In the era of precision medicine, clinical proteomics offers a real-time snapshot of what is happening within the body—far beyond what genomics alone can reveal. While genes serve as blueprints, proteins are the functional agents that execute life processes, dynamically responding to disease and therapeutic interventions. More than 95% of approved drugs target proteins, underscoring the central role of proteomics in understanding disease mechanisms and guiding clinical decisions [1]. Yet despite years of proteomics research and thousands of promising biomarkers, comprehensive protein analysis has not become routine in clinical settings. This article explores the journey of clinical proteomics from bench to bedside, examining the technological breakthroughs—from mass spectrometry and affinity-based platforms to AI integration—alongside the persistent challenges of sample complexity, standardization, and bioinformatics bottlenecks, and the innovative strategies that may finally realize the vision of proteomics-driven precision medicine.
Figure 1. Bridging the Gap Between Proteomics Technology and Clinical Application. Image reproduced from Albrecht, V., Müller-Reif, J., Nordmann, T. M. et al., 2024, Molecular & Cellular Proteomics: MCP, licensed under the Creative Commons Attribution License (CC BY 4.0).
1. The Promise of Clinical Proteomics: Why the Time is Now
1.1 Technological Advancements: Catalysts for Change
The Mass Spectrometry Revolution
In the past decade, mass spectrometry (MS)-based proteomics has undergone revolutionary advancements. High-resolution, accurate-mass (HR/AM) instruments, once confined to specialized core facilities, are now available in mainstream laboratories. These instruments allow for the detection and quantification of proteins across a vastly extended dynamic range. Modern hybrid high-speed mass spectrometers enable rapid and sensitive parallel reaction monitoring (PRM), with exceptional reproducibility and low coefficients of variation—sufficient to analyze many of the top 1000 plasma proteins [2].
This leap in technology has dramatically expanded the possibilities of proteomics. While earlier studies were restricted to abundant proteins, current instruments can access low-abundance proteins, which may hold the most clinical significance.
The Emergence of Affinity-Based Platforms
In parallel with mass spectrometry, affinity-based technologies have emerged as powerful tools in proteomics, complementing and enhancing traditional methods. Technologies like Olink and SomaScan are at the forefront of this evolution, enabling high-throughput, multiplexed protein detection with exceptional sensitivity and specificity. Olink, utilizing proximity extension assay (PEA) technology, allows for the simultaneous measurement of hundreds of proteins in a single sample, making it ideal for clinical applications requiring large-scale, reproducible analysis. Similarly, SomaScan, based on aptamer technology, offers the capability to detect thousands of proteins with high precision. These platforms are accelerating the translation of proteomics into clinical practice by providing scalable, standardized solutions for biomarker discovery, disease monitoring, and personalized medicine. Their ability to analyze protein panels associated with various disease states holds great promise for improving diagnostics, guiding therapeutic decisions, and monitoring treatment responses, ultimately enhancing the precision of clinical care. (Learn more at: MS-DIA vs. Olink vs. SomaScan)
Single-Cell and Spatial Proteomics
Single-cell and spatial proteomics are transforming clinical proteomics by offering detailed insights into cellular and tissue-specific protein expression. Single-cell proteomics allows for the analysis of individual cells, revealing heterogeneity within diseases like cancer and neurodegenerative disorders, where cellular diversity impacts disease progression and treatment response. Spatial proteomics, on the other hand, maps protein expression within tissue contexts, enhancing our understanding of disease mechanisms and guiding more precise, personalized therapies. Together, these technologies are set to revolutionize diagnostics and therapeutic strategies by providing a level of detail beyond what traditional bulk proteomics can offer.
1.2 Addressing Clinical Gaps: A Push from Medicine
Moving Beyond Single Biomarkers
The limitations of single-protein biomarkers are widely acknowledged. For example, PSA, while widely used, suffers from low specificity, leading to unnecessary biopsies and over-treatment. Troponin, while effective in diagnosing myocardial injury, provides little insight into disease mechanisms or prognosis. The need for multi-protein panels that capture the complexity of disease biology is clear [3].
