Proteomics experiments often yield hundreds or thousands of quantified proteins, but lists of fold changes and p-values rarely explain which biological programs are shifting. Pathway enrichment analysis addresses this gap by testing whether predefined gene or protein sets—such as pathways, cellular processes, or molecular functions—show coordinated evidence of perturbation. Two widely used strategies are over-representation analysis (ORA) and gene set enrichment analysis (GSEA). ORA asks whether a selected list of significant features is enriched for specific pathways, whereas GSEA evaluates enrichment across an entire ranked dataset without relying on a hard significance cutoff. Because the two methods use different inputs, null models, and assumptions, they can lead to different biological conclusions. Choosing the right approach is therefore essential for reliable interpretation of proteomics and other omics datasets. This article provides a detailed comparison of the two methods, practical guidance on when to apply each, and an overview of emerging alternatives that extend classical enrichment analysis.
1. OVER-REPRESENTATION ANALYSIS (ORA): PRINCIPLE AND WORKFLOW
ORA is the classic list-based approach to pathway enrichment. It asks a simple question: given a set of proteins selected as significant after differential analysis, do any curated pathways contain more of those proteins than expected by chance? That simplicity explains why ORA remains one of the most widely used functional interpretation strategies across genomics, transcriptomics, and proteomics (Khatri et al., 2012; Reimand et al., 2019).
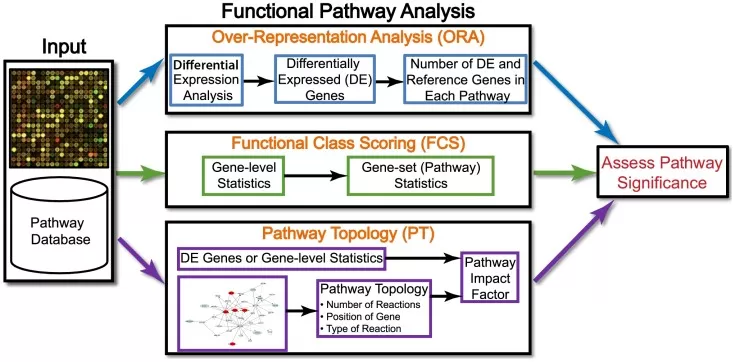
Figure 1. Overview of existing pathway analysis methods using gene expression data as an example. Image reproduced from Khatri et al., 2012, PLoS Computational Biology, 8(2), e1002375.
1.1 How ORA Works
A standard ORA workflow begins with differential analysis, followed by selection of a significant feature list using predefined thresholds such as adjusted p-value and fold change. For each pathway or gene set, the overlap between the selected proteins and the annotated members of the pathway is then evaluated with a 2 × 2 contingency table, most commonly using Fisher's exact test or an equivalent hypergeometric formulation. The resulting p-values are corrected for multiple testing, typically with the Benjamini-Hochberg false discovery rate (FDR) procedure (Khatri et al., 2012; Sherman et al., 2022). (Learn more at: p-Value vs FDR in Omics: Adjusted p-Value and q-Value Explained)
One critical but often underreported step is the definition of the background universe. In proteomics, the background should usually be the proteins that were confidently quantified and eligible for statistical testing in that experiment—not the entire genome or proteome. An inappropriate background can materially distort ORA results, especially when coverage is incomplete or biased toward abundant proteins (Wijesooriya et al., 2022; Zhao and Rhee, 2023).
1.2 Key Strengths of ORA
ORA is fast, computationally lightweight, and easy to explain to non-specialists. It is particularly useful when a study already has a clear list of strongly changing proteins and the immediate goal is to summarize those hits at the pathway level. The method is also supported by a mature software ecosystem, including DAVID, g:Profiler, and clusterProfiler, which makes it easy to integrate into routine bioinformatics workflows (Sherman et al., 2022; Kolberg et al., 2023; Wu et al., 2021).
1.3 Limitations and Common Pitfalls of ORA
The main limitation of ORA is threshold dependence. A protein just above a significance cutoff is treated as a full hit, while a protein just below the cutoff is excluded entirely. This binary decision discards rank information and effect-size gradients that may still be biologically meaningful. ORA also does not inherently distinguish pathway activation from pathway repression unless upregulated and downregulated feature lists are analyzed separately (Subramanian et al., 2005; Khatri et al., 2012).
