Spatial metabolomics, typically performed by MALDI mass spectrometry imaging (MALDI-MSI), enables in situ mapping of metabolite distributions across tissue sections. This capability has become increasingly valuable for investigating tumor microenvironments, drug distribution, and neurological disease pathology. However, the technique is highly sensitive to sample quality because metabolites are small (<1,000 Da), chemically labile, and prone to rapid degradation. As a result, any pre-treatment step between tissue collection and analysis can irreversibly alter the metabolomic profile. In practice, many submitted specimens have already undergone fixation, perfusion, decalcification, or formalin-fixed paraffin embedding (FFPE), rather than the preferred fresh-frozen workflow. This article reviews the reported effects of these four common pre-treatments on spatial metabolomics detection, helping researchers assess sample suitability and make more informed collection decisions.
1. Why Sample Preparation Matters in Spatial Metabolomics
Spatial metabolomics workflow typically involves tissue collection, preservation, sectioning, matrix application, laser desorption and ionization, and pixel-by-pixel mass spectral acquisition. This pipeline is designed to capture both the molecular identities and relative abundances of metabolites, as well as their spatial distributions within intact tissue architecture. The combination of abundance-related information and in situ localization is also what distinguishes MALDI-MSI from bulk metabolomics. Accordingly, preserving molecular composition and preserving spatial fidelity are equally important analytical objectives in spatial metabolomics. This dual requirement makes the method highly sensitive to pre-analytical handling, since sample pre-treatment can alter not only metabolite detectability and apparent abundance, but also the spatial distribution patterns observed across histological regions, lesion margins, and tissue microenvironments (Ngai et al., 2025).
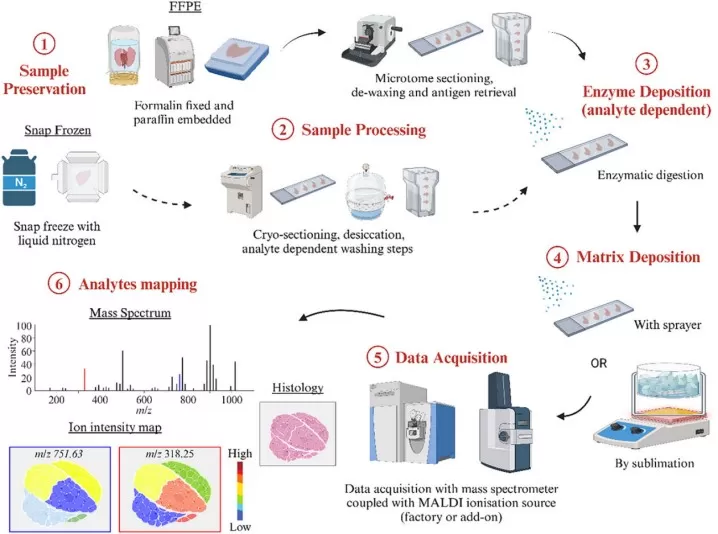
Figure 1. A typical MALDI-MSI workflow. Image reproduced from Ngai et al., 2025, Analytical Science Advances, 6(1), e70016.
At the mechanistic level, most pre-treatment-related artifacts in spatial metabolomics can be grouped into three categories: metabolic degradation over time, chemically induced molecular alteration, and physical loss or spatial redistribution of metabolites.
- The first is continued metabolism after tissue removal. Enzymatic activity does not stop at excision. Labile analytes such as ATP, glycolytic intermediates, and neurotransmitter-related compounds can shift rapidly during the post-mortem or post-excision interval if the tissue is not frozen immediately. Practical MALDI-MSI protocols therefore emphasize minimizing the interval between harvesting and freezing and avoiding unnecessary warming or handling steps (Veerasammy et al., 2020).
- The second is chemical modification. Formaldehyde can react with nucleophilic functional groups, especially primary amines, while acidic processing can promote hydrolysis or broader chemical degradation. For spatial metabolomics, these reactions matter because the detected ion is no longer guaranteed to represent the native molecule present in vivo.
- The third is physical redistribution or extraction. Water-soluble metabolites may diffuse during washes or prolonged immersion, and lipids can be removed during dehydration or clearing. In MSI, that is not merely a yield problem. It can also distort the biological story by shifting spatial contrast across histological regions.
