Quantitative proteomics experiments often identify hundreds to thousands of differentially expressed proteins (DEPs), yet a list of upregulated and downregulated proteins alone provides limited functional insight. Subcellular localization analysis helps address this gap by integrating curated databases with machine learning-based prediction tools to map DEPs to the cellular compartments where they are most likely to function, thereby converting a flat expression profile into a spatially informed view of biological change. This article provides a practical guide to subcellular localization analysis, covering the major bioinformatics resources and tools, a step-by-step workflow for analyzing the subcellular distribution of DEPs, and strategies for interpreting the results to generate meaningful biological hypotheses.
1. What Is Subcellular Localization in Proteomics?
Subcellular localization refers to the specific cellular compartment or structural region in which a protein resides and performs its function. In eukaryotic cells, proteins are not distributed randomly; instead, they are organized within a highly structured intracellular system composed of membrane-bound organelles and functionally specialized microenvironments. This spatial organization is a fundamental principle of cell biology, because protein activity depends not only on molecular identity and abundance, but also on where the protein is positioned within the cell. For example, transcriptional regulators must access the nucleus to control gene expression, whereas enzymes involved in oxidative phosphorylation must localize to mitochondria to participate in energy metabolism (Christopher et al., 2021).
From an analytical perspective, subcellular localization can be described at multiple levels, ranging from broad categories such as nuclear, cytoplasmic, membrane, and extracellular proteins to more refined organelle-level annotations such as endoplasmic reticulum, Golgi apparatus, lysosome, or peroxisome. Standardized classification systems, particularly Gene Ontology Cellular Component (GO-CC), provide a hierarchical vocabulary for organizing these localization categories and linking them to biological interpretation. In proteomics studies, this framework is especially valuable because proteins from the same compartment often participate in related pathways, structural programs, or trafficking networks. As a result, examining the localization distribution of differentially expressed proteins (DEPs) can reveal whether a biological perturbation is primarily associated with nuclear regulation, mitochondrial metabolism, secretory remodeling, membrane signaling, or intracellular degradation pathways, thereby helping convert protein lists into coherent mechanistic insight.
In practical proteomics research, subcellular localization analysis has become an important downstream step that bridges differential expression results with functional interpretation. Rather than treating DEPs as an undifferentiated list, researchers can use localization information to prioritize candidates, identify affected cellular systems, and generate hypotheses about the underlying biology driving observed expression changes. This spatially informed perspective is particularly useful in complex disease studies, where expression changes may span multiple cellular compartments and biological processes.
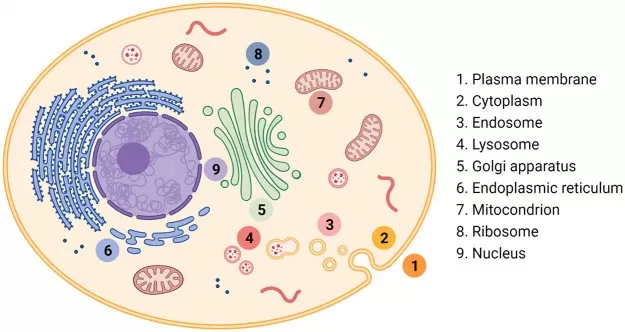
Figure 1. Key cellular compartments. Image reproduced from Zhou et al., 2022, Accounts of chemical research, 55(20), 2998–3009.
