High-throughput omics technologies can generate extensive lists of differentially expressed genes, proteins, post-translational modifications, and metabolites. Yet these lists, on their own, do not explain biology. The real challenge is moving from molecular changes to biological meaning: Which pathways are perturbed? Which regulatory programs are activated or suppressed? How do these individual changes fit into a broader cellular response? This is why pathway analysis has become a core step in modern omics research. Among the major pathway resources available today, Reactome stands out for its mechanistic depth, rigorous manual curation, and strong support for data interpretation across multiple omics layers.
Reactome is especially valuable because it represents biology as a connected series of molecular events. Instead of simply listing pathway members, it captures how molecules interact, change state, move between compartments, and regulate one another. For omics researchers, that makes Reactome more than a reference database; it is a practical framework for translating high-dimensional data into testable biological hypotheses. In this guide, we explain what Reactome is, which omics workflows it supports best, how it complements KEGG, and how to interpret common Reactome pathway-analysis visualizations in a biologically meaningful way.
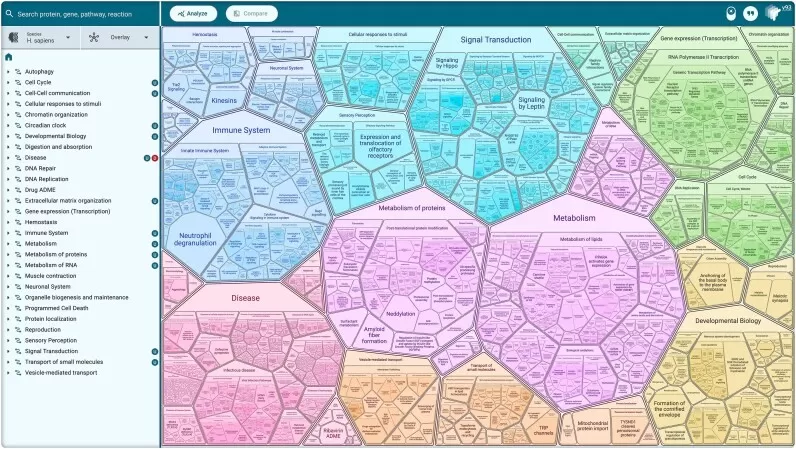
Figure 1. Reactome Pathway Space at a Glance. Image reproduced from Ragueneau, E., Gong, C., Sinquin, P. et al., 2026, Nucleic acids research, licensed under the Creative Commons Attribution License (CC BY 4.0).
1. What Is Reactome?
Reactome is an open-access, open-source, manually curated, and peer-reviewed pathway knowledgebase that describes biological processes as ordered networks of molecular reactions. These reactions include not only classical metabolic conversions, but also binding events, transport, translocation, activation, degradation, complex assembly, and other state changes that are central to cell biology.
1.1 Key Features of the Reactome Pathway Database
High-quality manual curation
A defining strength of Reactome is its emphasis on expert curation. Pathways are authored and reviewed by domain specialists and supported by published literature. This gives Reactome a level of biological granularity and interpretability that is particularly valuable in omics studies, where researchers need more than just a list of enriched terms.
Hierarchical pathway organization
Reactome is structured hierarchically. Individual reactions are grouped into pathways, and pathways are further organized into broader biological categories. This allows users to move seamlessly from global themes, such as immune signaling or metabolism, to specific subpathways and individual molecular events.
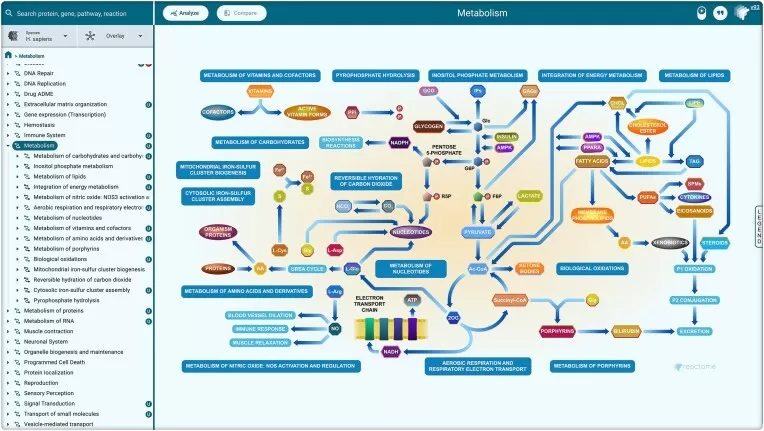
Figure 2. Hierarchical Pathway Organization in Reactome. Image reproduced from Ragueneau, E., Gong, C., Sinquin, P. et al., 2026, Nucleic acids research, licensed under the Creative Commons Attribution License (CC BY 4.0).