For discovery-oriented studies, rapid fresh-frozen preparation remains the reference standard because it arrests metabolism quickly and preserves native molecular localization more faithfully than any chemical preservation route (Ngai et al., 2025; Veerasammy et al., 2020).
2. Perfusion and Spatial Metabolomics Detection
Perfusion is widely used in animal studies, especially in neuroscience and cardiovascular research, to clear blood and interstitial fluid from the vasculature, thereby reducing blood-derived molecular contamination and improving morphological preservation. A typical protocol involves transcardial perfusion with phosphate-buffered saline (PBS) followed by paraformaldehyde (PFA) fixation. This two-step approach first flushes out blood components, then chemically stabilizes the tissue architecture. In structural biology, histology, and immunostaining workflows, this can be advantageous because blood-derived signals are reduced and tissue architecture is easier to evaluate.
2.1 How Perfusion Alters Metabolite Detection and Spatial Fidelity
For spatial metabolomics, perfusion is not a neutral cleaning step. The metabolic impact of perfusion operates through both temporal and chemical mechanisms. PBS flushing extends the ischemic period between tissue death and metabolic arrest, during which endogenous enzymes remain active. Veerasammy et al. (2020) documented that this extended ischemia significantly exacerbates concentration changes in labile metabolites compared with immediate snap-freezing. These effects are already important with buffer-only perfusion. When fixative is introduced through the vasculature, the sample is further shifted by fixation chemistry. The central issue is that the measured molecular map may reflect post-perfusion chemistry rather than the in vivo metabolite landscape.
2.2 Practical Guidance for Spatial Metabolomics Using Perfused Tissue
Perfused tissue is generally not the preferred starting material for untargeted, discovery-grade spatial metabolomics, because perfusion can disturb both metabolite levels and spatial distribution patterns before metabolism is fully arrested. It is most appropriate only when blood contamination poses a greater threat to data quality than the metabolic disturbance introduced by the perfusion procedure itself. When perfusion cannot be avoided, the most important safeguards are to minimize the interval to freezing or section preparation, apply the mildest effective fixation conditions, and record the protocol in detail. Measures such as shortening PFA exposure or considering faster metabolic arrest strategies, including microwave fixation, may help reduce damage. Even so, perfused tissue is usually more suitable for focused, hypothesis-driven studies or relatively stable analytes than for broad untargeted profiling. Any conclusions about native metabolic state should therefore be made cautiously and interpreted against matched controls processed in the same way, rather than against fresh-frozen tissue from a different workflow.
3. Chemical Fixation and Spatial Metabolomics Detection
Chemical fixation is routine in pathology and many translational workflows because it preserves tissue morphology, stabilizes cellular structure, and supports downstream histology or immunohistochemistry. The most common approach is immersion in neutral buffered formalin or another formaldehyde-based fixative, with 10% neutral buffered formalin (NBF) for 24 to 48 hours being the standard in many clinical protocols. This is often chosen when morphology, storage stability, or compatibility with pathology review is more important than preserving native small-molecule chemistry.
3.1 How Chemical Fixation Alters Metabolite Detection
The chemistry of formalin fixation centers on the reaction of formaldehyde with primary amines (NH2 groups) present in proteins, nucleic acids, and metabolites containing amino or thiol functional groups. This reaction produces methylene bridges (CH2 linkages) that cross-link biomolecules into a stabilized three-dimensional network. For spatial metabolomics, this matters because the detected ion is no longer guaranteed to represent the native metabolite present in vivo. The impact is class-dependent: amino acids, nucleotides, and many amine-containing metabolites are more vulnerable than some neutral lipid classes. Even when analytes remain detectable, fixation can introduce ion suppression, altered peak patterns, and reduced comparability with fresh-frozen tissue. Dannhorn et al. (2022) showed that formalin-fixed samples can still yield spatially resolved metabolite and drug signals, but the chemical space and interpretive confidence differ from those of fresh-frozen material.
3.2 Practical Guidance for Spatial Metabolomics Using Fixed Tissue
Fixed-but-unembedded tissue generally occupies an intermediate position between fresh-frozen and FFPE material. It may still support selected analytes and relative spatial patterns, but it should not be treated as a direct substitute for fresh-frozen collection in unbiased discovery studies. When chemically fixed tissue must be analyzed, short and well-documented fixation conditions are preferable, and study goals should focus on robust analytes, relative trends, or pathology-linked interpretation rather than full metabolic coverage. Whenever possible, adjacent fresh-frozen controls or orthogonal validation should be included.
