapes the visible phosphoproteome by favoring certain peptide chemistries. A focused tutorial and review of phosphopeptide enrichment materials emphasizes that choosing an enrichment strategy should be driven by sample type, desired breadth versus specificity, and the practical scale of the experiment, because no single chemistry captures all classes of phosphopeptides equally well [13].
Equally critical is preserving the in‑vivo phosphorylation state. Rapid quenching, phosphatase inhibition, consistent lysis conditions, and avoiding variable processing times across replicates are essential to prevent artifactual loss or gain of phosphorylation. Treat these steps as part of the biology: if phosphorylation is the language of signaling, sample handling is how you prevent the message from being rewritten after harvest.
7. From discovery to trait relevance: connecting phosphorylation to growth and resilience
Phosphoproteomics becomes most powerful when it bridges discovery and trait relevance—when regulated sites point to proteins that can be functionally tested and ultimately associated with agronomic performance. A strong pattern across successful studies is that they do not stop at reporting regulated sites; they connect site changes to protein stability, localization, or protein–protein interactions and then validate with targeted experiments. In cucumber, phosphorylation of CsIREH1 was shown to regulate DELLA protein behavior and thereby affect plant height, directly tying a specific phosphorylation event to a classic growth control node and a measurable trait [14]. This illustrates a general principle: phosphorylation sites are most valuable when they reside on “decision proteins” whose state determines whether growth proceeds or is restrained.
The same reasoning applies to immune resilience. Plant NLR receptors are tightly regulated by post‑translational modifications to balance defense and fitness costs, and synthesis alone cannot explain their rapid switching behavior. Reviews focused on NLR regulation highlight how PTMs govern activation thresholds and signaling competence, reinforcing that phosphorylation‑centric workflows are not merely descriptive—they are strategically aligned with the biology of rapid immune decisions [15].
Practically, this means the best project plans begin with an explicit decision point (“when does growth stop?”, “when does immunity activate?”) and then choose sampling, perturbations, and validation assays to interrogate that decision. When done well, phosphoproteomics is not an endpoint; it is the fastest route to causal, testable hypotheses in hormone and stress biology.
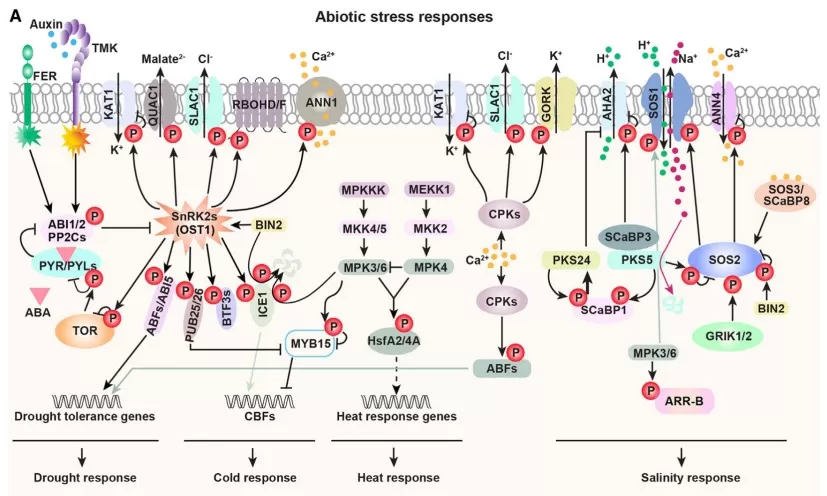
Protein phosphorylation in stress response
Image reproduced from Zhang et al., 2023, Cell Rep, licensed under the Creative Commons CC-BY license (CC BY).
Conclusion: Making Plant Hormone Signaling Visible with Phosphoproteomics
Plant hormone signaling is defined by speed and by state changes that often occur without changes in protein abundance. Proteomics provides the abundance layer needed to interpret longer‑term remodeling, but phosphoproteomics reveals the activity switches that drive receptor‑proximal decisions and shape transcription factor outputs. The most productive studies treat experimental design, enrichment, and interpretation as an integrated system: early time points capture transient switches, genetic or environmental perturbations support causality, and network‑aware annotation focuses attention on the sites most likely to matter.
If you are planning a hormone signaling project, start with a clear phenotypic decision window, select a time‑course or perturbation design that can capture it, and commit to reproducible sample handling and enrichment. With those foundations, phosphoproteomics can “make signaling visible” in a way that transcriptomics alone cannot—turning rapid phosphorylation events into mechanistic models and, ultimately, trait‑relevant hypotheses.
Reference
1. Kim SH, Bahk S, Nguyen NT, Pham MLA, Kadam US, Hong JC, Chung WS. Phosphorylation of the auxin signaling transcriptional repressor IAA15 by MPKs is required for the suppression of root development under drought stress in Arabidopsis. Nucleic Acids Res. 2022 Oct 14;50(18):10544-10561. doi: 10.1093/nar/gkac798.
2. Zhang WJ, Zhou Y, Zhang Y, Su YH, Xu T. Protein phosphorylation: A molecular switch in plant signaling. Cell Rep. 2023 Jul 25;42(7):112729. doi: 10.1016/j.celrep.2023.112729.