Table 1. Major Subcellular Localization Regions and Their Biological Relevance
| Compartment | Example Functions | Relevance to Proteomics |
|---|---|---|
| Nucleus | DNA replication, transcription, RNA processing, chromatin remodeling | Enrichment of nuclear DEPs suggests transcriptional regulation or genome maintenance; commonly linked to cancer and developmental studies |
| Cytoplasm | Signal transduction, metabolic reactions, cytoskeletal organization | Broad functional category; enrichment often reflects generalized cellular response or metabolic reprogramming |
| Mitochondria | Oxidative phosphorylation, apoptosis, fatty acid β-oxidation, TCA cycle | Mitochondrial DEP enrichment is a strong indicator of energy metabolism disruption, oxidative stress, or apoptotic signaling |
| Endoplasmic Reticulum (ER) | Protein folding, quality control, lipid biosynthesis, calcium storage | ER-localized DEPs may indicate ER stress, unfolded protein response activation, or secretory pathway changes |
| Golgi Apparatus | Protein sorting, glycosylation, vesicular trafficking | Enrichment of Golgi proteins can reflect alterations in protein trafficking, secretion, or post-translational modification |
| Lysosome | Autophagy, macromolecule degradation, antigen presentation | Lysosomal DEP enrichment is commonly observed in neurodegenerative diseases, lysosomal storage disorders, and immune activation |
| Peroxisome | Fatty acid oxidation, reactive oxygen species metabolism, plasmalogen synthesis | Peroxisomal protein changes may implicate redox balance or lipid metabolism pathways |
| Plasma Membrane | Cell adhesion, receptor signaling, transport, cell-cell communication | Membrane DEPs are strong biomarker candidates due to accessibility; enrichment suggests altered signaling or cellular interaction |
| Extracellular Space / Secreted | Immune signaling, cell-matrix interactions, paracrine communication | Secreted proteins are highly valuable as non-invasive biomarker candidates; enrichment may reflect intercellular signaling or tissue remodeling |
2. Key Databases and Prediction Tools for Subcellular Localization Analysis
Computational subcellular localization analysis relies on two complementary approaches: curated database annotations and in silico prediction tools. Curated databases provide a foundation of literature-supported and ontology-linked localization information, while prediction tools help extend coverage to proteins with incomplete or missing annotations. Together, these resources make it possible to interpret the spatial distribution of differentially expressed proteins (DEPs) in a systematic and scalable manner.
2.1 Curated Annotation Resources: UniProtKB and HPA
Among annotation resources, UniProtKB remains the most widely used foundation for subcellular localization analysis because it integrates curated subcellular location annotations, sequence features, and cross-referenced functional information, including Gene Ontology Cellular Component (GO-CC) terms (UniProt Consortium, 2025). For well-studied model organisms, these annotations cover a substantial proportion of the proteome. Researchers can retrieve subcellular location data programmatically via the UniProt REST API or dedicated R tools such as UniprotR.
For human proteins, the Human Protein Atlas (HPA) provides an important complementary resource by offering experimentally informed subcellular localization evidence derived largely from immunofluorescence-based mapping, making it especially useful when evaluating compartment-specific patterns in human or mammalian datasets (Thul et al., 2017). However, annotation coverage is not uniform across species or proteins, and many proteins identified in discovery proteomics—especially those from non-model organisms or poorly characterized gene families—still lack complete localization records. In these cases, computational prediction tools become essential for expanding interpretive coverage.
2.2 Prediction Tools: WoLF PSORT, DeepLoc, CELLO, and SignalP
Several prediction tools are commonly used to infer protein localization from amino acid sequence features, learned patterns, or targeting signals. These tools differ in algorithmic design, organism scope, output granularity, and suitability for multi-localized proteins, so they are best treated as complementary sources of evidence rather than interchangeable solutions.
Table 2. Commonly Used Prediction Tools for Subcellular Localization Analysis
| Tool | Method | Typical Application | Organism Coverage | Key Strength |
|---|---|---|---|---|
| WoLF PSORT | Sorting signal analysis combined with k-nearest neighbor classification | General prediction of likely subcellular compartment based on sequence-derived localization features | Primarily eukaryotes, including animal, plant, and fungal models | Widely used classical predictor; useful for broad compartment assignment and cross-checking results from newer tools (Horton et al., 2007) |
| DeepLoc 2.0 | Deep learning with protein language model-based representation | Prediction of eukaryotic protein localization, including proteins with more than one likely compartment | Eukaryotes | Supports multi-label prediction and offers strong overall performance for modern localization inference (Thumuluri et al., 2022) |
| CELLO | Two-level support vector machine (SVM) framework | Compartment prediction using organism-specific classification models | Eukaryotes and prokaryotes | Useful species-aware predictor with a long track record in localization studies (Yu et al., 2006) |
| SignalP 6.0 | Deep learning and protein language model-based signal peptide prediction | Identification of proteins likely to enter the secretory pathway through N-terminal signal peptides | Broad applicability across domains of life | Specialized for signal peptide type and cleavage site prediction; useful for recognizing secreted proteins or proteins entering the secretory pathway rather than general compartment assignment |
WoLF PSORT is one of the most established tools, converting protein sequences into numerical localization features based on sorting signals, amino acid composition, and functional motifs, and then applying a k-nearest neighbor algorithm for classification (Horton et al., 2007). DeepLoc 2.0 represents a more recent advance, leveraging protein language models to improve localization prediction and support multi-label output for proteins that may reside in more than one compartment (Thumuluri et al., 2022). CELLO employs a two-level support vector machine (SVM) framework and provides organism-specific models for different eukaryotic and prokaryotic datasets (Yu et al., 2006). SignalP, although not a general subcellular localization predictor, is particularly useful for identifying signal peptides and proteins likely to enter the secretory pathway (Teufel et al., 2022). A recent review by Gillani and Pollastri (2024) surveyed a broad range of subcellular localization prediction tools across eukaryotic, prokaryotic, and viral categories, underscoring the rapid expansion of this field.