4. FFPE Processing and Spatial Metabolomics Detection
FFPE tissue is the dominant format in clinical pathology because it supports long-term storage, sectioning quality, and routine histopathological evaluation. The complete FFPE process comprises four sequential steps, each imposing its own metabolic perturbations:
- Fixation: Tissue is immersed in 10% neutral buffered formalin for 24 to 48 hours, initiating the cross-linking reactions described in Chapter 3.
- Graded ethanol dehydration: Tissue is passed through an ascending ethanol series (typically 70%, 80%, 95%, and 100%), which progressively extracts water-soluble metabolites and lipids from the tissue.
- Clearing: Xylene or a xylene substitute displaces ethanol, further removing non-polar molecules from the tissue.
- Paraffin infiltration and embedding: Molten paraffin wax permeates the tissue, displacing remaining solvents and producing a solid block suitable for long-term room-temperature storage.
This route is especially common in retrospective clinical research, pathology archives, and translational studies where large annotated specimen collections already exist.
4.1 How FFPE Processing Changes Metabolite Detection and Lipid Recovery
FFPE is more disruptive than fixation alone because analytical bias accumulates across the entire workflow. Formalin fixation can chemically modify endogenous metabolites and reduce ionization efficiency through cross-linking, while graded ethanol dehydration extracts many polar small molecules and a large proportion of free lipids. Clearing and paraffin infiltration further narrow the retained chemical space, and later deparaffinization creates an additional opportunity for analyte loss. As a result, lipid coverage in FFPE tissue is markedly reduced relative to fresh-frozen samples, whereas polar small molecules are affected by both extraction and formaldehyde-related spectral complexity (Ngai et al., 2025; Denti et al., 2020; Dannhorn et al., 2022). However, not all molecular classes are affected to the same extent. A fraction of membrane-associated, solvent-resistant lipids may still produce detectable signals, although lipid recovery remains incomplete and workflow-dependent; peptides and N-glycans are also more compatible with FFPE-based spatial multiomics workflows than labile small-molecule metabolites (Denti et al., 2020; Denti et al., 2022). Selected metabolites and drug molecules can still be detected when workflows are carefully optimized, although FFPE cannot recapitulate the breadth of the fresh-frozen metabolome (Dannhorn et al., 2022).
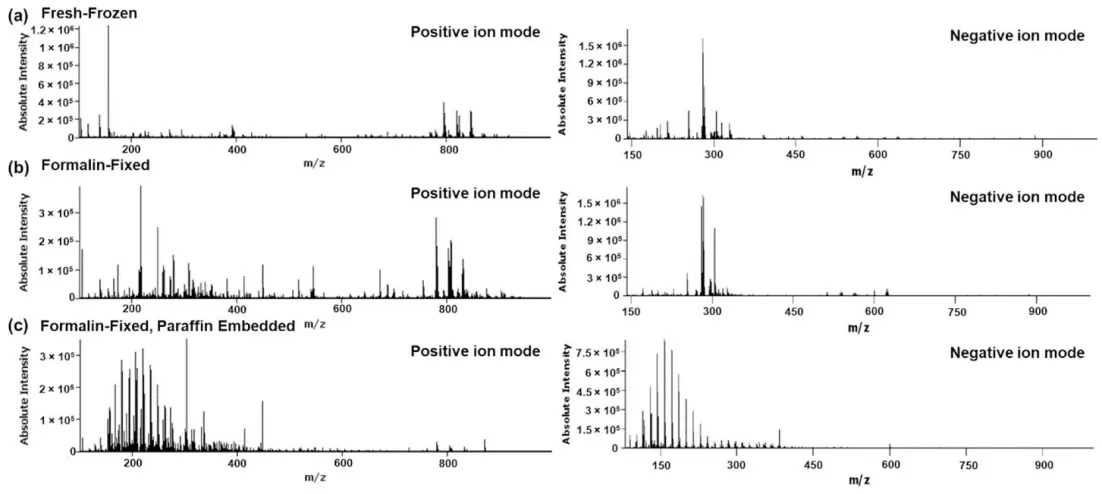
Figure 2. Representative mean spectra of (a) fresh-frozen, (b) formalin-fixed and (c) formalin-fixed, paraffin embedded rat liver specimens analyzed by DESI-MSI in positive and negative ion mode. Image reproduced from Dannhorn et al., 2022, Pharmaceuticals (Basel, Switzerland), 15(11), 1307.