Reaction-centered biological logic
Unlike databases built primarily around static pathway maps, Reactome uses a reaction-based data model. Each reaction can include inputs, outputs, catalysts, regulators, and compartment information. This makes it especially helpful for interpreting signaling cascades, transport processes, and other dynamic biological systems.
Data coverage
Reactome is curated primarily for human biology, and knowledge is extended to selected non-human species through orthology-based inference. This human-centered design is one of its major strengths for biomedical research, although it also means that users working in non-human systems should interpret inferred pathways with appropriate caution.
1.2 How to Access and Use Reactome
Reactome can be accessed in several researcher-friendly ways, making it useful for both beginners and advanced users. New users can start with the Reactome website and Pathway Browser to explore pathways visually and understand how molecules are connected in specific biological processes. Researchers who already have gene, protein, or metabolite lists can use Reactome's analysis tools to perform pathway enrichment and expression-based analysis directly. For more customized or large-scale workflows, Reactome also provides REST APIs, R packages such as ReactomePA and ReactomeGSA, as well as downloadable pathway files with identifier mapping resources. In practice, this means Reactome is flexible enough for quick interactive exploration while also supporting more advanced computational analysis in omics research.
2. Applications of Reactome in Omics Research
Reactome is highly versatile and can support a wide range of omics applications. Its value, however, differs slightly depending on the data type and the biological question.
2.1 Reactome in Transcriptomics
In transcriptomics, Reactome is commonly used to map differentially expressed genes or ranked gene lists onto curated biological pathways. This makes it well suited to RNA-seq and microarray studies, especially when researchers want to move beyond gene-level output and identify which biological programs are changing in a coordinated way.
Two common strategies are used here:
- Over-representation analysis (ORA), which tests whether a set of significant genes is enriched in particular pathways
- Gene set enrichment analysis (GSEA), which evaluates whether genes from a pathway tend to cluster at the top or bottom of a ranked gene list
Reactome is particularly useful in transcriptomics when the goal is mechanistic interpretation rather than simple term enrichment, because its hierarchical structure helps distinguish broad biological themes from more specific pathway branches.
2.2 Reactome in Proteomics
Reactome is also highly effective for proteomics, especially in studies focused on signaling, trafficking, immune biology, cell cycle control, and other protein-driven processes. Because Reactome explicitly represents proteins, complexes, catalysts, and regulators, enrichment analysis of differential proteins often yields pathway-level results that are easier to interpret mechanistically.
This is especially relevant for modified proteins, such as phosphoproteins or ubiquitinated proteins. In many cases, Reactome's reaction-based framework is better aligned with proteomics than a purely gene-centered model, because it reflects the fact that biological regulation often depends on protein state, localization, and interaction context.
2.3 Reactome in Metabolomics
Reactome can also be applied to metabolomics, especially when metabolites can be confidently mapped to standard identifiers such as ChEBI. It is most useful for targeted metabolomics or for datasets with relatively high-confidence compound annotation.
That said, Reactome should not always be treated as a standalone solution for untargeted metabolomics. In practice, it is often most informative when metabolite-level changes are interpreted together with pathway enzymes, transporters, or transcript/protein data from the same biological system.
2.4 Reactome for Multi-Omics Integration
Reactome is particularly powerful in multi-omics research because it helps integrate molecular changes from different data layers into a unified pathway-level view. When transcriptomics, proteomics, and metabolomics are analyzed separately, researchers often obtain partially overlapping findings that are difficult to interpret as a single biological story. Reactome addresses this challenge by mapping these signals onto shared pathways, making it easier to identify biological processes that are consistently perturbed across the system. A key tool in this context is ReactomeGSA (Gene Set Analysis), which was developed for comparative pathway analysis across multi-omics datasets and can help reveal pathway-level patterns that are more robust and biologically informative than results derived from any single omics layer alone. In practical terms, this means a pathway supported by coordinated changes in gene expression, protein abundance or modification, and metabolite levels is often more compelling than one identified from only one data type.