3. Bioinformatics Workflow for Subcellular Localization Analysis
The following workflow outlines a standard computational pipeline for mapping DEPs to their subcellular compartments. This pipeline integrates curated annotations with computational predictions to achieve comprehensive and reliable coverage.
Step 1. Standardize Protein IDs and Retrieve Sequences
The first step involves converting protein identifiers from the proteomics experiment (e.g., gene symbols, RefSeq IDs, or Ensembl IDs) into UniProt accessions, which serve as the standard identifier for downstream retrieval. Tools such as the UniProt ID mapping service or the R function clusterProfiler::bitr can perform this conversion efficiently. Once UniProt accessions are obtained, the corresponding protein sequences in FASTA format are retrieved for proteins that will require computational prediction.
Step 2. Retrieve Curated Localization Annotations
With the mapped UniProt accessions, subcellular location annotations are retrieved in bulk via the UniProt REST API or batch query interface. The returned data includes both the curated "SUBCELLULAR LOCATION" text and the associated GO Cellular Component terms. The GO CC terms are particularly useful for downstream enrichment analysis because they follow a standardized ontology with defined hierarchical relationships. At this stage, a proportion of DEPs will have well-documented localization, while others may carry partial or no annotation — particularly for less-studied proteins or non-model organisms.
Step 3. Predict Localization for Unannotated Proteins
For proteins lacking experimental annotations, computational prediction tools are deployed. DeepLoc 2.0 can be used as the primary predictor for eukaryotic proteins owing to its multi-label capability and strong performance benchmarks (Thumuluri et al., 2022). WoLF PSORT provides a useful cross-validation: agreement between multiple tools increases confidence in the prediction, while disagreement flags cases that may require manual inspection. Results from UniProt annotations and prediction tools are merged into a unified localization table, organized by confidence tier: experimentally validated (Tier 1), multi-tool agreement (Tier 2), and single-tool prediction (Tier 3).
Step 4. Perform GO Cellular Component Enrichment Analysis
Beyond assigning individual localizations, enrichment analysis identifies compartments that are statistically over-represented among the DEPs relative to a background set. This is typically performed using R/Bioconductor packages such as clusterProfiler or topGO, which implement over-representation analysis (ORA) or gene set enrichment analysis (GSEA). The background set should ideally comprise all proteins detected in the proteomics experiment rather than the entire theoretical proteome, as this accounts for the detection bias inherent in mass spectrometry. Multiple testing correction (e.g., Benjamini-Hochberg FDR) is applied to control false positives, and results are visualized as dot plots, bar charts, or enrichment network diagrams.
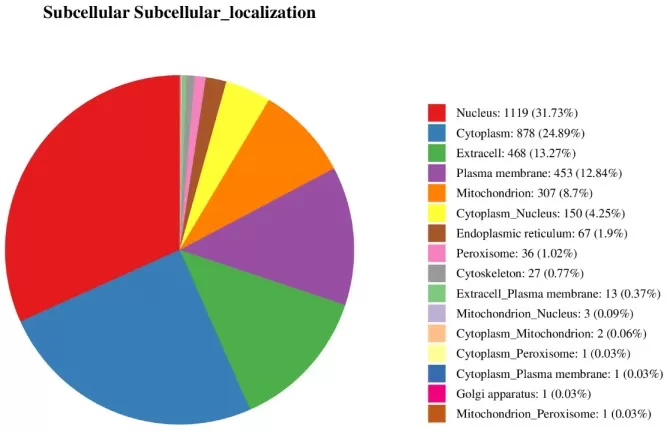
Figure 2. Subcellular Distribution of Differentially Expressed Proteins.