Recent advances suggest that targeted protein panels—carefully selected sets of proteins measured with high analytical rigor—may soon replace traditional antibody-based assays in clinical settings. These panels offer multiplexing capabilities, enhanced sensitivity, and reproducibility, making them ideal candidates for clinical applications [3].
Reclassifying Disease Through Proteomics
Proteomics is driving a fundamental shift in how diseases are classified. In oncology, proteomic profiling has uncovered subtypes within traditional cancer types, each with distinct behaviors and therapeutic responses. Beyond oncology, proteomic signatures are being developed for complex neurological disorders like Alzheimer's disease, where early diagnosis remains a significant challenge. In cardiovascular medicine, proteomics is being used for early detection and risk stratification, helping identify at-risk individuals before clinical symptoms manifest [4].
Proteomics in Companion Diagnostics and Monitoring Therapy
With the expansion of targeted and immunotherapies in oncology, the need for dynamic biomarkers that can track treatment response and predict resistance is increasingly urgent. Proteomics provides a unique advantage by enabling the monitoring of pathway activation states, detecting early signs of therapeutic efficacy, and identifying resistance mechanisms before they become clinically apparent.
2. Challenges: The Complex Journey from Bench to Bedside
2.1 Sample Complexity: The Challenge of Low-Abundance Proteins
The Dynamic Range Problem
Blood plasma, the most commonly used clinical sample, presents a substantial analytical challenge due to its vast dynamic range. Protein concentrations vary by more than ten orders of magnitude, with abundant proteins like albumin constituting roughly 55% of total protein mass, while lower-abundance proteins may exist at concentrations billions of times lower. This wide range exceeds the detection capabilities of even the most advanced mass spectrometers, necessitating complex fractionation or depletion strategies that introduce variability and increase complexity [5].
Heterogeneity Across Sample Types
Clinical specimens are highly heterogeneous—serum, plasma, urine, tissue biopsies, and cerebrospinal fluid each have unique pre-analytical variables that impact protein measurement. Even within a single sample type, factors like anticoagulant choice, processing time, and storage conditions can significantly influence the detectable proteome. Standardizing these variables across different clinical sites remains a major challenge [5].
Characteristics and Analytical Challenges of Common Clinical Sample Types in Proteomics
| Sample Type | Protein Complexity | Dynamic Range | Major Abundant Proteins | Key Analytical Challenges |
|---|---|---|---|---|
| Serum | Medium | Very High | Albumin, immunoglobulins | Abundant proteins mask low-abundance biomarkers; variability in sample collection and storage |
| Plasma | High | Very High | Albumin, immunoglobulins | Similar to serum, with additional complexity from coagulation-related proteins; processing time critical |
| Urine | Low to Medium | Moderate | Uromodulin, albumin | Low total protein content; high variability due to hydration and diurnal factors; potential contamination from non-renal sources |
| Tissue Biopsies | Very High | High | Varies by tissue type | Limited sample quantity; need for rapid processing to prevent degradation; tissue heterogeneity within biopsy |
| Cerebrospinal Fluid (CSF) | Low to Medium | High | Albumin, IgG | Low sample volume; extremely low protein concentration; protein loss during collection |
2.2 Achieving Standardization and Reproducibility
In research settings, methodological flexibility is a virtue—it enables innovation and adaptation to specific biological questions. In clinical diagnostics, however, flexibility is the enemy of reliability. Clinical laboratories require assays that produce identical results across different operators, instruments, and days.
Achieving this level of reproducibility in proteomic assays has proven difficult. The field lacks consensus on reference materials and performance metrics that would enable robust inter-laboratory comparisons. As noted in an early NHLBI report, key barriers include false positive associations that occur with analysis of high-dimensional datasets and the limited understanding of the effects of growth, development, and age on the normal plasma proteome [5].