ORA is also sensitive to annotation redundancy and pathway size. Broad, overlapping gene sets can dominate the results and create the illusion of multiple independent findings when the signal is actually concentrated in one biological theme. For this reason, ORA should be interpreted as a starting point for biological summarization rather than as definitive proof of pathway activation (Reimand et al., 2019; Wijesooriya et al., 2022).
2. GENE SET ENRICHMENT ANALYSIS (GSEA): PRINCIPLE AND WORKFLOW
GSEA was developed to address the information loss that occurs when a ranked omics dataset is reduced to a binary list of significant hits. Instead of testing only a filtered subset, GSEA evaluates whether members of a predefined gene set are preferentially concentrated near the top or bottom of a ranked feature list. Although the original method was introduced for transcriptomics, the same ranked-set logic is widely applied to proteomics once quantified proteins are mapped to pathway annotations (Subramanian et al., 2005; Zhao and Rhee, 2023).
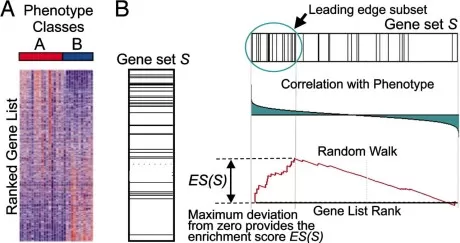
Figure 2. A GSEA overview illustrating the method. Image reproduced from Subramanian et al., 2005, Proceedings of the National Academy of Sciences of the United States of America, 102(43), 15545–15550.
2.1 How GSEA Works
In a typical GSEA workflow, all retained proteins are ranked by a statistic that captures their association with the phenotype of interest—for example, signal-to-noise ratio, t statistic, or log2 fold change. For each pathway, GSEA walks down the ranked list with a running-sum statistic, increasing the score when a pathway member is encountered and decreasing it otherwise. The maximum deviation from zero is the enrichment score (ES). Because raw ES values are influenced by gene set size, they are normalized to produce a normalized enrichment score (NES), and significance is estimated through permutation testing (Subramanian et al., 2005).
Permutation strategy matters. In classic two-class GSEA, phenotype permutation is generally preferred when sample sizes are adequate, whereas gene-set permutation or preranked workflows are more common when sample numbers are limited or when the ranking statistic is generated outside the GSEA software. These choices affect the null model and therefore the interpretation of FDR thresholds (Reimand et al., 2019; Zhao and Rhee, 2023).
2.2 Interpreting GSEA Results
A positive NES indicates that pathway members accumulate near the top of the ranked list, whereas a negative NES indicates enrichment near the bottom. The leading-edge subset pinpoints the core members that contribute most strongly to the enrichment signal. This can be highly informative in proteomics because it helps distinguish broad pathway labels from the specific proteins that are driving the result (Subramanian et al., 2005).
2.3 What GSEA Captures That ORA Often Misses
GSEA is especially useful when many proteins in a pathway move modestly but coherently in the same direction. Such coordinated, distributed shifts are common in signaling, stress-response, and immune pathways, yet they may be missed by ORA if too few individual proteins cross a significance threshold. By preserving the ranked structure of the dataset, GSEA can detect pathway-level regulation that remains invisible in a threshold-based analysis (Subramanian et al., 2005; Khatri et al., 2012).
That said, GSEA is not automatically more rigorous than ORA. Its conclusions depend on the quality of the ranking statistic, the handling of ties and missing values, the permutation scheme, and the gene-set database used. GSEA does not require simple imputation by definition; rather, it requires a defensible ranked list of retained features. In proteomics, preprocessing decisions should therefore be documented just as carefully as the enrichment method itself (Reimand et al., 2019; Wijesooriya et al., 2022).
