In multi-group omics analysis, the real challenge is rarely whether you remembered to run ANOVA. The bigger issue is whether the result is trustworthy enough to support what comes next: feature selection, pathway enrichment, biomarker ranking, and biological interpretation. When group variances differ, sample sizes are uneven, or the data have strong skew and outliers, a default one-way ANOVA can look formal while still answering the wrong question. In practice, the key issue is whether the selected test truly matches the data structure. To clarify this, this article compares ordinary ANOVA, Welch ANOVA, and Kruskal-Wallis for multi-group omics data, especially in situations involving unequal variances, unbalanced sample sizes, skewed distributions, or outliers.
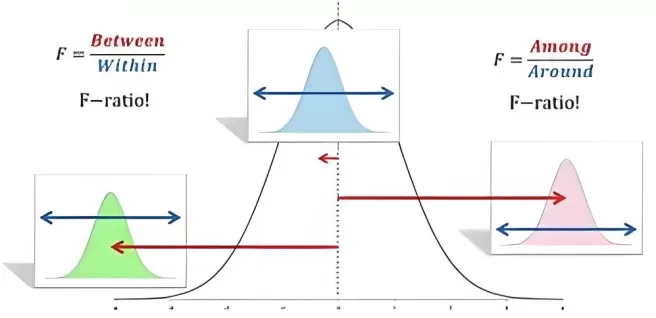
Figure 1. ANOVA Analysis of Variance.
1. MULTI-GROUP OMICS IS NOT JUST A LARGER TWO-GROUP PROBLEM
Multi-group designs are routine in omics: disease stage I/II/III/IV, multiple dose levels, tissue compartments, or distinct spatial niches such as tumor core, invasive margin, stroma, and immune-rich regions. But once the number of groups increases, the analysis becomes more fragile. Unequal variance is easier to miss, sample sizes are more likely to drift out of balance, and the number of pairwise follow-up comparisons grows quickly. In proteomics workflows, this is often compounded by multiple runs, missing values, and technical variation across batches [1]. So the main question is not simply whether an omnibus p-value is significant. It is whether that p-value reflects the biology of interest or a mismatch between the test and the structure of the data.
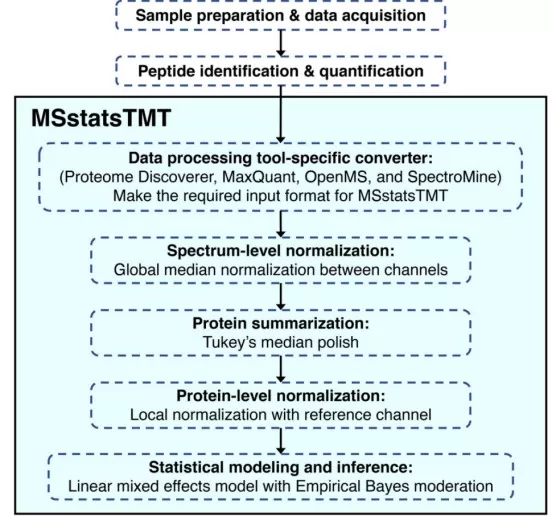
Figure 2. Statistical Detection of Differentially Abundant Proteins in Experiments. Image reproduced from Huang et al., 2022, Mol Cell Proteomics, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
2. ANOVA VS WELCH ANOVA VS KRUSKAL-WALLIS: WHAT DOES EACH TEST COMPARE?
Ordinary one-way ANOVA compares group means under the assumption that the groups have similar variances. Welch ANOVA asks the same mean-based question but relaxes that equal-variance assumption, making it a better fit when one group is clearly more variable than another [2]. Kruskal-Wallis takes a different route. It works on ranks rather than raw values and asks whether the groups differ in their overall distributions [3]. That distinction matters. ANOVA and Welch ANOVA are primarily about mean differences. Kruskal-Wallis is more robust to skew and extreme values, but its interpretation is broader, especially when group shapes differ. A significant omnibus result from any of the three methods still needs a suitable post hoc procedure to identify which groups actually differ.