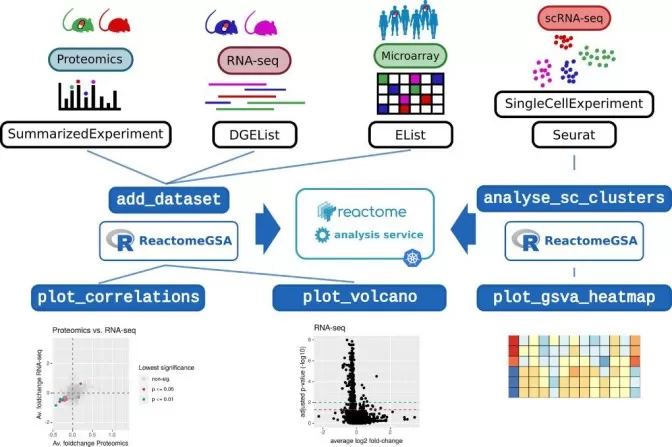
Figure 3. Multi-Omics Pathway Integration Using ReactomeGSA. Image reproduced from Griss, J., Viteri, G., Sidiropoulos, K. et al., 2020, Molecular & cellular proteomics : MCP, licensed under the Creative Commons Attribution License (CC BY 4.0).
A useful example is a cancer drug-resistance study. RNA-seq may identify altered expression of genes involved in MAPK signaling, phosphoproteomics may capture activation of upstream regulators, and metabolomics may reveal downstream metabolic rewiring associated with treatment adaptation. Viewed separately, these datasets provide valuable but incomplete information. When interpreted together through a Reactome-based multi-omics workflow—particularly with tools such as ReactomeGSA—they can highlight pathways that show coordinated changes across multiple molecular layers, providing a more integrated and systems-level view of the resistance mechanism.
3. Reactome vs. KEGG: How They Compare and Complement Each Other
Reactome and KEGG are both widely used pathway databases, and both are commonly used for enrichment analysis. However, they are built with different priorities, and in practice they are often most powerful when used together.
3.1 Similarities in Pathway Analysis
Both Reactome and KEGG:
- organize biological knowledge into pathways
- support enrichment analysis
- help interpret gene, protein, or metabolite lists
- are widely used in omics workflows
3.2 Major Differences in Design and Use
The main difference is conceptual. Reactome is reaction-centered and human-focused. It is especially strong in mechanistic detail, signaling logic, regulatory relationships, and hierarchical pathway structure. KEGG is often more map-centered and broadly organism-oriented. It has long been especially useful for metabolism, organismal pathway mapping, and cross-species pathway interpretation.
Table 1. Reactome vs. KEGG: A Practical Comparison.
| Aspect | Reactome | KEGG |
|---|---|---|
| Core design | Reaction-based, mechanistic, event-centered | Map-based, reference pathway oriented |
| Main strength | Signaling, regulation, reaction logic, pathway hierarchy | Metabolism, organism-wide mapping, broad pathway coverage |
| Species strategy | Primarily human, with inferred pathways for selected non-human species | Broad organism coverage across many taxa |
| Curation emphasis | Expert manual curation with pathway detail | Curated reference maps with strong systems-level coverage |
| Visualization style | Interactive browser, hierarchical views, detailed event-level diagrams | Canonical pathway maps and organism-specific overlays |
| Best use case | Human-centric mechanistic interpretation | Broad functional overview, especially for metabolism and comparative studies |
3.3 When to Use Reactome, KEGG, or Both
For many omics projects, the most informative approach is not choosing Reactome or KEGG, but using Reactome alongside KEGG.
A practical strategy is:
- use Reactome when you want detailed mechanistic interpretation, especially for signaling and regulation
- use KEGG when you want broad pathway coverage, especially in metabolism or cross-species studies
- compare results across both resources to increase confidence and identify consistent biological themes
When the same pathway-level conclusion appears across databases, the result is often more convincing. When they differ, the discrepancy can still be informative, because it may reflect differences in pathway granularity, curation philosophy, or species modeling.
