Label-free quantification (LFQ) is one of the most widely used quantitative strategies in proteomics. Owing to its label-free design, flexible sample handling, relatively low cost, and scalability for larger sample cohorts, LFQ is broadly applied in biomarker discovery, disease research, and functional proteomics studies. The workflow is built on high-resolution mass spectrometry data and converts peptide-level signals into protein-level abundance estimates through a series of computational steps. This article explains the complete LFQ workflow, from MS1 signal detection to final protein quantification, with a focus on the analytical principles behind each step.
1. LFQ Fundamentals: What Does Label-Free Quantification Measure?
LFQ is not a mass spectrometry acquisition mode, but rather a quantification strategy. In practice, LFQ data are most commonly generated using data-dependent acquisition (DDA). As illustrated in Figure 1, the mass spectrometer records continuous peptide signal intensities at the MS1 level and then selects a subset of precursor ions, typically using a topN strategy, for fragmentation at the MS2 level to enable peptide identification. LFQ is therefore built on the integration of MS1 signal information anchored by MS2-based peptide identification.
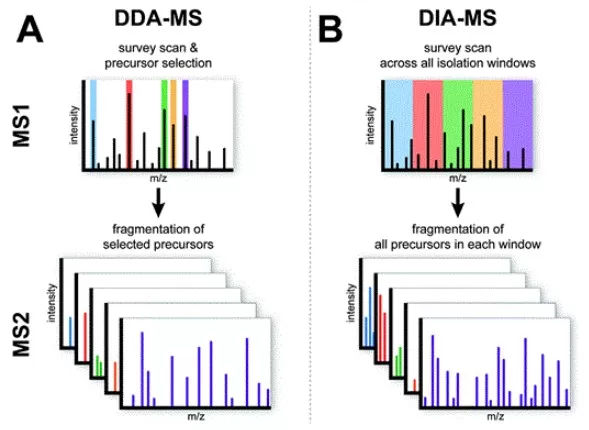
Figure 1. Schematic comparison of DDA-MS and DIA-MS acquisition modes. Left: In DDA, a subset of precursor ions is selected for MS2 fragmentation after each MS1 survey scan. Right: In DIA, all precursors within predefined m/z windows are systematically fragmented. Image reproduced from Krasny & Huang, 2021, Molecular Omics, 17(1), 29–42.
The quantitative foundation of LFQ mainly relies on two strategies, which together connect raw mass spectrometric signals to protein-level quantitative information.
1.1 Main LFQ Quantification Strategies
| Strategy | Principle | Strengths | Limitations |
|---|---|---|---|
| MS1 intensity-based quantification | Estimates relative peptide abundance by measuring chromatographic peak area or ion intensity in MS1. High-resolution MS1 data allow accurate extraction of peptide XICs, and the integrated area is proportional to peptide abundance. | Currently the dominant LFQ strategy; widely implemented in MaxQuant, Proteome Discoverer, FragPipe, and related platforms. | Highly dependent on robust feature detection, peak integration, and alignment quality. |
| Spectral counting-based quantification | Infers protein abundance from the number of times a peptide is fragmented and successfully identified in MS/MS, typically expressed as peptide-spectrum match (PSM) counts. | Useful for preliminary screening and for comparing relatively abundant proteins. | Much less stable for low-abundance proteins and generally less quantitative than intensity-based methods. |
2. Label-Free Quantification Workflow
Although LFQ can be implemented through both MS1 intensity-based quantification and spectral counting, these two strategies differ substantially in analytical depth and practical use. For a step-by-step explanation of LFQ data analysis, MS1 intensity-based LFQ provides the more representative and informative framework, because it captures the core computational workflow that links peptide identification, feature alignment, signal extraction, normalization, and protein-level quantification. Since this strategy is most commonly performed in a DDA-based setting, the following sections focus specifically on DDA-based, MS1 intensity-based label-free quantification (LFQ) and explain its major analytical steps in detail.