2.3 Throughput and Cost Considerations
Comprehensive proteomic profiling, whether through discovery-based mass spectrometry or high-plex affinity assays, remains expensive and time-consuming. Clinical laboratories process large volumes of samples daily, yet current proteomic technologies do not offer the throughput required. Despite the promise of targeted MS-based laboratory-developed tests (LDTs), scalability and ease of use must be improved for these technologies to become widely adopted [3]. Moreover, healthcare reimbursement models present further challenges, as novel proteomic tests must demonstrate both clinical utility and cost-effectiveness relative to existing diagnostic methods.
2.4 The Bioinformatics Bottleneck
The complex nature of mass spectrometry-generated data poses a significant bioinformatics challenge. Converting raw data into meaningful peptide identifications, quantifying protein abundances, and applying appropriate statistical analyses requires specialized expertise that is scarce in clinical settings. Moreover, distinguishing biologically relevant protein changes from technical noise requires sophisticated analytical tools and a deep understanding of disease biology.
3. Accelerating Clinical Proteomics: Solutions and Future Pathways
3.1 Targeted Proteomics: The Path to Clinical Translation
The growing recognition that discovery and clinical proteomics require distinct approaches is shaping the field's evolution. Discovery-based proteomics (data-dependent acquisition, DDA, and data-independent acquisition, DIA) aims to capture a broad spectrum of proteins from a sample. In contrast, targeted proteomics (multiple reaction monitoring, MRM, and parallel reaction monitoring, PRM) focuses on the precise quantification of a predefined set of proteins, ensuring maximum analytical rigor [3]. (Learn more at: Mass Spectrometry Acquisition Mode Showdown: DDA vs. DIA vs. MRM vs. PRM)
The transition from discovery to targeted proteomics is the most promising pathway for clinical implementation. Recent advances in hybrid mass spectrometry technologies have shown that these modern instruments can effectively bridge the gap between discovery proteomics and routine clinical testing, thereby enhancing the diagnostic and prognostic value of protein biomarkers [2]. By concentrating on rigorously validated protein panels tailored to specific clinical questions, targeted proteomics can provide the high analytical performance required by both regulatory agencies and clinicians.
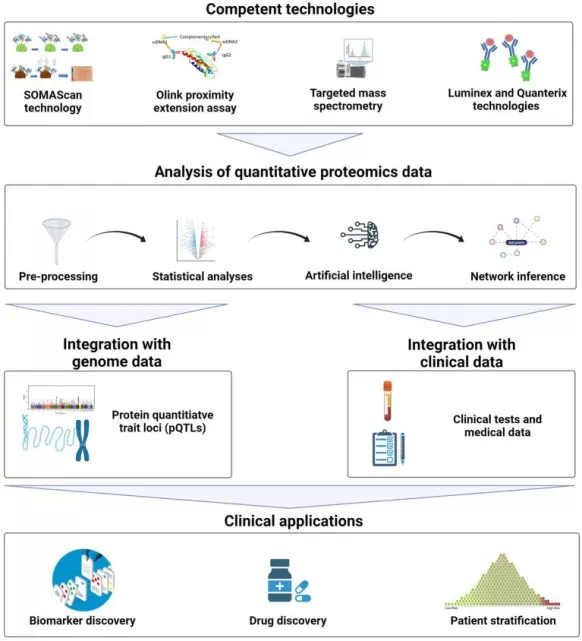
Figure 2. Integrated Workflow of Clinical Proteomics. Image reproduced from Correa Rojo, A., Heylen, D., Aerts, J. et al., 2021, Frontiers in Physiology, licensed under the Creative Commons Attribution License (CC BY 4.0).
3.2 Advancements in Automation and Microfluidics
Automation plays a key role in improving reproducibility and throughput in proteomics. Automated systems for sample preparation, protein extraction, and digestion are reducing human error and variability. Commercially available, fully automated mass spectrometry platforms are now capable of replacing labor-intensive manual workflows, streamlining processes, and supporting faster, standardized care.