4.2 Practical Guidance for Spatial Metabolomics Using FFPE Samples
FFPE tissue can support spatial metabolomics only within a restricted analytical scope. Before MALDI-MSI, sections require xylene deparaffinization and graded ethanol rehydration, but these steps can themselves remove residual analytes, so conditions must be optimized to balance wax removal against metabolite retention (Ly et al., 2016). Heat-induced retrieval can improve access to retained lipid signals in FFPE sections, especially in lipid-focused workflows, while high-resolution platforms are preferred for distinguishing fixation-related adducts from native signals (Denti et al., 2020; Ly et al., 2016). In practice, FFPE is most suitable for retrospective pathology-based studies, exploratory drug localization, and multimodal designs using adjacent archived sections (Dannhorn et al., 2022; Rittel et al., 2023). It is less suitable for broad untargeted profiling, labile small molecules, or highly accurate quantitative metabolomics, and conclusions should be framed as partial molecular recovery rather than equivalence to fresh-frozen tissue.
5. Decalcification and Spatial Metabolomics Detection
Bone, teeth, and other mineralized tissues often cannot be sectioned for routine MSI workflows without prior removal of calcium-rich material. Decalcification is therefore commonly used in skeletal pathology, bone research, and calcified lesion studies. The main approaches are EDTA-based chelation, formic acid treatment, and stronger acid methods such as hydrochloric acid. These options differ substantially in speed, chemical harshness, and preservation performance.
- EDTA decalcification: A chelation-based approach that sequesters calcium ions in solution. This method is the gentlest but also the slowest, typically requiring days to weeks depending on tissue size and density.
- Formic acid decalcification: An intermediate approach that dissolves calcium salts through acid-mediated dissolution. Processing times range from hours to days.
- Hydrochloric acid decalcification: The fastest method, capable of decalcifying specimens within hours. However, its strongly acidic conditions are the most destructive to biomolecules.
5.1 How Decalcification Chemistry Affects Molecular Preservation
Direct spatial metabolomics benchmarking of decalcification workflows remains limited, so current recommendations rely partly on chemical reasoning and broader molecular preservation studies. Strong acids are fast but introduce conditions that are unfavorable for labile small molecules and prolonged aqueous exposure can promote diffusion or loss. In a comparative pathology study, hydrochloric acid and longer formic acid protocols preserved antigenicity and nucleic-acid readouts less well than EDTA-based decalcification, while EDTA provided the best overall preservation among the tested methods (Miquelestorena-Standley et al., 2020). Li et al. (2024) further showed that accelerated EDTA-based protocols can improve workflow speed while maintaining better preservation of tissue morphology and multiple molecular readouts than harsher chemistries. These studies are not metabolomics benchmarks, but they support that preservation-oriented decalcification chemistry is more compatible with future MSI than aggressive acid treatment.
5.2 Practical Guidance for Spatial Metabolomics Using Decalcified Samples
When decalcification is unavoidable, EDTA-based workflows are currently the most defensible default for spatial metabolomics-oriented studies. Treatment time should still be minimized, because long immersion can itself encourage analyte loss. A pilot study is advisable when sample availability allows, especially for bone projects that depend on labile metabolites. Where technical infrastructure is available, undecalcified hard-tissue sectioning or cryo-based approaches deserve early consideration because they avoid one major source of chemical damage altogether.
6. Sample Selection Framework for Spatial Metabolomics Projects
The best sample type depends on the study question, but one principle remains consistent: fresh-frozen tissue offers the highest metabolite retention and the most defensible spatial interpretation. All other pre-treatments represent trade-offs between morphology, logistics, archival value, and chemical fidelity.