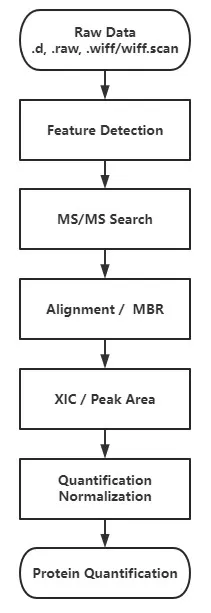
Figure 2. General database search workflow for MS/MS-based peptide identification.
Step 1: Spectral Interpretation and Peptide Feature Detection
This is the starting point for mass spectrometry data processing. The key question at this stage is how to identify and track each peptide-derived signal unit across continuous MS1 scans. These signal units are referred to as features. An MS1 feature is a stable signal entity in three-dimensional m/z-RT-intensity space and is typically defined by the following parameters:
- m/z: mass-to-charge ratio
- Retention time (RT): chromatographic retention time
- Intensity: ion signal intensity
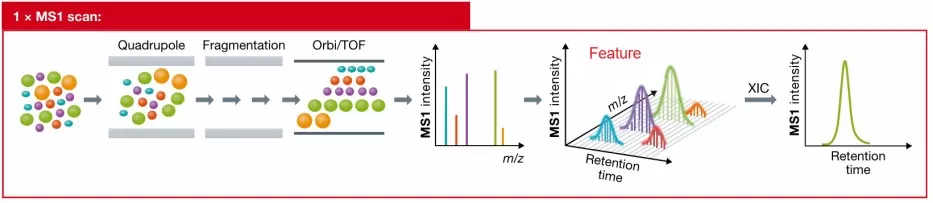
Figure 3. MS1 features and the XIC curve. Image reproduced from Ludwig et al., 2018, Molecular Systems Biology, 14(8), e8126.
At the implementation level, feature detection generally includes centroiding, noise filtering, isotope clustering, charge-state assignment, and signal linking across scans. Together, these steps generate MS1 features that can subsequently be used for matching and quantification.
Step 2: Peptide and Protein Identification (MS/MS Search)
This is the core qualitative stage of the workflow. The central question here is which peptides and proteins the detected MS1 features actually represent. In a DDA label-free workflow, this is achieved by matching tandem mass spectra (MS/MS) against a protein sequence database.
From a computational perspective, the qualitative stage takes as input:
- MS/MS spectra
- Associated precursor information, including m/z, charge state, and RT, which are derived from the corresponding MS1 features
The identification process can be summarized in three major steps.
Theoretical-to-Experimental Spectrum Matching
First, theoretical peptides are generated by performing in silico digestion of the protein database while accounting for allowed missed cleavages and defined modification settings, including both fixed and variable modifications. Next, theoretical fragment ions, such as b and y ions, are calculated according to fragmentation rules. Finally, the experimental MS/MS spectra are compared against the predicted fragment patterns, and a score is assigned based on the degree of agreement between experimental and theoretical spectra.
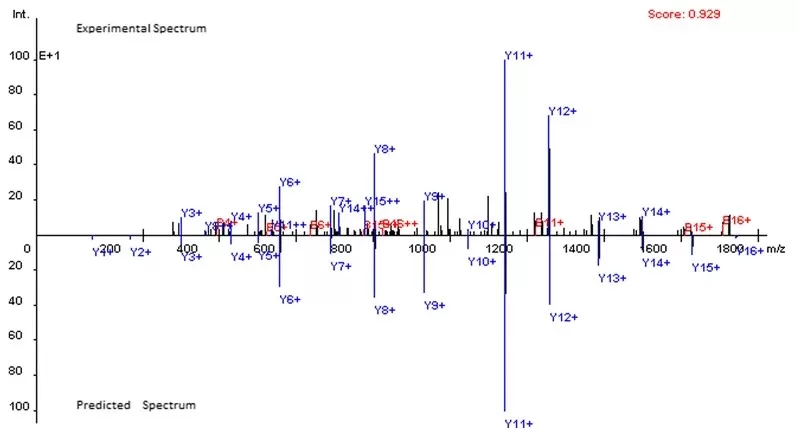
Figure 4. Comparison of the experimental spectrum and the predicted spectrum for peptide EIELEDPLENMGAQMVK. Image reproduced from Wang et al., 2015, BMC Bioinformatics, 16, 110.