3.3 The Role of Artificial Intelligence in Proteomics
Artificial intelligence (AI) and machine learning are transforming proteomics workflows by enhancing both the accuracy and efficiency of data analysis. AI-driven deep learning models are now integral to tasks such as peptide identification, spectrum interpretation, and chromatography prediction, significantly improving the speed and reliability of these processes. Furthermore, AI facilitates the integration of proteomic data with other omics layers, including genomics, transcriptomics, and metabolomics, providing multi-dimensional insights into disease biology. By combining these datasets, AI algorithms can identify complex biomarker signatures and predict disease progression with greater precision. As proteomics technologies advance, future research will focus on expanding AI applications to the human proteome, improving multi-source omics integration, and refining predictive models for clinical use [1]. This integration will enable more personalized diagnostic and therapeutic strategies, driving the future of precision medicine.
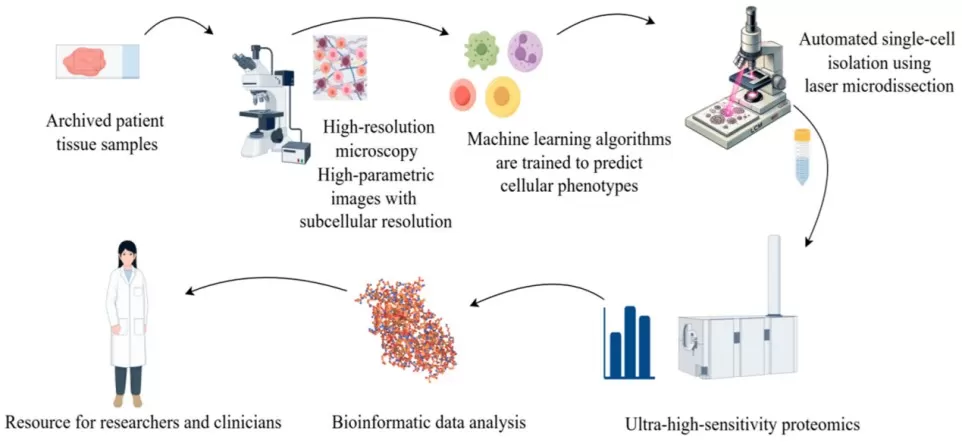
Figure 3. AI-Driven Workflow in Proteomics: Integrating High-Resolution Microscopy, Machine Learning, and Ultra-High-Sensitivity Technologies. Image reproduced from Xie, Y., Chen, X., Xu, M. et al., 2025, Biomedicines, licensed under the Creative Commons Attribution License (CC BY 4.0).
3.4 Collaborative Approaches: Bridging Research, Clinical, and Industrial Gaps
Successful clinical proteomics requires collaboration between researchers, clinicians, and industry partners. Proteomic research must be rooted in clinically relevant questions, with input from healthcare professionals who will ultimately use the tests. Conversely, clinicians must understand the limitations of proteomics technologies to formulate meaningful questions and interpret results appropriately.
4. The Future of Precision Medicine: A Vision for Clinical Proteomics
Clinical proteomics stands at a pivotal juncture. It is not yet fully established as either a research tool or a routine clinical practice. Like a developing field, it possesses immense potential but faces significant challenges that require meticulous development to realize its full impact. Despite these hurdles, the future of clinical proteomics is promising. Ongoing technological advancements will address critical issues such as sample complexity and bioinformatics bottlenecks, while efforts toward standardization will establish consistent metrics. Automation will enhance throughput, and artificial intelligence, coupled with collaborative ecosystems, will accelerate the field's progression. These innovations will enable proteomics to become an integral part of personalized medicine.
Envision a future where a routine health check includes a comprehensive proteomic profile, providing insights not only into functional protein networks but also into cardiovascular risk and the early detection of neurodegenerative diseases—years before clinical symptoms manifest. In oncology, proteomic analysis of tumors and liquid biopsies will guide treatment decisions, while serial proteomic monitoring will detect emerging resistance, facilitating timely therapeutic adjustments. This vision is within reach. As proteomics technologies continue to evolve, they hold the promise of advancing diagnostics, enabling personalized therapies, and ultimately improving patient outcomes. Overcoming existing challenges will be essential for the full integration of proteomics into clinical practice.