3. ORA VS GSEA: A DIRECT COMPARISON
ORA and GSEA answer related but non-identical questions. ORA asks whether a selected hit list is enriched for pathways; GSEA asks whether a pathway shows a non-random distribution across an entire ranked dataset. This difference has practical consequences for sensitivity, interpretability, and reproducibility (Khatri et al., 2012; Subramanian et al., 2005).
| Feature | ORA | GSEA |
|---|---|---|
| Input data | Significant protein list | All retained proteins ranked by a continuous statistic |
| Threshold dependence | High | Low |
| Statistical framework | Contingency table with Fisher/hypergeometric test | Running-sum enrichment statistic with permutation testing |
| Sensitivity to effect size | Limited | Retains rank and weighting information |
| Direction of regulation | Requires separate up/down analyses | Directly reflected by NES sign |
| Detection of coordinated weak shifts | Often poor | Strong |
| Background dependence | Very high | Still important, but typically less explicit than in ORA |
| Computational cost | Low | Moderate |
| Interpretability | Very intuitive | More nuanced, but often more informative |
In practice, ORA tends to highlight the most obvious pathway signals among already significant proteins, while GSEA is better at capturing pathway-wide shifts that are statistically diffuse. Deep, well-quantified proteomics datasets often benefit from GSEA, whereas smaller or more targeted studies may still favor the transparency and speed of ORA. In many real projects, the most robust interpretation comes from comparing both views rather than treating them as mutually exclusive alternatives.
4. CHOOSING BETWEEN ORA AND GSEA: A PRACTICAL GUIDE
4.1 When ORA Is the Better Fit
ORA is a strong choice when the analysis already yields a confident list of differentially abundant proteins and the goal is to summarize those hits quickly. It is also useful in targeted proteomics or other lower-dimensional datasets where the ranked list is short, making permutation-based enrichment less stable or less informative. When the biological question centers on the most strongly altered proteins, ORA often provides the cleanest first pass (Khatri et al., 2012; Zhao and Rhee, 2023).
4.2 When GSEA Is the Better Fit
GSEA is usually preferable when pathway-level behavior matters more than the exact boundary of a significant hit list. That includes studies of signaling, drug response, stress adaptation, immune activation, and other settings in which coordinated moderate shifts may be biologically meaningful. GSEA is also valuable when the cutoff for "significance" would otherwise feel arbitrary or unstable across related comparisons (Subramanian et al., 2005; Reimand et al., 2019).
4.3 Why Many Studies Run Both
Running ORA and GSEA in parallel is often the most informative strategy. Pathways supported by both methods usually represent the most reproducible biological themes. Pathways unique to ORA may reflect strong, sparse signals concentrated in a few proteins, whereas pathways unique to GSEA may indicate broader but subtler regulation. Reporting this convergence and divergence is often more transparent than presenting either method as the single "correct" answer (Khatri et al., 2012; Zhao and Rhee, 2023).
5. BEYOND ORA AND GSEA: USEFUL EXTENSIONS
5.1 ssGSEA and Sample-Level Pathway Scoring
Single-sample GSEA (ssGSEA) extends the GSEA concept from group comparisons to individual samples, generating one enrichment score per sample-gene set pair. This is useful for clustering, patient stratification, longitudinal profiling, and multi-sample visualization of pathway activity, especially when heterogeneity within a cohort is biologically important (Barbie et al., 2009).
5.2 Correlation-Aware Competitive Tests
Methods such as CAMERA address a common statistical issue in enrichment analysis: genes or proteins within the same pathway are often correlated, and ignoring that correlation can inflate significance. CAMERA explicitly adjusts for inter-gene correlation and is particularly relevant when the goal is rigorous differential pathway testing under a linear-model framework (Wu and Smyth, 2012).
5.3 Topology-Aware Methods
Topology-aware methods such as SPIA go beyond simple membership testing by considering how pathway components interact. In principle, this can yield more mechanistic insight than ORA or GSEA alone because activation, inhibition, and network position are incorporated into the analysis. These methods are most attractive when curated pathway topology is directly relevant to the biological question (Tarca et al., 2009).