Identification Confidence and FDR Control
Because the search space in proteomics is extremely large, random matches are unavoidable. For this reason, all mainstream database search engines apply statistical filtering procedures, most commonly including:
- the target-decoy strategy
- false discovery rate (FDR) control at the PSM, peptide, and protein levels
Only identifications that pass the defined FDR threshold are retained for downstream quantitative analysis.
Protein Inference
Once peptides have been identified, peptide-level evidence must be consolidated at the protein level. In practice, shared peptides, which can map to more than one protein, are used to construct the minimal explanatory protein set, often referred to as a protein group.
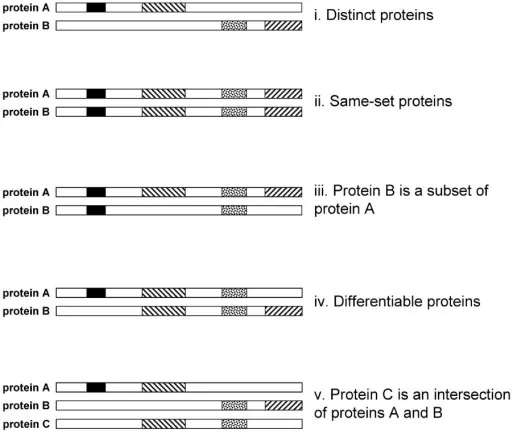
Figure 5. Principle of protein inference. According to the parsimony principle, protein A and protein B are retained, whereas protein C is downgraded or removed from the final report. Image reproduced from Koskinen et al., 2011, Molecular & Cellular Proteomics, 10(6), M110.003822.
Step 3: Cross-Run Alignment and Match Between Runs
Two major challenges arise in DDA-based label-free analysis. First, the retention time of the same peptide may shift systematically across runs because of factors such as column aging, flow-rate variation, or small gradient drifts. Second, stochastic precursor selection in DDA can lead to missing MS/MS events, meaning that a peptide fragmented in one run may not be selected for fragmentation in another.
To address these issues, cross-run alignment generally includes two components:
- Cross-Run RT Alignment: Retention time alignment establishes a common coordinate system across runs. Typical strategies include selecting anchor features that appear reproducibly across multiple runs, usually high-intensity peptides, fitting a linear or nonlinear RT mapping function, and projecting raw RT values from each run into a shared aligned RT space. This step makes it possible to determine whether a feature observed at RT = 34.8 min in Run A corresponds to the same peptide as a feature observed at RT = 36.1 min in Run B.
- Match Between Runs: Once alignment has been performed, features can be matched across runs in aligned RT-m/z space. The core principle is straightforward: if a peptide has been identified by MS/MS in Run A and therefore has a defined m/z and aligned RT window, then an MS1 feature in Run B with matching m/z and RT can inherit that peptide identification even if it was not fragmented in Run B.
It is important to note that missing values in LFQ are not caused solely by stochastic DDA sampling. They are the combined result of RT alignment quality, feature detection performance, and feature matching strategy.