MetwareBio: Your Trusted Partner for Proteomics Analysis
MetwareBio offers cutting-edge proteomics services designed to bridge the gap between research discovery and clinical application. Our expert team leverages state-of-the-art mass spectrometry platforms and affinity-based technologies to deliver high-quality, reproducible protein quantification across diverse sample types.
MetwareBio's Proteomics Analysis Service provides comprehensive solutions for biomarker discovery, protein panel development, and multi-omics integration, supporting your journey from bench to bedside with rigorous analytical standards and personalized scientific support.
If you are interested in clinical proteomics or would like to learn more about our services, please do not hesitate to contact us.
Contact UsReferences
- Xie, Y., Chen, X., Xu, M., & Zheng, X. (2025). Application of the Human Proteome in Disease, Diagnosis, and Translation into Precision Medicine: Current Status and Future Prospects. Biomedicines, 13(3), 681. https://doi.org/10.3390/biomedicines13030681
- Wahle, M., Remes, P. M., Albrecht, V., Baggio Lorenz, M., Mueller-Reif, J., Steigerwald, S., Heymann, T., Niu, L., Lössl, P., Horning, S., Jacob, C. C., & Mann, M. (2025). A Novel Hybrid High-Speed Mass Spectrometer Allows Rapid Translation From Biomarker Candidates to Targeted Clinical Tests Using 15N-Labeled Proteins. Molecular & Cellular Proteomics: MCP, 24(9), 101050. https://doi.org/10.1016/j.mcpro.2025.101050
- Ramesh, P., & Prasad, T. S. K. (2025). Future of clinical proteomics: could targeted multi-protein panels supplant low-throughput methods? Expert Review of Proteomics, 22(11-12), 447–452. https://doi.org/10.1080/14789450.2025.2604161
- Alshehri, S., Vitorino, R., Saleh, O., Al-Harthi, S., Alahmadi, A., Alotibi, R., da Silva Rosa, S. C., Osama, A., Magedeldin, S., Alhattab, D., Emwas, A. H., & Jaremko, M. (2025). Advances and applications of clinical proteomics in precision medicine. Expert Review of Proteomics, 22(10), 401–420. https://doi.org/10.1080/14789450.2025.2560919
- Gerszten, R. E., Accurso, F., Bernard, G. R., Caprioli, R. M., Klee, E. W., Klee, G. G., Kullo, I., Laguna, T. A., Roth, F. P., Sabatine, M., Srinivas, P., Wang, T. J., & Ware, L. B. (2008). Challenges in translating plasma proteomics from bench to bedside: update from the NHLBI Clinical Proteomics Programs. American Journal of Physiology. Lung Cellular and Molecular Physiology, 295(1), L16–L22. https://doi.org/10.1152/ajplung.00044.2008
- Albrecht, V., Müller-Reif, J., Nordmann, T. M., Mund, A., Schweizer, L., Geyer, P. E., Niu, L., Wang, J., Post, F., Oeller, M., Metousis, A., Bach Nielsen, A., Steger, M., Wewer Albrechtsen, N. J., & Mann, M. (2024). Bridging the Gap From Proteomics Technology to Clinical Application: Highlights From the 68th Benzon Foundation Symposium. Molecular & Cellular Proteomics: MCP, 23(12), 100877. https://doi.org/10.1016/j.mcpro.2024.100877
- Correa Rojo, A., Heylen, D., Aerts, J., Thas, O., Hooyberghs, J., Ertaylan, G., & Valkenborg, D. (2021). Towards Building a Quantitative Proteomics Toolbox in Precision Medicine: A Mini-Review. Frontiers in Physiology, 12, 723510. https://doi.org/10.3389/fphys.2021.723510