The table below provides a qualitative planning summary. The star ratings are relative and intended for project design rather than as quantitative benchmarks.
| Sample Type | Tissue Morphology | Metabolite Retention | Spatial Fidelity | Best Fit | Main Caution |
|---|---|---|---|---|---|
| Fresh-frozen | ★★★★☆ | ★★★★★ | ★★★★★ | Discovery spatial metabolomics and lipidomics | Requires immediate freezing and disciplined handling |
| Perfused | ★★★★★ | ★★☆☆☆ | ★★☆☆☆ | Morphology-led animal studies with limited targeted MSI | Wash effects and delayed metabolic arrest can distort native maps |
| Formalin-fixed | ★★★★★ | ★★☆☆☆ | ★★★☆☆ | Selected robust analytes, fixed-but-unembedded tissue, pathology-linked interpretation | Cross-linking, adduct formation, and ion suppression reduce comparability |
| FFPE | ★★★★★ | ★☆☆☆☆ | ★★☆☆☆ | Archived retrospective studies, multimodal pathology projects | Multi-step processing narrows chemical space and increases extraction bias |
| Decalcified (EDTA-preferred) | ★★★★☆ | ★★★☆☆ | ★★★☆☆ | Calcified tissues when decalcification cannot be avoided | Direct MSI evidence remains limited; protocol choice strongly matters |
For prospective discovery studies, fresh-frozen collection remains the preferred route. If pathology review is also required, a parallel design is often the strongest option: one portion is snap-frozen for MSI, while another is processed for FFPE histology or related pathology workflows. Multimodal strategies built around fresh-frozen tissue plus routine pathology can substantially improve study flexibility (Rittel et al., 2023).
For perfused or chemically fixed tissues, the key question is whether the project can tolerate a chemical state that has already diverged from the native one. These samples may still support selected analytes or carefully framed biological comparisons, but they are usually a poor basis for broad discovery claims.
For FFPE archives, the main decision is whether partial chemical recovery is sufficient for the scientific question. If the answer depends on archived clinical material, selected compound classes, or spatial trends linked to pathology, FFPE can be valuable. If the answer depends on full coverage of labile metabolites, FFPE is usually the wrong substrate.
For bone and calcified tissues, the decision point is whether decalcification is unavoidable. When it is, EDTA-based workflows are the safest starting point; when it is not, hard-tissue sectioning deserves serious evaluation during study design.
7. How MetwareBio Supports Spatial Metabolomics Study Design
Because sample history often determines whether spatial metabolomics data will be biologically interpretable, early technical review is critical before sample submission. MetwareBio provides spatial metabolomics services across both MALDI-MSI and AFADESI-MSI workflows, allowing project design to be matched more closely to the biological question, tissue type, analyte class, and desired balance between spatial resolution and metabolite coverage. In addition to discovery-oriented spatial metabolomics and spatial lipidomics, MetwareBio has developed specialized MALDI-based workflows for phytohormone spatial profiling and neurotransmitter spatial profiling, supported by an extensive in-house MALDI metabolite database and optimized derivatization and identification strategies for low-abundance analytes. The team has experience across a wide range of specimen types, including plant tissues, animal tissues, and archived pathological materials, and MetwareBio-supported studies have been reported in high-impact journals, including PNAS and Nature Cancer, among many other peer-reviewed outlets. Researchers planning spatial metabolomics studies, especially those involving challenging or non-standard samples, are welcome to contact MetwareBio for technical evaluation and project design support.
Contact UsRead More: Spatial Metabolomics and Sample Preparation
From tissue handling to workflow design, these articles extend the current discussion on spatial metabolomics sample preparation, covering MALDI-MSI workflows, embedding media selection, and spatial omics integration.
Comprehensive guidelines for MALDI-based spatial metabolomics covering tissue sampling, embedding media, sectioning, preservation, and transport, providing practical protocols that complement the pre-treatment considerations discussed above.
An overview of the complete MALDI-MSI workflow from matrix application to data interpretation, helping you understand how upstream sample decisions flow into downstream analytical outcomes.
A classification and comparison of spatial omics platforms, including MALDI-MSI, DESI, and AFADESI, contextualizing spatial metabolomics within the broader spatial omics technology landscape.
Explores how spatial metabolomics data can be integrated with spatial transcriptomics and proteomics for multi-modal discovery, especially relevant when working with FFPE archives that support multimodal workflows.