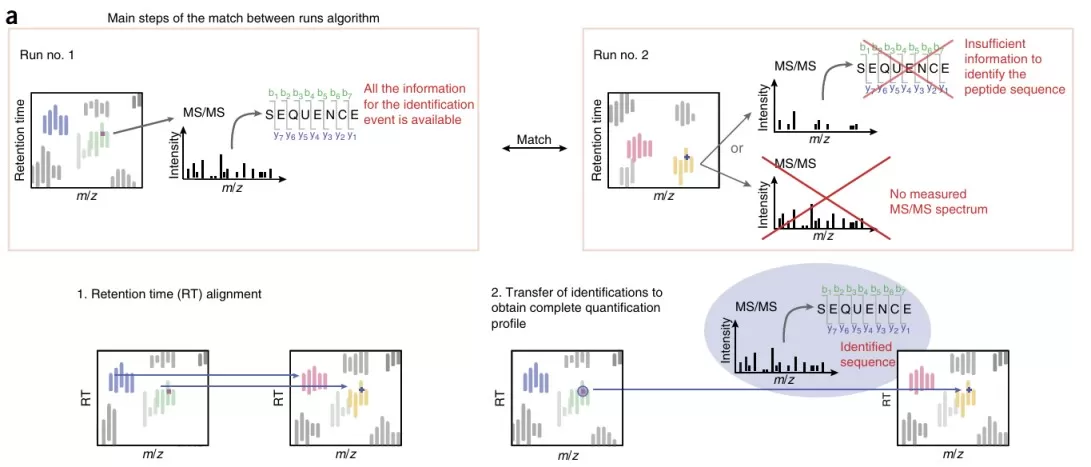
Figure 6. Match between runs algorithm. Image reproduced from Tyanova et al., 2016, Nature Protocols, 11(12), 2301–2319.
Step 4: Extracted Ion Chromatograms (XICs) and Peptide Quantification
This step is the core of quantitative analysis. The main question is how a final quantitative value for a peptide in a single sample is derived from MS1 data.
An extracted ion chromatogram (XIC) is an intensity-versus-time curve generated by extracting signals from continuous MS1 scans across the RT dimension within a specified m/z tolerance, as illustrated in Figure 3. In practical terms, it represents the chromatographic MS1 peak formed by a peptide over its full elution window.
In peptide quantification, the most commonly used metric is peak area, which captures the cumulative peptide signal across the chromatographic peak. Peak height is used less frequently because it is more sensitive to noise and peak-shape variation. Importantly, XIC integration is performed at the MS1 feature level rather than at the PSM level, because PSMs belong to the identification stage rather than the quantification stage.
Step 5: Quantitative Normalization
Normalization is essential for reducing technical bias. In real experiments, systematic variation may arise from small differences in sample loading, instrument response drift, or changes in ionization efficiency. The goal of normalization is therefore to remove these global biases and place all samples on a comparable quantitative scale. Common normalization strategies include:
- Total intensity / total peptide amount normalization, which scales each sample so that the total peptide signal is equivalent across samples
- Median normalization, which aligns samples by adjusting the median intensity and is generally more robust to outliers
- Ratio-based normalization, which uses the distribution of peptide intensity ratios between samples to correct global scaling differences
Step 6: Protein Quantification
After protein identification and grouping have been established, the next task is to summarize quantified peptide signals into a protein-level abundance value according to predefined rules.
The input for protein quantification includes the established protein groups, the quantified peptides assigned to each group, and the peptide intensity values measured in each sample, typically as XIC peak areas or equivalent metrics.
The choice of peptides used for protein quantification is critical. Unique peptides map exclusively to a single protein group and therefore provide the clearest evidence. Shared peptides are shared across multiple proteins but are assigned to the protein group supported by the strongest overall evidence. Additional quality filters are often applied to remove peptides with high missingness, unusually large between-sample variability, or poor signal quality.
Peptide-level intensities can then be combined into protein abundance values using several strategies, including summation, mean or median aggregation, top N peptide selection, or ratio-driven methods such as MaxLFQ. From a data-structure perspective, protein quantification is essentially a mapping and summarization problem that converts Peptide × Sample information into Protein × Sample abundance estimates.
3. Conclusion: LFQ as a Chain of Quantitative Evidence
When the full DDA label-free workflow is considered as a whole, it becomes clear that LFQ is not the output of a single quantification algorithm. Instead, it is a stepwise computational process in which each stage depends strongly on the reliability of the previous one.