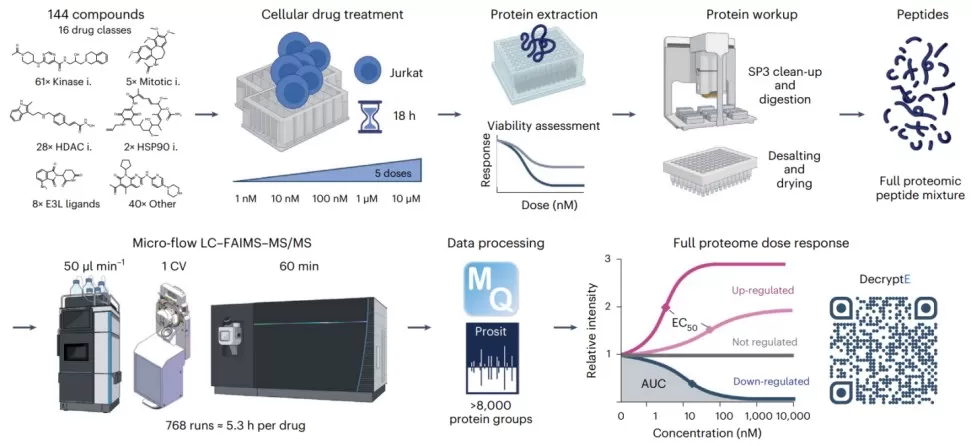
Figure 3. Drug-induced protein expression changes. Image reproduced from Eckert et al., 2025, Nat Biotechnol, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
3. WHY ORDINARY ANOVA CAN MISLEAD OMICS RESULTS
In real omics projects, statistical choices propagate downstream. A different omnibus test can change which proteins, metabolites, or genes are called significant, which in turn changes enrichment results and the biological story built on top of them. That is why variance structure matters so much. If one treatment group is much more variable than the others, a pooled-variance ANOVA can be too optimistic or simply unstable. This is not a rare edge case. In large proteomics pipelines, variation can differ substantially across features and experimental settings [1]. In multi-dose proteomics, regulated proteins can show much larger coefficients of variation than non-regulated proteins, which makes equal-variance assumptions hard to defend across groups [2]. A p-value is only as reliable as the model behind it.
4. REAL EXAMPLES FROM PROTEOMICS, METABOLOMICS, AND SPATIAL OMICS
Published studies make the point clearly. In dose-resolved proteomics, the decryptE study profiled 144 drugs across roughly 8,000 proteins and observed higher variability among regulated proteins than among non-regulated ones [2]. In that setting, Welch ANOVA is often more defensible than ordinary ANOVA when the goal is to compare mean abundance across dose groups. Metabolomics provides a different example. In a lung adenocarcinoma progression study, researchers compared benign lesions and multiple histological stages and identified altered metabolites across four groups [3]. For features with marked skew or uneven spread, a rank-based strategy such as Kruskal-Wallis may be more stable, but the conclusion should be framed as a distributional difference rather than a simple shift in means. Spatial omics adds yet another layer: the key signal may be regional rather than global, so collapsing all measurements into a single average can hide biologically important local enrichment patterns [4].
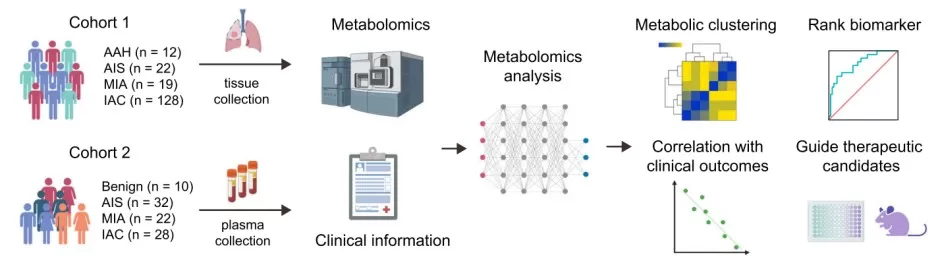
Figure 4. Evolutionary metabolic landscape from preneoplasia to invasive lung adenocarcinoma. Image reproduced from Nie et al., 2021, Nat Commun, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
5. WHY SPATIAL MULTI-OMICS NEEDS EXTRA STATISTICAL CAUTION
Spatial data deserve separate treatment because they violate one of the quiet assumptions behind standard omnibus tests: independence. Nearby spots or cells in a tissue section are often correlated. That means a naive test can underestimate uncertainty and exaggerate significance. Recent work using spatial mixed models showed that accounting for spatial correlation can reduce false positives and produce more credible differential-expression results in spatial transcriptomics [4]. The practical lesson is straightforward. If the biological question is about differences between annotated regions, it is often better to aggregate to biologically meaningful ROIs or patient-level summaries first. If the question is truly spot-level or cell-level, then a spatially aware model is usually more appropriate than treating each nearby observation as independent. In spatial omics, the statistical method should follow the tissue architecture, not ignore it.
6. HOW TO CHOOSE BETWEEN ANOVA, WELCH ANOVA, AND KRUSKAL-WALLIS
As a practical rule, Welch ANOVA is often the safer default for multi-group omics data when you still care about mean differences but cannot justify equal variance across groups. Kruskal-Wallis is useful when the data are strongly skewed or heavily influenced by extreme values, but it should not be treated as a universal substitute for every problematic ANOVA.
