In spatial metabolomics research, selecting regions of interest (ROI) is a deceptively simple yet critically important step that can determine the success of a study. A single tissue section contains tens of thousands of spatial pixels, each harboring rich metabolic information. However, due to limitations in instrument analysis time, sample availability, and experimental costs, conducting full-coverage, high-depth analysis across the entire section is often impractical. How can researchers accurately pinpoint the most biologically meaningful regions from such vast spatial data? This decision not only affects experimental efficiency but also directly influences the depth and reliability of subsequent biological discoveries. This article outlines strategies and methodologies for ROI selection, aiming to assist researchers in optimizing spatial analysis and maximizing the interpretability of molecular heterogeneity.
1. Why ROI Selection Is Critical in Spatial Metabolomics
The central paradigm in spatial metabolomics experiments is to "observe the global context before examining local details," ensuring that biologically relevant regions are accurately captured for downstream analysis. In a typical mass spectrometry imaging (MSI) workflow, the process generally involves:
- Conducting a low-resolution pre-scan of the entire tissue section to obtain an overview of metabolite distributions;
- Defining ROIs based on pre-scan results or tissue morphological images, guided by anatomical and functional features;
- Performing high-resolution acquisition or subsequent multi-omics analyses within the selected ROIs.
The quality of ROI selection directly influences the ability to detect spatial heterogeneity, identify region-specific metabolites, and uncover pathway variations. Inadequate selection can lead to missing key metabolic transition zones, mixing distinct cellular populations, or inefficient use of costly samples and instrument time. Careful ROI planning maximizes data interpretability, experimental efficiency, and reproducibility across studies, enabling more reliable discovery of spatially resolved biomarkers.
2. Limitations of Traditional ROI Selection Methods
Historically, ROI selection has primarily relied on two approaches, each with inherent limitations:
Morphology-Guided Manual Annotation
Researchers manually delineate regions on tissue sections using H&E or other histological staining images, guided by pathological expertise. While this method is intuitive and aligns with clinical pathology, it is highly subjective and shows poor reproducibility. Different researchers may generate completely different ROIs on the same tissue section, making cross-study comparisons challenging and limiting standardization.
MSI Feature Similarity-Based Automated Segmentation
Clustering algorithms group pixels with similar spectral features to generate segmented maps. Although automated segmentation enhances throughput and reduces manual labor, its outcomes are often sensitive to algorithm parameters and susceptible to instrument noise or imaging artifacts. This can lead to inconsistencies and questionable biological reliability. Both traditional approaches may fail to capture subtle metabolic heterogeneity or rare cell populations, highlighting the need for more advanced, reproducible ROI selection strategies in spatial metabolomics.
3. Advanced ROI Selection Techniques in Spatial Metabolomics
Between 2024 and 2025, spatial metabolomics witnessed significant methodological advancements in ROI selection, driven by the need for more accurate, reproducible, and high-throughput analysis of tissue sections. Several innovative strategies have emerged that leverage metabolite-guided multi-omics, AI-based automated selection, and multi-modal data integration to improve precision and biological relevance. These approaches represent some of the most impactful innovations in ROI selection, combining fundamental principles with practical advantages and applications that enhance precision, reproducibility, and biological insight in modern spatial metabolomics workflows.
3.1 Metabolomics-Guided Multi-Omics Strategy
A key approach in spatial metabolomics involves using metabolite information to guide the integration of multiple omics layers on the same tissue section. Non-destructive pre-analysis of the tissue using desorption electrospray ionization MSI (DESI-MSI) allows segmentation maps to be generated from metabolite distribution patterns, which can then guide ROI selection for in-depth spatial proteomics. This workflow enables seamless integration of spatial metabolomics and proteomics within a single section. DESI-MSI requires minimal matrix and introduces negligible sample interference, making it an effective strategy for accurate multi-omics spatial targeting.
In a 2025 study published in Analytical Chemistry, rat brain tissue slices were analyzed with DESI-MSI, yielding 160 lipid annotations (SwissLipids database, FDR ≤ 20%). Segmentation maps derived from these metabolite images facilitated precise selection of 200 μm × 200 μm ROIs for subsequent proteomic analysis, resulting in identification of 3,888–4,717 proteins per ROI.
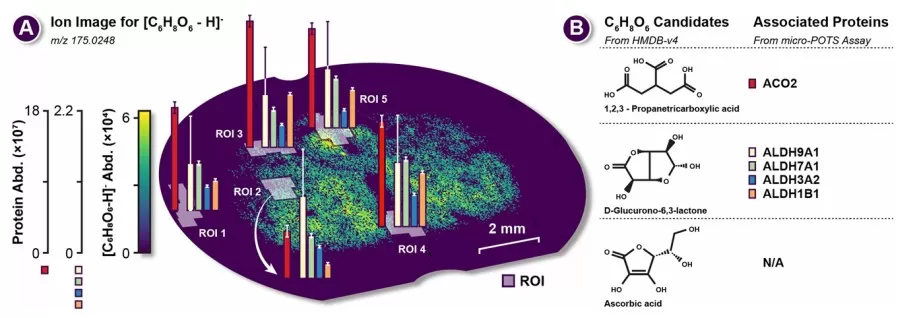
Figure 1. Example of spatial multi-omics ROI for C6H8O6 and associated proteins. Image adapted from Vandergrift et al. (2025), Analytical Chemistry, 97(1), 392–400.