MS1 feature detection determines which signals exist and can be quantified. MS/MS identification assigns those features a credible biological identity. Cross-run alignment and Match Between Runs ensure that the same peptide is treated as the same analytical entity across multiple runs. XIC integration provides a quantitative estimate of peptide abundance within each sample. Normalization places all samples onto a common quantitative scale. Finally, protein quantification summarizes multiple peptide-level measurements into a protein-level abundance estimate under an established protein grouping framework.
In this sense, label-free quantification is not simply a value produced by software. It is the natural outcome of correctly interpreting MS1-derived evidence at every stage of the analysis pipeline.
Support Your LFQ Proteomics Research with Reliable Data Analysis
LFQ proteomics depends on a complete chain of evidence, from MS1 signal extraction to protein-level abundance estimation. MetwareBio provides quantitative proteomics services that support DDA-based label-free quantification, differential protein expression analysis, pathway interpretation, and multi-omics integration.
If you need reliable LFQ data analysis for biomarker discovery, disease research, or functional proteomics studies, contact us to discuss your project.
Read More: Proteomics Quantification and Data Analysis
After understanding the LFQ workflow from MS spectra to protein abundance, explore these related articles covering acquisition strategies, quantitative methods, downstream statistical analysis, and multi-omics integration in proteomics research.
Once you have protein abundance estimates from LFQ, volcano plots help visualize differential expression results by combining fold-change and statistical significance. Learn how to set appropriate thresholds and interpret volcano plot patterns for proteomics datasets.
LFQ quantifies proteins at the group level, but biologically distinct proteoforms—arising from PTMs, splice variants, and genetic variation—may behave differently. Understand what proteoforms are, why they matter, and which detection strategies complement label-free quantification.
After normalizing protein abundance data with LFQ, multivariate methods like PLS-DA can reveal group separation patterns. This guide covers PLS-DA principles, validation strategies, and practical tips applicable to both metabolomics and proteomics data.
Building on PLS-DA fundamentals, learn when OPLS-DA offers advantages for separating predictive variation from orthogonal noise. A practical comparison to help you select the right method for your quantitative proteomics or metabolomics datasets.
LFQ is a key quantitative strategy in clinical proteomics for biomarker discovery and patient stratification. Explore how quantitative proteomics methods, including label-free approaches, are being translated into precision medicine applications and the challenges involved.
Quantitative proteomics data from LFQ workflows often gain deeper biological meaning when integrated with metabolomics. See how multi-omics strategies combining protein and metabolite abundance data drive discoveries in plant science and agricultural research.
References
- Krasny, L., & Huang, P. H. (2021). Data-independent acquisition mass spectrometry (DIA-MS) for proteomic applications in oncology. Molecular Omics, 17(1), 29–42. https://doi.org/10.1039/d0mo00072h
- Ludwig, C., Gillet, L., Rosenberger, G., Amon, S., Collins, B. C., & Aebersold, R. (2018). Data-independent acquisition-based SWATH-MS for quantitative proteomics: a tutorial. Molecular Systems Biology, 14(8), e8126. https://doi.org/10.15252/msb.20178126
- Wang, Y., Yang, F., Wu, P., Bu, D., & Sun, S. (2015). OpenMS-Simulator: an open-source software for theoretical tandem mass spectrum prediction. BMC Bioinformatics, 16, 110. https://doi.org/10.1186/s12859-015-0540-1
- Koskinen, V. R., Emery, P. A., Creasy, D. M., & Cottrell, J. S. (2011). Hierarchical clustering of shotgun proteomics data. Molecular & Cellular Proteomics, 10(6), M110.003822. https://doi.org/10.1074/mcp.M110.003822
- Tyanova, S., Temu, T., & Cox, J. (2016). The MaxQuant computational platform for mass spectrometry-based shotgun proteomics. Nature Protocols, 11(12), 2301–2319. https://doi.org/10.1038/nprot.2016.136