A case study demonstrating spatial multi-omics in liver disease research, illustrating how optimized tissue handling and spatial metabolomics work together in translational pathology settings.
Looks at the convergence of spatial metabolomics with single-cell technologies, highlighting how sample quality decisions at the tissue level affect multi-scale molecular discovery.
References
- Dannhorn, A., Swales, J. G., Hamm, G., Strittmatter, N., Kudo, H., Maglennon, G., Goodwin, R. J. A., & Takats, Z. (2022). Evaluation of Formalin-Fixed and FFPE Tissues for Spatially Resolved Metabolomics and Drug Distribution Studies. Pharmaceuticals (Basel, Switzerland), 15(11), 1307. https://doi.org/10.3390/ph15111307
- Denti, V., Capitoli, G., Piga, I., Clerici, F., Pagani, L., Criscuolo, L., Bindi, G., Principi, L., Chinello, C., Paglia, G., Magni, F., & Smith, A. (2022). Spatial Multiomics of Lipids, N-Glycans, and Tryptic Peptides on a Single FFPE Tissue Section. Journal of Proteome Research, 21(11), 2798–2809. https://doi.org/10.1021/acs.jproteome.2c00601
- Denti, V., Piga, I., Guarnerio, S., Clerici, F., Ivanova, M., Chinello, C., Paglia, G., Magni, F., & Smith, A. (2020). Antigen Retrieval and Its Effect on the MALDI-MSI of Lipids in Formalin-Fixed Paraffin-Embedded Tissue. Journal of the American Society for Mass Spectrometry, 31(8), 1619–1624. https://doi.org/10.1021/jasms.0c00208
- Li, Z., Wenhart, C., Reimann, A., Cho, Y.-L., Adler, K., & Muench, G. (2024). Hypertonic saline- and detergent-accelerated EDTA-based decalcification better preserves mRNA of bones. Scientific Reports, 14, 10888. https://doi.org/10.1038/s41598-024-61459-8
- Ly, A., Buck, A., Balluff, B., Sun, N., Gorzolka, K., Feuchtinger, A., Janssen, K. P., Kuppen, P. J., van de Velde, C. J., Weirich, G., Erlmeier, F., Langer, R., Aubele, M., Zitzelsberger, H., McDonnell, L., Aichler, M., & Walch, A. (2016). High-mass-resolution MALDI mass spectrometry imaging of metabolites from formalin-fixed paraffin-embedded tissue. Nature Protocols, 11(8), 1428–1443. https://doi.org/10.1038/nprot.2016.081
- Miquelestorena-Standley, E., Jourdan, M. L., Collin, C., Bouvier, C., Larousserie, F., Aubert, S., Gomez-Brouchet, A., Guinebretière, J. M., Tallegas, M., Brulin, B., Le Nail, L. R., Tallet, A., Le Loarer, F., Massiere, J., Galant, C., & de Pinieux, G. (2020). Effect of decalcification protocols on immunohistochemistry and molecular analyses of bone samples. Modern Pathology, 33(8), 1505–1517. https://doi.org/10.1038/s41379-020-0503-6
- Ngai, Y. T., Lau, D., Mittal, P., & Hoffmann, P. (2025). Mini Review: Highlight of Recent Advances and Applications of MALDI Mass Spectrometry Imaging in 2024. Analytical Science Advances, 6(1), e70016. https://doi.org/10.1002/ansa.70016
- Rittel, M. F., Schmidt, S., Weis, C. A., Birgin, E., van Marwick, B., Rädle, M., Diehl, S. J., Rahbari, N. N., Marx, A., & Hopf, C. (2023). Spatial Omics Imaging of Fresh-Frozen Tissue and Routine FFPE Histopathology of a Single Cancer Needle Core Biopsy: A Freezing Device and Multimodal Workflow. Cancers, 15(10), 2676. https://doi.org/10.3390/cancers15102676
- Veerasammy, K., Chen, Y. X., Sauma, S., Pruvost, M., Dansu, D. K., Choetso, T., Zhong, T., Marechal, D., Casaccia, P., Abzalimov, R., & He, Y. (2020). Sample Preparation for Metabolic Profiling using MALDI Mass Spectrometry Imaging. Journal of Visualized Experiments: JoVE, (166), 10.3791/62008. https://doi.org/10.3791/62008