3.2 AI-Driven Automated ROI Selection
AI-driven automated ROI selection leverages standard H&E images to guide precise identification of biologically relevant regions. Pre-trained pathology models extract features from the images to automatically select ROIs that maximize molecular information content. This approach provides a reproducible, quantitative, and scalable method for integrating spatial information, enabling efficient capture of molecular heterogeneity across tissue sections.
A 2025 cover study in Nature Cell Biology introduced the "Smart Spatial omics (S2-omics)" workflow. Validation across multiple tissues—including breast, colon, kidney, liver, and stomach—and platforms such as Xenium, Visium HD, and CosMx demonstrated that ROIs selected by S2-omics often captured molecular heterogeneity as well as or better than expert manual annotation. The workflow reliably identifies tumor regions and tertiary lymphoid structures while accurately predicting cell types and cell–cell interaction networks with limited sampling. Quantifiable ROI scores further ensure maximum biological return on costly experiments, particularly in large-scale, multi-center studies.
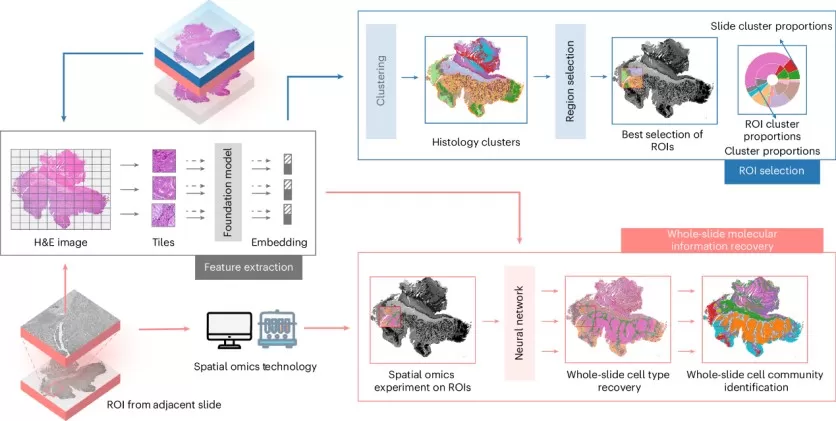
Figure 2. Workflow, ROI selection and cell label broadcasting of S2-omics. Image adapted from Yuan et al. (2025), Nature Cell Biology, 27(12), 2225–2238.
3.3 Multi-Modal Fusion for Spatial Segmentation
Multi-modal fusion for spatial segmentation combines MSI molecular data with H&E-derived tissue morphology to improve the accuracy of ROI delineation. Deep convolutional neural networks extract a "tissue morphology spectrum," enabling mutual validation between modalities and allowing ROI boundaries to be defined more reliably. This approach reduces subjectivity in algorithm parameter selection and provides a robust framework for integrating molecular and histological information.
A 2024 study published in GigaScience proposed this multi-modal fusion method, using Cohen's kappa to assess consistency between MSI-based and histology-based ROIs and selecting clustering parameters that maximized agreement. Only regions validated by both modalities were designated as reliable ROIs. In mouse kidney and tumor samples, ROIs generated with this approach closely matched actual anatomical labels. The core code has been made publicly available, enabling researchers to apply the method to their own datasets.
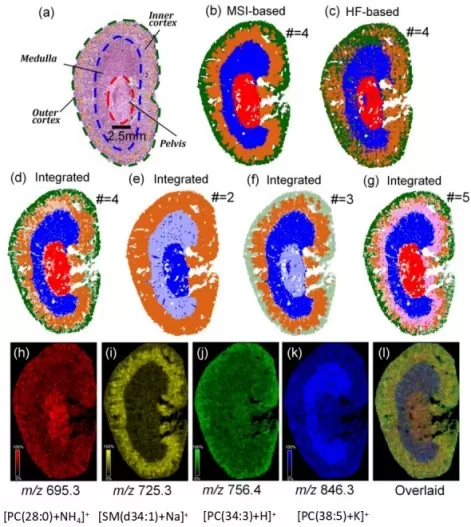
Figure 3. Automatic tissue segmentation by multi-modal fusion: mouse kidney ROI mapping and characteristic molecular distribution. Image adapted from Guo et al. (2022), GigaScience, 12, giad021.
