Untargeted Spatial Metabolomics
Untargeted Spatial Metabolomics
Overview of Untargeted Spatial Metabolomics
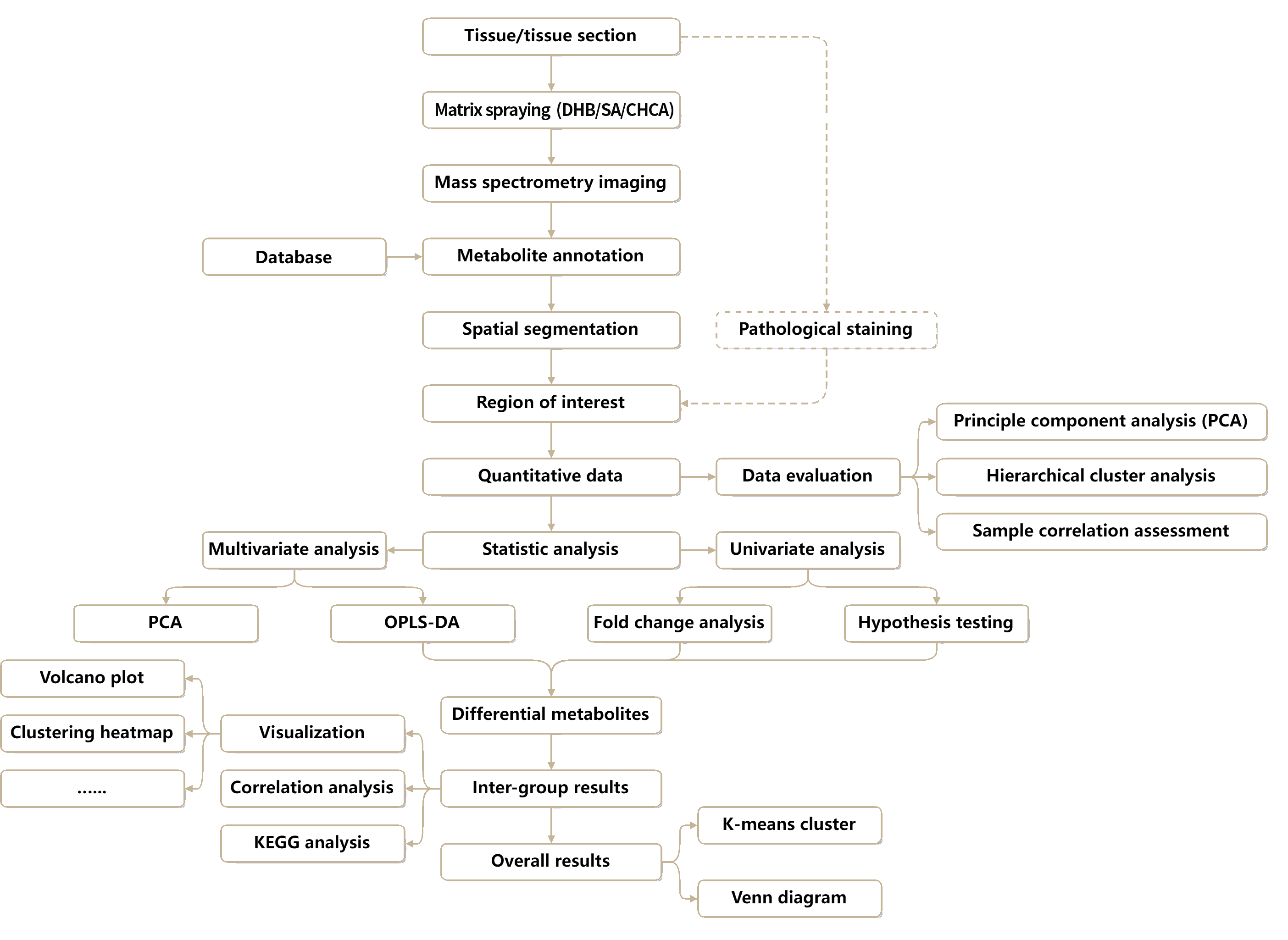
The Technical Workflow of Spatial Metabolomics Based on MALDI or AFADESI
Why Choose MetwareBio for Untargeted Spatial Metabolomics?