4. How to Interpret Subcellular Localization Results in Proteomics
Subcellular localization analysis becomes most informative when localization patterns are translated into biological meaning rather than treated as simple compartment labels. Interpreting these results requires attention not only to where proteins are enriched, but also to how spatial patterns align with functional pathways, regulatory programs, and the dynamic behavior of multi-localized proteins.
4.1 Analyze Compartment Distribution Patterns
The first level of interpretation involves examining the overall distribution of DEPs across subcellular compartments. A bar chart or pie chart showing the proportion of upregulated and downregulated proteins in each compartment provides an immediate visual summary. Over-represented compartments suggest that the biological perturbation under study preferentially affects proteins in those locations. For example, an enrichment of ER-localized DEPs in a proteotoxic stress experiment may indicate activation of the unfolded protein response, while an accumulation of nuclear proteins could reflect widespread transcriptional reprogramming. The direction of change also matters: upregulated membrane proteins may signal increased cellular communication, whereas their downregulation may suggest suppressed receptor signaling.
4.2 Integrate Localization with Functional Enrichment
Subcellular localization results gain additional significance when cross-referenced with other functional enrichment analyses. If GO Biological Process (BP) enrichment reveals terms related to "oxidative phosphorylation" and the CC enrichment concurrently highlights mitochondrial localization, the combined evidence provides a stronger basis for biological interpretation than either analysis alone. Similarly, KEGG pathway enrichment results should be examined for consistency with compartment-level patterns: pathways primarily mediated by plasma membrane receptors should align with plasma membrane enrichment at the CC level. This triangulation across multiple annotation dimensions — molecular function, biological process, cellular component, and pathway — yields the most robust and convincing biological narrative.
4.3 Address Multi-Localization and Ambiguity
A significant challenge in interpreting subcellular localization data is the prevalence of multi-localized proteins. Many proteins genuinely reside in more than one compartment or shuttle between locations in a regulated manner. When a protein is annotated to multiple compartments, it should be counted toward each relevant category in enrichment analysis rather than arbitrarily assigned to a single location. DeepLoc 2.0 handles this explicitly through its multi-label output, and UniProt annotations frequently list multiple locations with supporting evidence. A practical strategy is to assign confidence tiers: Tier 1 experimental annotations are preferred over predictions, and discrepancies between tools are flagged for manual review. In enrichment analysis, multi-localized proteins contribute to each compartment they occupy, which more accurately reflects their biological role.
5. Research Applications of Subcellular Localization Analysis
Once localization patterns have been interpreted in context, their practical value becomes clearer across a range of research settings. From biomarker screening to target prioritization and mechanism discovery, compartment-level information can help connect proteomics results to biologically and clinically relevant questions.
5.1 Cancer Biomarker Discovery
Subcellular localization analysis can substantially improve mass spectrometry-based cancer biomarker discovery by helping researchers prioritize proteins that are more likely to be detectable, clinically accessible, or biologically relevant to tumor progression. In particular, proteins localized to the plasma membrane, extracellular space, or secretory pathway are often of greatest interest because they are more likely to be released into biofluids, exposed on the cell surface, or involved in tumor–microenvironment communication. When integrated with differential proteomics, localization information adds an important layer of biological context that helps distinguish mechanistically meaningful candidates from large background protein lists. This strategy has been applied across multiple proteomics-based biomarker studies in cancer research (Kume et al., 2014; Birse et al., 2015; Wang et al., 2023).