Start by deciding what biological question you want the statistics to answer. If you want to compare means and the groups differ in variance or sample size, Welch ANOVA is usually a better default than ordinary ANOVA. If the data are highly skewed or the signal is better represented by ranks, Kruskal-Wallis may be more appropriate, but the interpretation should stay aligned with that choice. In all cases, the omnibus test is only one part of the workflow. Good practice also requires sensible preprocessing, QC, normalization, missing-value handling, and post hoc testing.
| Method | Best used when | Main strength | Main caution |
|---|---|---|---|
| Ordinary ANOVA | Group variances are reasonably similar and the design is fairly balanced | Direct test of mean differences across groups | Can be unreliable when heteroscedasticity and uneven n are present |
| Welch ANOVA | Variances differ across groups or sample sizes are unbalanced | More robust mean comparison under unequal variances | Still answers a mean-based question |
| Kruskal-Wallis | Data are strongly skewed, outlier-prone, or better treated as ranks | More robust to non-normality and extreme values | Omnibus only, and interpretation is about rank/distribution differences rather than means |
7. CONCLUSION
Ordinary ANOVA is not obsolete, but in omics it is often too brittle to serve as the automatic default. Welch ANOVA is frequently the more credible choice for mean-based multi-group comparisons because real datasets rarely satisfy equal-variance assumptions cleanly. Kruskal-Wallis remains valuable when robustness is the priority, especially for skewed or rank-like data, but it answers a different question. For spatial multi-omics, the bar is even higher: the analysis has to respect tissue structure and spatial dependence. In other words, the right test is not the one that is most familiar. It is the one that preserves the biological meaning of the result.
Need a more reliable statistical workflow for multi-group omics data?
Choosing the right test is only one part of getting trustworthy biological results. MetwareBio supports proteomics, metabolomics, lipidomics, spatial omics, and multi-omics studies with integrated data analysis workflows designed to improve result interpretation and downstream discovery.
Explore our multi-omics analysis services or contact us to discuss your project.
Contact UsRead More: Related Guides for Omics Statistical Analysis
Want to build a more reliable omics analysis workflow beyond a single omnibus test? These related articles cover broader statistical test selection, two-group comparisons, feature prioritization, and FDR control for downstream interpretation.
Statistical Tests for Differential Protein Expression in Proteomics
Get a broader view of the major statistical tests used in proteomics and how method selection shapes differential expression results.
T-Test vs Welch's T-Test vs Mann–Whitney U: Which Test Should You Use in Omics?
Learn how to choose the right method for two-group omics comparisons when variance, distribution, and robustness all matter.
Differential Feature Screening in Omics: Why the Best Candidate Is Not Always the One with the Smallest p-Value
See why p-values alone are not enough when ranking proteins or metabolites for downstream interpretation and biomarker discovery.
FDR Control in Proteomics: Principles, Calculation Methods, and Threshold Selection
Follow up with a practical guide to multiple-testing correction and more credible significance threshold setting in omics studies.
References
- Huang T, Choi M, Tzouros M, Golling S, Pandya NJ, Banfai B, Dunkley T, Vitek O. MSstatsTMT: Statistical Detection of Differentially Abundant Proteins in Experiments with Isobaric Labeling and Multiple Mixtures. Mol Cell Proteomics. 2020 Oct;19(10):1706-1723. https://doi.org/10.1074/mcp.RA120.002105
- Eckert S, Berner N, Kramer K, Schneider A, Muller J, Lechner S, Brajkovic S, Sakhteman A, Graetz C, Fackler J, Dudek M, Pfaffl MW, Knolle P, Wilhelm S, Kuster B. Decrypting the molecular basis of cellular drug phenotypes by dose-resolved expression proteomics. Nat Biotechnol. 2025 Mar;43(3):406-415. https://doi.org/10.1038/s41587-024-02218-y
- Nie M, Yao K, Zhu X, Chen N, Xiao N, Wang Y, Peng B, Yao L, Li P, Zhang P, Hu Z. Evolutionary metabolic landscape from preneoplasia to invasive lung adenocarcinoma. Nat Commun. 2021 Nov 10;12(1):6479. https://doi.org/10.1038/s41467-021-26685-y
- Ospina OE, Soupir AC, Manjarres-Betancur R, Gonzalez-Calderon G, Yu X, Fridley BL. Differential gene expression analysis of spatial transcriptomic experiments using spatial mixed models. Sci Rep. 2024 May 14;14(1):10967. https://doi.org/10.1038/s41598-024-61758-0