MetwareBio’s Spatial Metabolomics In-house Database
| Untargeted Spatial Metabolomics Database (Human/Animal) | |
| Compound Class | Number |
| Amino acid and Its metabolites | 470+ |
| Organic acid and Its derivatives | 200+ |
| Nucleotide and Its metabolites | 140+ |
| Hormones and related compounds | 60+ |
| Carbohydrates and Its metabolites | 50+ |
| CoEnzyme and vitamins | 40+ |
| Bile Acids | 25+ |
| Tryptamines, Cholines, Pigments | 10+ |
| Lipids | 80+ |
| Benzene and substituted derivatives | 410+ |
| Heterocyclic compounds | 160+ |
| Alcohol and amines | 40+ |
| Others | 110+ |
| Total | 1800+ |
| Untargeted Spatial Metabolomics Database (Plant) | |
| Compound Class | Number |
| Flavonoids | 360+ |
| Alkaloids | 160+ |
| Phenolic acids | 140+ |
| Terpenoids | 75+ |
| Lignans and coumarins | 65+ |
| Quinones | 25+ |
| Steroids | 15+ |
| Tannins | 10+ |
| Amino acids and derivatives | 110+ |
| Nucleotides and derivatives | 95+ |
| Organic acids | 45+ |
| Lipids | 40+ |
| Others | 315+ |
| Total | 1450+ |
Untargeted Spatial Metabolomics Analysis Workflow
Deliverables of Untargeted Spatial Metabolomics Service
Project Experience of Untargeted Spatial Metabolomics Analysis
Number of Metabolites Mapped Across Human and Animal Tissues by MALDI Untargeted Spatial Metabolomics at 50 µm Resolution in Positive-Ion Mode
Number of Metabolites Mapped Across Various Plant Tissues by MALDI Untargeted Spatial Metabolomics at 50 µm Resolution in Positive-Ion Mode
Applications of Untargeted Spatial Metabolomics Analysis
Untargeted spatial metabolomics enables in situ characterization of metabolic reprogramming within complex tissues, supporting research on disease mechanisms, tumor microenvironment (TME), biomarker discovery, and region-specific pathological metabolism. By preserving tissue context, this approach helps reveal localized metabolic alterations associated with disease progression and molecular heterogeneity.
In pharmacology and translational research, untargeted spatial metabolomics supports the study of spatial DMPK, drug tissue distribution, toxicity and safety assessment, and target-related metabolic responses. Spatially resolved metabolite profiling provides valuable insight into how drugs affect different tissue regions, supporting mechanism-of-action studies and preclinical evaluation.
For animal research, untargeted spatial metabolomics is well suited for investigating reproductive development, disease mechanisms, nutrient metabolism, and toxicological responses across diverse tissue types. This approach enables spatial analysis of metabolic organization and tissue-specific biochemical changes, supporting both basic biology and preclinical studies.
In plant research, untargeted spatial metabolomics enables direct visualization of active compound distribution, reproductive development, abiotic stress responses, and metabolism associated with crop traits. By mapping metabolites within intact plant tissues, this technology supports deeper understanding of tissue-specific metabolism, adaptive regulation, and agricultural improvement.
Case Studies of Untargeted Spatial Metabolomics Analysis
Case 1: Spatial Metabolomics Reveals Region-Specific Metabolic Remodeling in Human Liver Disease
A Nature Genetics study applied MetwareBio’s MSI-based spatial metabolomics alongside spatial transcriptomics and spatial proteomics to profile 61 human liver samples spanning control, MASL, and MASH stages. The study revealed MASLD-specific phospholipid accumulation and linked these metabolic changes to lipid-associated macrophages and disease-associated fibrotic regions, demonstrating the value of spatial metabolomics for resolving disease mechanisms, metabolic reprogramming, and tissue microenvironment heterogeneity.
Source: Li, Z., Luo, G., Gan, C. et al. Spatially resolved multi-omics of human metabolic dysfunction-associated steatotic liver disease. Nat Genet 57, 3112–3125 (2025). https://doi.org/10.1038/s41588-025-02407-8
MASLD-associated metabolic modules revealed by Hotspot (Li et al., Nat Genet. 2025)
Case 2: Spatial Metabolomics Reveals Dynamic Metabolite Distribution During Tomato Fruit Ripening
A Food Research International study used MetwareBio’s MALDI-MSI to visualize metabolite distributions in tomato fruit across four maturity stages, from green to red ripening. The analysis showed that soluble sugars, organic acids, and amino acids were broadly distributed throughout the fruit, while several volatile organic compounds (VOCs) shifted from widespread localization in the flesh to stronger accumulation near the peel during maturation. These findings demonstrate the utility of spatial metabolomics for studying plant development, metabolite localization, fruit quality, and crop trait biology.
Source: Guan S, Liu C, Yao Z, et al. Visualizing the spatial distribution of metabolites in tomato fruit at different maturity stages by matrix-assisted laser desorption/ionization mass spectrometry imaging. Food Res Int. 206:115973 (2025). https://doi.org/10.1016/j.foodres.2025.115973
The MALDI-MSI of targeted primary metabolites and relative content in different mature stages in the tomato fruit (Guan et al., Food Res Int. 2025)
Sample Requirements & Submission Guidelines for Spatial Metabolomics
- Embedding Medium: CMC is recommended for optimal tissue support during sectioning.
- Tissue Cross-section Size: 1.5 × 1.5 mm (min) – 50 × 30 mm (max)
- Tissue Height: 2 mm (min) – 25 mm (max)
- Storage & Shipment: Store samples in a frozen tissue storage box and ship on dry ice to maintain sample integrity.