5.2 Drug Target Prioritization
In drug discovery, subcellular localization directly informs target druggability. Proteins located on the cell surface or secreted into the extracellular space are generally more accessible to antibody-based therapeutics (such as monoclonal antibodies and antibody-drug conjugates) than intracellular proteins. Localization analysis of DEPs in disease contexts can reveal which dysregulated proteins occupy drug-accessible compartments, thereby streamlining the target selection process. Receptor tyrosine kinases and G-protein-coupled receptors enriched among DEPs, for instance, would be immediately prioritized given their well-established pharmacological tractability.
5.3 Disease Mechanism Discovery
Beyond biomarker and drug target identification, subcellular localization patterns can illuminate fundamental disease mechanisms. A redistribution of mitochondrial proteins in neurodegenerative disease models may indicate impaired mitochondrial dynamics or bioenergetic dysfunction. An accumulation of nuclear proteins with DNA repair functions in cancer cells might reflect genomic instability. Similarly, enrichment of lysosomal proteins in lysosomal storage disorders directly implicates the degradation pathway in disease pathology. By connecting localization shifts to compartment-specific biology, researchers can generate focused, testable mechanistic hypotheses.
6. Practical Tips and Common Pitfalls for Subcellular Localization Analysis
Several practical considerations can significantly affect the quality and interpretability of subcellular localization analysis:
- Database version consistency: Ensure that all retrieval steps (ID mapping, annotation retrieval, background set definition) use the same version of UniProt. Annotations are updated periodically, and inconsistent versions can introduce discrepancies that propagate through the analysis pipeline.
- Species-specific tool selection: Not all prediction tools perform equally well across organisms. WoLF PSORT and CELLO offer species-specific models (animal, plant, fungal), while DeepLoc 2.0 is optimized for eukaryotes but trained predominantly on human and model organism data. For non-model organisms, cross-tool consensus becomes especially important for reliable predictions.
- Sequence-level vs. gene-level analysis: Many proteomics experiments report results at the gene level, collapsing protein isoforms into a single entry. However, isoforms of the same gene can differ in subcellular localization due to alternative splicing affecting targeting signals. When isoform-resolved data is available, predictions should be performed at the sequence level to capture these biologically meaningful differences.
- Background set selection: For GO CC enrichment analysis, the choice of background set significantly influences statistical results. Using all detected proteins from the proteomics experiment as the background generally produces more reliable results than using the whole proteome, as it accounts for detection limitations inherent in the experimental setup.
- Computational predictions as hypotheses: Regardless of the tool or confidence tier, computational predictions are not substitutes for experimental validation. Localization results should be treated as hypotheses to guide further investigation, with key findings confirmed through immunofluorescence microscopy, subcellular fractionation coupled with western blotting, or proximity labeling techniques (Christopher et al., 2021).
How MetwareBio Supports Subcellular Localization Analysis in Proteomics
Subcellular localization analysis requires not only the right tools but also expertise in data processing, statistical analysis, and biological interpretation. MetwareBio provides end-to-end proteomics services that encompass quantitative protein identification, differential expression analysis, and comprehensive downstream bioinformatics analysis, including subcellular localization mapping and multi-dimensional functional enrichment.
With a dedicated team of bioinformaticians and access to advanced computational infrastructure, MetwareBio ensures that every proteomics project delivers both statistical rigor and biological relevance. From experimental design consultation through to publication-ready figures and interpretation, the integrated service workflow is designed to maximize the scientific value of proteomics data. For researchers seeking a streamlined, expert-guided approach to proteomics data analysis, the MetwareBio platform and support team offer a reliable solution. Visit MetwareBio to explore the full range of available services, or access the MetwareBio Cloud Platform for interactive data analysis tools.
Contact UsRead More: Proteomics Data Analysis and Functional Interpretation
Subcellular localization analysis is one piece of a broader proteomics data interpretation pipeline. The articles below cover upstream differential analysis, downstream functional enrichment, and related multi-omics strategies to help you build a complete analytical workflow.
A solid localization analysis starts with reliable differential expression testing. This article reviews the practical choices between t-tests, ANOVA, and non-parametric alternatives for proteomics datasets, including guidance on multiple testing correction and effect size estimation.
Before mapping DEPs to cellular compartments, it is essential to understand how fold change, p-value, FDR, and VIP scores each contribute to feature selection. This guide explains why a multi-criteria approach produces more robust candidate lists.