4. Reactome Pathway Analysis Visualization and Result Interpretation
Generating a list of enriched Reactome pathways is only the first step in pathway analysis. The more important task is learning how to read those results in a biologically meaningful way. In practice, pathway interpretation depends not only on statistical significance, but also on pathway size, the number of mapped molecules, the hierarchical structure of Reactome, and the broader experimental context. Different visualization formats highlight different aspects of the data, so choosing the right plot can make a major difference in how clearly the results are understood. Bar charts are useful for rapid ranking, dot plots provide a more balanced view of significance and coverage, and network-style visualizations help reveal pathway overlap, shared molecular drivers, and potential regulatory connections. Used together, these visual formats can turn a long enrichment table into a more coherent biological story.
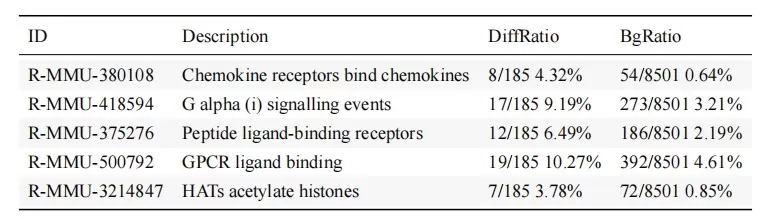
Figure 4. Example Reactome Pathway Enrichment Results for Differentially Expressed Genes.
4.1 Bar Chart Interpretation
A bar chart is often the most straightforward way to summarize Reactome pathway enrichment results. In most analyses, pathways are ranked by a statistic such as -log10(p-value), adjusted p-value, or enrichment score, allowing researchers to quickly identify the most significantly enriched biological processes in the dataset. Because of its simplicity and strong visual contrast, the bar chart is especially useful for initial screening and prioritization. However, statistical significance alone should not be overinterpreted. A highly ranked pathway should still be evaluated in the context of pathway size, the number of matched genes or proteins, the experimental background, and its position within the Reactome hierarchy. In many cases, a more specific child pathway may provide clearer biological insight than a broad parent category.
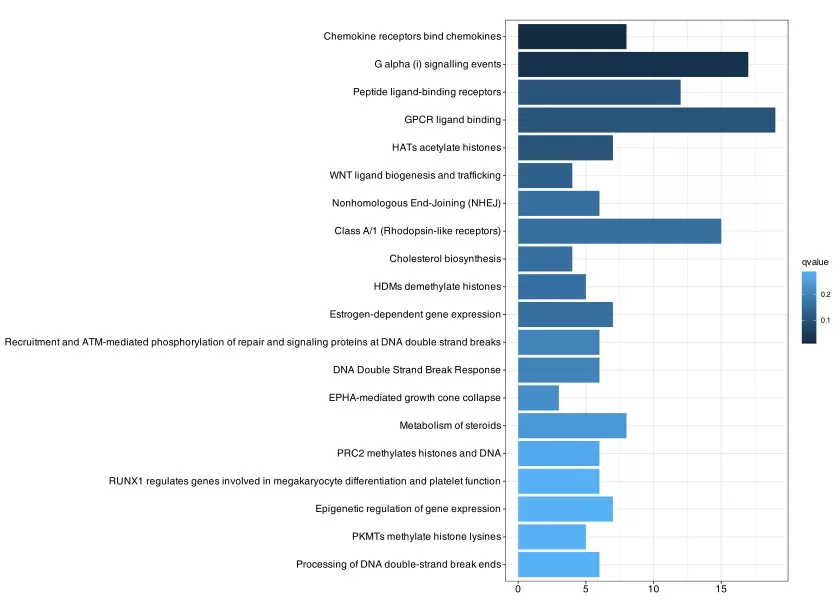
Figure 5. Bar Chart of Enriched Reactome Entries in Differentially Expressed Genes. The x-axis represents the number of differentially expressed genes annotated to each Reactome term, while the y-axis shows the names of the Reactome terms. Darker colors indicate more significant enrichment.
4.2 Dot Plot Interpretation
Compared with a bar chart, a dot plot provides a more information-rich view of Reactome enrichment results by displaying pathway significance and pathway coverage simultaneously. In a typical dot plot, the size of each dot reflects the number of mapped genes or proteins, while color indicates significance, often represented by adjusted p-value. This makes dot plots particularly useful for identifying pathways that are not only statistically significant but also broadly represented in the dataset. In practice, researchers should look for pathways that show a good balance between significance and coverage. A small but highly significant pathway may indicate a focused regulatory event, whereas a larger pathway with moderate significance may reflect a broader biological shift. As a result, dot plots are well suited for refining candidate pathways after an initial round of ranking.