3. Yamashita K, Katagiri S, Takase H, Li Y, Oishi A, Otoguro A, Kamiyama Y, Yamauchi S, Wang YS, Takemiya A, Mori IC, Kollist H, Umezawa T. MAP4K1 and MAP4K2 regulate ABA-induced and Ca2+-mediated stomatal closure in Arabidopsis. Sci Adv. 2025 Dec 19;11(51):eadt4916. doi: 10.1126/sciadv.adt4916.
4. Wang Y, Tong X, Qiu J, Li Z, Zhao J, Hou Y, Tang L, Zhang J. A phosphoproteomic landscape of rice (Oryza sativa) tissues. Physiol Plant. 2017 Aug;160(4):458-475. doi: 10.1111/ppl.12574.
5. Urban J. A review on recent trends in the phosphoproteomics workflow. From sample preparation to data analysis. Anal Chim Acta. 2022 Mar 22;1199:338857. doi: 10.1016/j.aca.2021.338857.
6. Hsu CC, Zhu Y, Arrington JV, Paez JS, Wang P, Zhu P, Chen IH, Zhu JK, Tao WA. Universal Plant Phosphoproteomics Workflow and Its Application to Tomato Signaling in Response to Cold Stress. Mol Cell Proteomics. 2018 Oct;17(10):2068-2080. doi: 10.1074/mcp.TIR118.000702.
7. Friml J, Gallei M, Gelová Z, Johnson A, Mazur E, Monzer A, Rodriguez L, Roosjen M, Verstraeten I, Živanović BD, Zou M, Fiedler L, Giannini C, Grones P, Hrtyan M, Kaufmann WA, Kuhn A, Narasimhan M, Randuch M, Rýdza N, Takahashi K, Tan S, Teplova A, Kinoshita T, Weijers D, Rakusová H. ABP1-TMK auxin perception for global phosphorylation and auxin canalization. Nature. 2022 Sep;609(7927):575-581. doi: 10.1038/s41586-022-05187-x.
8. Cruz-Mireles N, Osés-Ruiz M, Derbyshire P, Jégousse C, Ryder LS, Bautista MJA, Eseola A, Sklenar J, Tang B, Yan X, Ma W, Findlay KC, Were V, MacLean D, Talbot NJ, Menke FLH. The phosphorylation landscape of infection-related development by the rice blast fungus. Cell. 2024 May 9;187(10):2557-2573.e18. doi: 10.1016/j.cell.2024.04.007.
9. Yao Q, Ge H, Wu S, Zhang N, Chen W, Xu C, Gao J, Thelen JJ, Xu D. P³DB 3.0: From plant phosphorylation sites to protein networks. Nucleic Acids Res. 2014 Jan;42(Database issue):D1206-13. doi: 10.1093/nar/gkt1135. Epub 2013 Nov 15.
10. Zhu C, Jing B, Lin T, Li X, Zhang M, Zhou Y, Yu J, Hu Z. Phosphorylation of sugar transporter TST2 by protein kinase CPK27 enhances drought tolerance in tomato. Plant Physiol. 2024 May 31;195(2):1005-1024. doi: 10.1093/plphys/kiae124.
11. Chen L, Zhang Y, Bu Y, Zhou J, Man Y, Wu X, Yang H, Lin J, Wang X, Jing Y. Imaging the spatial distribution of structurally diverse plant hormones. J Exp Bot. 2024 Dec 4;75(22):6980-6997. doi: 10.1093/jxb/erae384.
12. Wang G, Chen X, Yu C, Shi X, Lan W, Gao C, Yang J, Dai H, Zhang X, Zhang H, Zhao B, Xie Q, Yu N, He Z, Zhang Y, Wang E. Release of a ubiquitin brake activates OsCERK1-triggered immunity in rice. Nature. 2024 May;629(8014):1158-1164. doi: 10.1038/s41586-024-07418-9.
13. Qiu W, Evans CA, Landels A, Pham TK, Wright PC. Phosphopeptide enrichment for phosphoproteomic analysis - A tutorial and review of novel materials. Anal Chim Acta. 2020 Sep 8;1129:158-180. doi: 10.1016/j.aca.2020.04.053.
14. Zhao H, Sun P, Tong C, Li X, Yang T, Jiang Y, Zhao B, Dong J, Jiang B, Shen J, Li Z. CsIREH1 phosphorylation regulates DELLA protein affecting plant height in cucumber (Cucumis sativus). New Phytol. 2025 Feb;245(4):1528-1546. doi: 10.1111/nph.20309.
15. Zhong C, Wang X, Li Y, Dinesh-Kumar SP, Zhang Y. Regulation of plant NLRs by post-translational modifications. Trends Biochem Sci. 2026 Jan;51(1):51-63. doi: 10.1016/j.tibs.2025.10.008.
Next-Generation Omics Solutions:
Proteomics & Metabolomics
Ready to get started? Submit your inquiry or contact us at support-global@metwarebio.com.