- Slide Type: ITO-coated slides are required for MALDI-MSI analysis.
- Maximum Scanning Area: 65 × 40 mm
- Section Thickness: 8–50 µm
- Storage & Shipment: Store sections in a slide box and ship on dry ice.
FAQ about Untargeted Spatial Metabolomics Service
Spatial metabolomics is a cutting-edge technology that combines metabolomics with imaging techniques to map the spatial distribution of metabolites within biological samples. In a typical workflow, metabolites are ionized from defined positions on the sample surface and recorded as spatially resolved molecular signals, which are then reconstructed into metabolite distribution maps. This enables qualitative, quantitative and localization analysis of thousands of metabolites in biological tissues.
Spatial metabolomics offers detailed, spatially resolved metabolic information, which is essential for understanding localized metabolic processes and heterogeneity within tissues. Bulk metabolomics, on the other hand, provides a broad, averaged overview of metabolites across a homogenized sample, making it suitable for general metabolic profiling without spatial context. The key difference in experimental procedures is that spatial metabolomics involves freezing and sectioning the sample, applying a specific matrix, and then performing mass spectrometry imaging (MSI) for analysis. In contrast, bulk metabolomics requires homogenizing fresh samples, extracting metabolites, and analyzing them using LC-MS.
Both MALDI-MSI and AFADESI-MSI are widely used for spatial metabolomics, but they differ primarily in their ionization mechanisms. MALDI-MSI uses a matrix-coated tissue surface and laser irradiation to desorb and ionize metabolites from defined spatial coordinates, making it well suited for high-spatial-resolution imaging. In contrast, AFADESI-MSI is an ambient ionization technique that generates ions from the tissue surface through airflow-assisted desorption and electrospray ionization, without requiring laser-based matrix desorption. Because of these differences, the two technologies show distinct characteristics in sample preparation, spatial resolution, scanning area, and metabolite coverage.
Choice depends on sample size, spatial resolution requirements, and research goals. MALDI-MSI supports spatial resolution down to 5 µm, offers more flexible resolution settings, and supports MS/MS-based identification, making it the preferred option when sample compatibility allows. AFADESI-MSI is often recommended for larger tissue sections, such as whole-body or large-format samples that exceed the standard MALDI scanning area. In most cases, MALDI-MSI is recommended first if the sample is suitable.
Samples can be submitted as either fresh-frozen tissue blocks or prepared tissue sections. For tissue blocks, rapid dissection under cold conditions is recommended, with removal of excess hair, blood, and other contaminants before embedding in a suitable medium. For tissue sections, fresh-frozen blocks should be cryosectioned to 8–50 µm thickness (8–12 µm is commonly recommended for animal tissues) and mounted on ITO-coated slides for MALDI-MSI or standard adhesive slides for AFADESI-MSI. If sections are submitted, four sections per sample are recommended: one for H&E staining and three for MSI analysis or backup. Both tissue blocks and sections should be sealed in appropriate frozen storage containers, stored at −80°C, and shipped on dry ice.
FFPE samples can be analyzed, but they are generally not recommended for spatial metabolomics. During FFPE processing, metabolites may undergo loss, diffusion, or relocation, which can compromise the original spatial distribution of small molecules. For more reliable spatial metabolomics results, fresh-frozen tissues are strongly preferred.
CMC and FSC22 are recommended embedding media for untargeted spatial metabolomics. OCT can be compatible in some cases, but it may interfere with metabolite detection and data quality. For integrated spatial multi-omics analysis, FSC22 is generally recommended because it offers better compatibility across different spatial omics workflows.
For MALDI-MSI, available spatial resolutions include 5, 10, 20, 50, and 100 µm. For AFADESI-MSI, available spatial resolutions include 50 and 100 µm. The optimal setting depends on tissue structure, sample size, and the level of spatial detail required for the study.
The resolution should be chosen based on the sample type, the spatial heterogeneity of the metabolites, and the desired level of detail for analysis. For large or heterogeneous samples (e.g., organs), lower resolutions (100 µm to 50 µm) are typically sufficient to observe broad metabolite distributions, while higher resolutions (10 µm to 5 µm) are necessary for smaller, more homogeneous samples (e.g., cellular or subcellular structures) to capture finer details. If metabolites are widely distributed, a lower resolution is adequate, but if they are localized to small regions (e.g., organelles or specific cell types), a higher resolution provides better visualization of spatial heterogeneity.
Typical Resolutions Used:
- 100 µm: Suitable for large tissue sections or general metabolite distribution mapping.
- 50 µm: Provides a good balance between resolution and data volume for tissues or larger regions of interest.
- 20 µm: Used for more detailed mapping of tissue microstructures.
- 10 µm or 5 µm: High-resolution imaging for single cells or specific subcellular structures.
Next-Generation Omics Solutions:
Proteomics & Metabolomics
Ready to get started? Submit your inquiry or contact us at support-global@metwarebio.com.