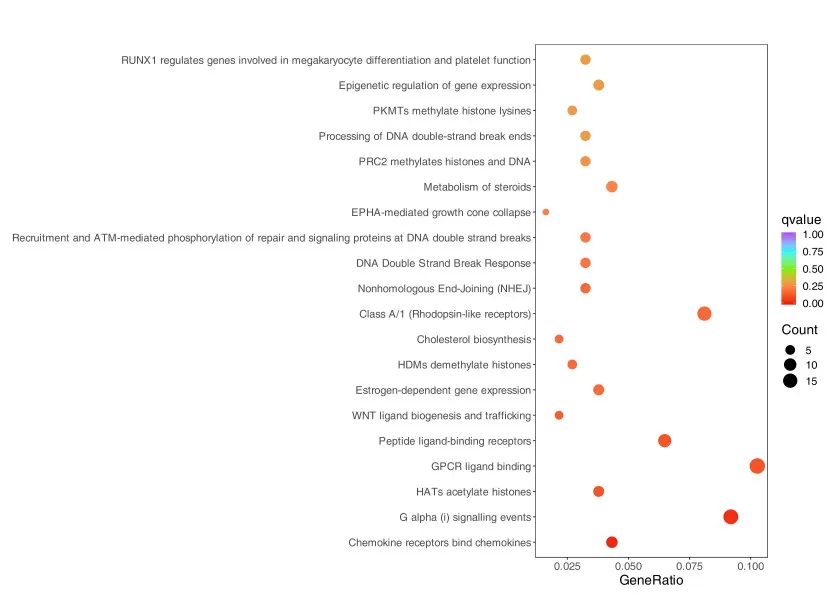
Figure 6. Reactome Enrichment Results Dot Plot. The y-axis represents the Reactome terms, and the x-axis represents the GeneRatio. A larger GeneRatio indicates a higher degree of enrichment. Larger dots represent a greater number of differentially expressed genes enriched in the pathway, while redder dots indicate more significant enrichment.
4.3 Chord Plot and Network Interpretation
Chord plots and network-based visualizations are especially valuable when the goal is to understand how pathways are connected rather than simply how they are ranked. These visualizations highlight shared genes or proteins across enriched pathways, making it easier to identify overlap, cross-talk, and central regulatory nodes. This is particularly important when several pathways appear enriched because of a common set of driving molecules. In such cases, the analysis can move beyond a list of pathway names toward a more integrated view of biological organization. Researchers should pay special attention to hub genes or proteins, pathway overlap patterns, and, when quantitative information is included, the direction and consistency of molecular changes. In multi-omics studies, network-style visualization is even more informative because it can reveal whether different omics layers converge on the same pathway structure or highlight distinct parts of a biological network.
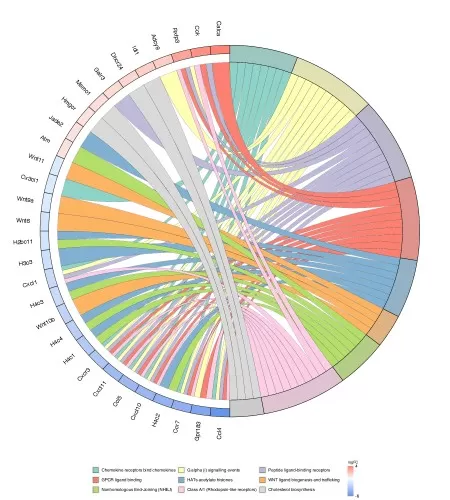
Figure 7. Reactome Enrichment Chord Plot. The 9 most significantly enriched Reactome entries are shown on the right side of the figure, the 90 genes with the largest fold change |logFC| in these 9 Reactome entries are shown on the left side, and the middle line represents the correspondence between pathways and genes.
4.4 Best Practices for Reading Reactome Results
The choice of visualization should always depend on the biological question. Bar charts are most useful for rapid ranking, dot plots are better for evaluating significance together with pathway coverage, and chord plots or network views are more suitable for exploring pathway connectivity and shared molecular drivers. In practice, a staged workflow often works best: researchers can begin with a bar chart to identify major candidate pathways, use a dot plot to assess the breadth of pathway representation, and then move to network-style visualizations to examine pathway relationships in greater depth. It is also important to interpret Reactome results in light of pathway hierarchy, since significant high-level categories may be driven by only one or two specific subpathways. Whenever possible, key findings should be cross-checked with other pathway resources such as KEGG, particularly in metabolism-focused or cross-species studies. Most importantly, pathway interpretation should not rely on p-values alone; robust conclusions require statistical significance to be considered together with pathway coverage, experimental context, and biological plausibility.