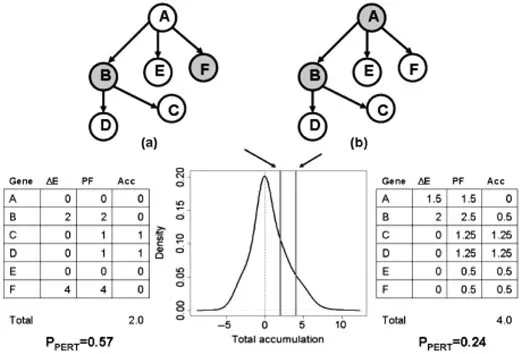
Figure 3. Capturing the topology of the pathways and the position of the gene through the perturbation analysis. Image reproduced from Tarca et al., 2009, Bioinformatics, 25(1), 75–82.
6. BEST PRACTICES FOR PATHWAY ENRICHMENT ANALYSIS
- Define the correct background universe. For ORA in particular, the background should reflect the proteins that were actually measurable and testable in the experiment, not the entire proteome by default (Wijesooriya et al., 2022; Zhao and Rhee, 2023).
- Choose current and biologically relevant annotation resources. Database versioning matters, and it should be reported alongside the analysis. Common resources include MSigDB and tool ecosystems such as DAVID, g:Profiler, and clusterProfiler (Liberzon et al., 2011; Kolberg et al., 2023; Sherman et al., 2022; Wu et al., 2021).
- Document identifier mapping. Proteomics data are often mapped onto gene-centric pathway resources, so the ID conversion strategy should be explicit and reproducible (Reimand et al., 2019; Zhao and Rhee, 2023).
- Report the parameters that materially affect interpretation, including ranking metric, permutation type, minimum and maximum gene-set size, multiple-testing correction, and FDR threshold (Reimand et al., 2019; Wijesooriya et al., 2022).
- Manage redundancy in enriched terms. Hierarchical resources such as Gene Ontology can produce many overlapping hits, so summarization, clustering, or network-based visualization is often needed before biological interpretation (Reimand et al., 2019; Wu et al., 2021).
- Treat enrichment as hypothesis generation, not final proof. Pathway results should be integrated with the underlying protein-level evidence, prior biology, and, when possible, orthogonal validation experiments (Khatri et al., 2012; Zhao and Rhee, 2023).
How MetwareBio Supports Proteomics Interpretation
Pathway enrichment is only as reliable as the data and preprocessing that feed into it. MetwareBio provides proteomics, metabolomics, lipidomics, and integrated multi-omics services designed to support hypothesis generation and biological interpretation from high-dimensional datasets. Its public service portfolio includes DIA and DDA quantitative proteomics as well as multi-omics combinations such as proteomics + metabolomics and transcriptomics + proteomics + metabolomics.
For teams that want to move from raw abundance tables to interpretable biology more efficiently, the Metware Cloud Platform offers re-analysis and visualization tools that can complement enrichment workflows. Contact us to discuss study design, data generation, or downstream interpretation for your next proteomics or multi-omics project.
Contact UsPathway and Multivariate Analysis for Proteomics and Multi-Omics
Enrichment analysis is only one step in a broader workflow—from careful statistical testing and feature screening through pathway interpretation and multivariate exploration. These related guides walk you through the methods that most often accompany ORA and GSEA in real proteomics and multi-omics projects.
After running ORA or GSEA, learn how to read the output: understand Gene Count, Rich Factor, p-values, and FDR-adjusted significance to build biologically meaningful pathway narratives from your enrichment results.
A solid enrichment analysis starts with sound differential expression testing. Review the practical choices between t-tests, ANOVA, and non-parametric alternatives, including guidance on normality testing and effect size estimation for proteomics datasets.
Whether you use ORA or GSEA, understanding how fold change, FDR, and VIP work together helps you avoid over-relying on any single metric when building candidate feature lists from omics data.
Complement pathway enrichment with correlation analysis. Learn when Pearson vs Spearman is appropriate, how hierarchical clustering adds biological insight, and how heatmaps reveal coordinated behavior across proteins or metabolites.
Like GSEA, multivariate methods preserve full-rank information without hard thresholds. Compare unsupervised PCA for exploration with supervised PLS-DA and OPLS-DA for discriminant analysis and biomarker discovery in proteomics and metabolomics.
Go beyond pathway lists with weighted gene co-expression network analysis. WGCNA identifies modules of coordinated proteins or genes and links them to phenotypes—a powerful complement to both ORA and GSEA for hypothesis generation in omics studies.