GO Cellular Component enrichment is a direct extension of subcellular localization analysis. This comparison guide explains when to use GO, KEGG, and COG/KOG annotations, and how to combine them for a comprehensive functional interpretation of your DEPs.
After identifying enriched compartments, KEGG pathway analysis provides the next layer of functional context. Learn how to read enrichment output — including Gene Count, Rich Factor, p-values, and FDR — to build biologically meaningful pathway narratives.
Existing proteomics datasets can be reanalyzed with updated annotations and tools, including subcellular localization predictors. This guide covers practical strategies for extracting new insights from previously published mass spectrometry data.
Weighted gene co-expression network analysis can complement subcellular localization by identifying modules of co-expressed proteins that may share compartment-level organization and functional roles in disease biology.
References
- Birse, C. E., Lagier, R. J., FitzHugh, W., … Ruben, S. M. (2015). Blood-based lung cancer biomarkers identified through proteomic discovery in cancer tissues, cell lines and conditioned medium. Clinical proteomics, 12(1), 18. https://doi.org/10.1186/s12014-015-9090-9
- Christopher, J. A., Stadler, C., Martin, C. E., Morgenstern, M., ... Lundberg, E. (2021). Subcellular proteomics. Nature Reviews Methods Primers, 1, 32. https://doi.org/10.1038/s43586-021-00029-y
- Gillani, M., & Pollastri, G. (2024). Protein subcellular localization prediction tools. Computational and Structural Biotechnology Journal, 23, 1796-1807. https://doi.org/10.1016/j.csbj.2024.04.032
- Horton, P., Park, K. J., Obayashi, T., ... Nakai, K. (2007). WoLF PSORT: Protein localization predictor. Nucleic Acids Research, 35(suppl_2), W585-W587. https://doi.org/10.1093/nar/gkm259
- Kume, H., Muraoka, S., Kuga, T., …Tomonaga, T. (2014). Discovery of colorectal cancer biomarker candidates by membrane proteomic analysis and subsequent verification using selected reaction monitoring (SRM) and tissue microarray (TMA) analysis. Molecular & cellular proteomics : MCP, 13(6), 1471–1484. https://doi.org/10.1074/mcp.M113.037093
- Teufel, F., Almagro Armenteros, J. J., Johansen, A. R., Gíslason, M. H., Pihl, S. I., Tsirigos, K. D., Winther, O., Brunak, S., von Heijne, G., & Nielsen, H. (2022). SignalP 6.0 predicts all five types of signal peptides using protein language models. Nature biotechnology, 40(7), 1023–1025. https://doi.org/10.1038/s41587-021-01156-3
- Thul, P. J., Åkesson, L., Wiking, M., ... Lundberg, E. (2017). A subcellular map of the human proteome. Science, 356(6340), eaal3321. https://doi.org/10.1126/science.aal3321
- Thumuluri, V., Almagro Armenteros, J. J., Johansen, A. R., Nielsen, H., & Winther, O. (2022). DeepLoc 2.0: Multi-label subcellular localization prediction using protein language models. Nucleic Acids Research, 50(W1), W228-W234. https://doi.org/10.1093/nar/gkac278
- UniProt Consortium (2025). UniProt: the Universal Protein Knowledgebase in 2025. Nucleic acids research, 53(D1), D609–D617. https://doi.org/10.1093/nar/gkae1010
- Wang, W., Huang, G., Lin, H., Ren, L., Fu, L., & Mao, X. (2023). Label-free LC-MS/MS proteomics analyses reveal CLIC1 as a predictive biomarker for bladder cancer staging and prognosis. Frontiers in oncology, 12, 1102392. https://doi.org/10.3389/fonc.2022.1102392
- Yu, C. S., Chen, Y. C., Lu, C. H., & Hwang, J. K. (2006). Prediction of protein subcellular localization. Proteins: Structure, Function, and Bioinformatics, 64(3), 643-651. https://doi.org/10.1002/prot.21018
- Zhou, Z., Maxeiner, K., Ng, D. Y. W., & Weil, T. (2022). Polymer Chemistry in Living Cells. Accounts of chemical research, 55(20), 2998–3009. https://doi.org/10.1021/acs.accounts.2c00420