4. ROI Selection Tools Across Mainstream Spatial Omics Platforms
Different spatial omics platforms provide specialized ROI selection capabilities, each optimized for distinct experimental workflows and tissue analysis objectives. These platforms vary in how regions of interest are defined, whether through morphology-guided annotation, automated segmentation, or bioinformatics-driven clustering. The following table summarizes the key ROI selection functionalities, their technical approaches, and the experimental scenarios in which they are most effectively applied, providing a clear comparison across mainstream spatial omics technologies.
| Platform/Technology | ROI Selection Features | Suitable Scenarios |
|---|---|---|
| NanoString GeoMx | Fluorescence-guided manual annotation and immunofluorescence-guided automated segmentation; ROIs can be geometric or free-form based on cellular morphology | Targeted region analysis using morphological markers, e.g., tumor infiltration margins or specific cell subpopulations |
| 10x Genomics Visium | ROI selection corresponds to pre-arranged capture points; HD version provides higher resolution | Unbiased exploration of tissue regions; suitable for discovering novel spatial structures |
| MALDI-MSI / DESI-MSI | Bioinformatics-based segmentation; SCiLS Lab and METASPACE offer spatially aware clustering algorithms | Discovery-driven studies; functional regions defined by molecular distribution patterns |
5. From ROI Selection to Biological Discovery
Selecting ROIs represents the initial step in spatial metabolomics, with the true biological value arising from detailed analysis of differences between regions.
Regional biomarker discovery leverages algorithms such as the "spatially aware nearest-shrinking centroid" to compute t-statistics for each ROI, enabling identification of metabolites that are highly specific to particular tissue regions. Higher t-values correspond to a greater contribution to regional specificity, supporting targeted hypotheses about tissue function and pathology.
Spatial heterogeneity in metabolic pathways is further revealed by mapping ROI-specific metabolites onto KEGG pathways or other metabolic databases. For example, in prostate cancer studies, comparisons between benign and malignant epithelial ROIs uncovered significant enrichment differences in glycolysis, fatty acid biosynthesis, and the TCA cycle. Such analyses highlight the spatial organization of metabolic reprogramming in tumors, providing critical insights for biomarker identification, pathway-based therapeutic strategies, and integrative multi-omics analyses.
6. ROI Selection Recommendations for Spatial Metabolomics Researchers
Beginner Principles
- Examine tissue staining first: H&E or immunofluorescence serves as the "first eye" for ROI selection. Understanding tissue morphology is fundamental to interpreting metabolic spatial patterns.
- Utilize segmentation tools: Run unsupervised segmentation in SCiLS Lab or METASPACE as a reference for ROI selection.
- Maintain buffer zones: Include a margin around ROI boundaries to prevent missing regions due to tissue deformation or instrument drift.
Expert Strategies
- Multi-modal cross-validation: Align molecular and morphological data and quantify ROI reliability using metrics such as Cohen's kappa.
- Integrate automated workflows: Incorporate AI tools like S2-omics into your pipeline to make ROI selection traceable, reproducible, and quantifiable.
- Omics-guided nested design: Follow Vandergrift et al.'s strategy, using non-destructive DESI-MSI to guide deep proteomics or spatial transcriptomics for multi-level information extraction on the same section.
From Better ROI Selection to Better Spatial Insights
As ROI selection in spatial metabolomics becomes more data-driven, reproducible, and multi-modal, researchers also need workflows that preserve spatial context and support confident biological interpretation. MetwareBio provides end-to-end spatial metabolomics support, from tissue preparation and MSI workflow design to data analysis and result interpretation, helping turn well-defined ROIs into more reliable biological insights.
Contact UsRead More: Related Guides for Spatial Metabolomics Workflows
Want to go beyond ROI selection? These articles cover the upstream and downstream decisions that shape spatial metabolomics results, from MSI platform choice and sample preparation to data analysis.
Spatial Metabolomics Explained: How It Works and Its Role in Cancer Research
Start with a practical overview of spatial metabolomics, including its core workflow, research value, and why spatial context matters for biological interpretation.
MALDI, DESI, or SIMS? How to Choose the Best MSI Techniques for Spatial Metabolomics
Compare major MSI platforms and see how instrument choice can influence ROI definition, spatial resolution, metabolite coverage, and study design.
How to Prepare Samples for Spatial Metabolomics: The Essential Guide You Need
Learn the sample preparation fundamentals that directly affect tissue integrity, signal quality, and the reliability of ROI-based spatial analysis.
Advanced Data Analysis in Spatial Metabolomics
Explore the analytical strategies used after ROI selection, including segmentation, visualization, and interpretation of region-specific metabolic patterns.
References
- Vandergrift, G. W., Veličković, M., Day, L. Z., Gorman, B. L., Williams, S. M., Shrestha, B., & Anderton, C. R. (2025). Untargeted Spatial Metabolomics and Spatial Proteomics on the Same Tissue Section. Analytical Chemistry, 97(1), 392–400. https://doi.org/10.1021/acs.analchem.4c04462
- Yuan, M., Jin, K., Yan, H., Schroeder, A., Luo, C., Yao, S., et al. (2025). Smart spatial omics (S2-omics) optimizes region of interest selection to capture molecular heterogeneity in diverse tissues. Nature Cell Biology, 27(12), 2225–2238. https://doi.org/10.1038/s41556-025-01811-w
- Guo, A., Chen, Z., Li, F., & Luo, Q. (2022). Delineating regions of interest for mass spectrometry imaging by multimodally corroborated spatial segmentation. GigaScience, 12, giad021. https://doi.org/10.1093/gigascience/giad021