References
- Barbie, D. A., Tamayo, P., Boehm, J. S., Kim, S. Y., Moody, S. E., Dunn, I. F., Schinzel, A. C., Sandy, P., Meylan, E., Scholl, C., Fröhling, S., Chan, E. M., Sos, M. L., Michel, K., Mermel, C., Silver, S. J., Weir, B. A., Reiling, J. H., Sheng, Q., ... Hahn, W. C. (2009). Systematic RNA interference reveals that oncogenic KRAS-driven cancers require TBK1. Nature, 462(7269), 108–112. https://doi.org/10.1038/nature08460
- Khatri, P., Sirota, M., & Butte, A. J. (2012). Ten years of pathway analysis: Current approaches and outstanding challenges. PLoS Computational Biology, 8(2), e1002375. https://doi.org/10.1371/journal.pcbi.1002375
- Kolberg, L., Raudvere, U., Kuzmin, I., Adler, P., Vilo, J., & Peterson, H. (2023). g:Profiler—interoperable web service for functional enrichment analysis and gene identifier mapping (2023 update). Nucleic Acids Research, 51(W1), W207–W212. https://doi.org/10.1093/nar/gkad347
- Liberzon, A., Subramanian, A., Pinchback, R., Thorvaldsdóttir, H., Tamayo, P., & Mesirov, J. P. (2011). Molecular signatures database (MSigDB) 3.0. Bioinformatics, 27(12), 1739–1740. https://doi.org/10.1093/bioinformatics/btr260
- Reimand, J., Isserlin, R., Voisin, V., Kucera, M., Tannus-Lopes, C., Rostamianfar, A., Wadi, L., Meyer, M., Wong, J., Xu, C., Merico, D., & Bader, G. D. (2019). Pathway enrichment analysis and visualization of omics data using g:Profiler, GSEA, Cytoscape and EnrichmentMap. Nature Protocols, 14(2), 482–517. https://doi.org/10.1038/s41596-018-0103-9
- Sherman, B. T., Hao, M., Qiu, J., Jiao, X., Baseler, M. W., Lane, H. C., Imamichi, T., & Chang, W. (2022). DAVID: a web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Research, 50(W1), W216–W221. https://doi.org/10.1093/nar/gkac194
- Subramanian, A., Tamayo, P., Mootha, V. K., Mukherjee, S., Ebert, B. L., Gillette, M. A., Paulovich, A., Pomeroy, S. L., Golub, T. R., Lander, E. S., & Mesirov, J. P. (2005). Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proceedings of the National Academy of Sciences of the United States of America, 102(43), 15545–15550. https://doi.org/10.1073/pnas.0506580102
- Tarca, A. L., Draghici, S., Khatri, P., Hassan, S. S., Mittal, P., Kim, J.-S., Kim, C. J., Kusanovic, J. P., & Romero, R. (2009). A novel signaling pathway impact analysis. Bioinformatics, 25(1), 75–82. https://doi.org/10.1093/bioinformatics/btn577
- Wijesooriya, K., Jadaan, S. A., Perera, K. L., Kaur, T., & Ziemann, M. (2022). Urgent need for consistent standards in functional enrichment analysis. PLoS Computational Biology, 18(3), e1009935. https://doi.org/10.1371/journal.pcbi.1009935
- Wu, D., & Smyth, G. K. (2012). Camera: a competitive gene set test accounting for inter-gene correlation. Nucleic Acids Research, 40(17), e133. https://doi.org/10.1093/nar/gks461
- Wu, T., Hu, E., Xu, S., Chen, M., Guo, P., Dai, Z., Feng, T., Zhou, L., Tang, W., Zhan, L., Fu, X., Liu, S., Bo, X., & Yu, G. (2021). clusterProfiler 4.0: A universal enrichment tool for interpreting omics data. The Innovation, 2(3), 100141. https://doi.org/10.1016/j.xinn.2021.100141
- Zhao, K., & Rhee, S. Y. (2023). Interpreting omics data with pathway enrichment analysis. Trends in Genetics, 39(4), 308–319. https://doi.org/10.1016/j.tig.2023.01.003