5. Key Takeaways and Practical Tools for Reactome Pathway Analysis
Reactome is one of the most valuable pathway-analysis resources available to omics researchers because it combines expert curation, mechanistic depth, hierarchical organization, and flexible tools for data interpretation. Its reaction-centered framework is particularly useful for transcriptomics, proteomics, and multi-omics studies, where researchers need to move beyond lists of altered molecules and understand how those changes fit into broader biological processes. At the same time, its limitations should also be recognized: Reactome is primarily optimized for human biology, non-human pathway annotations often rely on orthology-based inference, and metabolomics applications are most informative when compound identities can be assigned with confidence. In addition, its depth and complexity can create a steeper learning curve for new users. Even so, as omics research continues to shift toward integrative and systems-level analysis, Reactome is likely to become even more important for pathway-level interpretation, disease mechanism studies, multi-omics integration, and the generation of biologically grounded hypotheses for downstream validation. Ultimately, Reactome is most powerful when used not simply as an enrichment resource, but as a framework for linking omics data to mechanism, helping researchers move from molecular change to biological insight.
MetwareBio: Your Trusted Partner for Multi-Omics Data Analysis
For researchers seeking a faster and more accessible way to perform Reactome pathway analysis, the Metware Cloud Platform offers a practical one-click solution for streamlined data interpretation. Backed by an experienced multi-omics bioinformatics team, the platform supports free registration, enables users to upload data in the required format, and provides access to more than 50 omics analysis tools, together with step-by-step text instructions and video tutorials, making Reactome-based pathway exploration easier and more efficient.
MetwareBio's Multi-Omics Integration Analysis services provide comprehensive support for transcriptomics, proteomics, and metabolomics pathway analysis, helping researchers translate complex multi-layer data into actionable biological insights. If you are interested in multi-omics pathway analysis, please do not hesitate to contact us.
Contact UsRead More: Pathway Analysis and Omics Data Interpretation
Pathway analysis is a cornerstone of omics research, bridging the gap between molecular data and biological meaning. These related articles explore complementary enrichment methods, visualization techniques, and multi-omics integration strategies to help you build a more complete analytical workflow.
Understand the key differences between GO, KEGG, and GSEA enrichment methods, and learn when to apply each one depending on your research question, data type, and desired level of biological interpretation.
A practical introduction to KEGG pathway analysis covering database navigation, enrichment workflows, and result interpretation—the ideal companion to Reactome for cross-database validation.
Learn how GSEA differs from ORA, how to prepare ranked gene lists, and how to interpret enrichment scores and leading-edge subsets in your transcriptomics and proteomics studies.
Discover how to combine multiple omics layers into a unified biological narrative using pathway-level integration approaches, including ReactomeGSA for comparative multi-omics pathway analysis.
Explore CCA as a complementary statistical method for identifying cross-omics associations, helping you uncover coordinated molecular changes that pathway enrichment alone may not reveal.
Before running pathway enrichment, you need to identify significantly changed molecules. This guide covers how to create and interpret volcano plots for differential analysis in metabolomics and proteomics.
References
- Ragueneau, E., Gong, C., Sinquin, P., Sevilla, C., Beavers, D., Grentner, A., Griss, J., Hogue, G. F. J., Li, N. T., Matthews, L., May, B., Milacic, M., Mohammadi, H., Petryszak, R., Rothfels, K., Shamovsky, V., Stephan, R., Tiwari, K., Weiser, J., Wright, A., & D'Eustachio, P. (2026). The Reactome Knowledgebase 2026. Nucleic acids research, 54(D1), D673–D681. https://doi.org/10.1093/nar/gkaf1223
- Griss, J., Viteri, G., Sidiropoulos, K., Nguyen, V., Fabregat, A., & Hermjakob, H. (2020). ReactomeGSA - Efficient Multi-Omics Comparative Pathway Analysis. Molecular & cellular proteomics : MCP, 19(12), 2115–2125. https://doi.org/10.1074/mcp.TIR120.002155


