High-throughput technologies such as proteomics, metabolomics, and lipidomics assays have enabled comprehensive profiling of biomolecules across diverse biological systems. However, the resulting datasets, which are typically characterized by numerous variables relative to limited sample sizes, present substantial analytical challenges. Multivariate analysis methods have become indispensable tools for extracting biologically meaningful patterns from such complex data. Among these, Partial Least Squares Discriminant Analysis (PLS-DA) and its derivative Orthogonal Partial Least Squares Discriminant Analysis (OPLS-DA) stand as two of the most widely implemented supervised techniques in omics research. This article provides a rigorous comparison of these methods, elucidating their respective principles, advantages, limitations, and appropriate applications to guide researchers in selecting the most suitable approach for their specific analytical objectives.
1. Why Multivariate Analysis is Essential for Omics Data
Modern omics technologies—including metabolomics, proteomics, and lipidomics—routinely generate datasets comprising thousands of measured features from relatively few biological samples. This high-dimensional structure, characterized by multicollinearity among variables and the "large p, small n" problem, renders conventional univariate statistical approaches inadequate for capturing the complex interrelationships inherent in biological systems. Multivariate analysis addresses this challenge by reducing data dimensionality while preserving critical information about sample groupings and biochemical patterns.
Multivariate methods are broadly categorized into unsupervised and supervised techniques. Unsupervised methods, such as Principal Component Analysis (PCA) and hierarchical clustering, explore inherent data structure without utilizing class labels. These approaches are invaluable for initial data inspection, outlier detection, and visualization of natural groupings. However, because they disregard experimental design information, unsupervised methods may overlook subtle but biologically meaningful patterns associated with specific phenotypes. Supervised multivariate methods, by contrast, incorporate prior knowledge of class membership—such as disease versus control groups, treatment conditions, or time points—to build models that maximize separation between predefined classes. This category includes techniques like Partial Least Squares Discriminant Analysis (PLS-DA), Orthogonal Partial Least Squares Discriminant Analysis (OPLS-DA), and various machine learning algorithms including support vector machines and random forests. Among these, PLS-DA and OPLS-DA have gained particular prominence in omics research due to their ability to handle highly collinear data, their inherent dimensionality reduction capabilities, and their interpretable outputs that facilitate biomarker discovery. (Learn more at: PCA vs PLS-DA vs OPLS-DA)
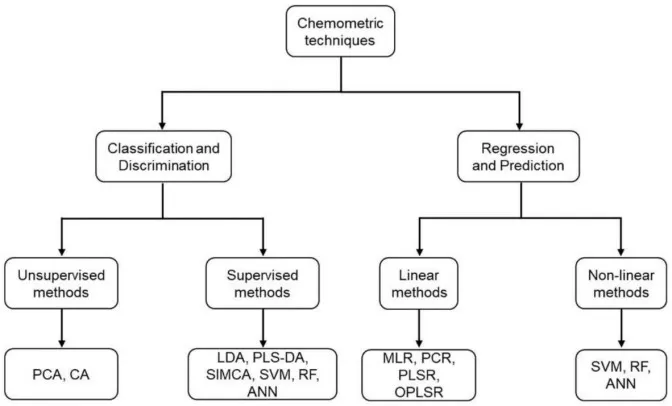
Figure 1. Classification of the chemometric techniques commonly used in food science. Image reproduced from González-Domínguez (2022), Foods, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
2. PLS‑DA: Principles, Limitations and Applications in Omics
As a foundational supervised multivariate method in chemometrics, Partial Least Squares Discriminant Analysis (PLS-DA) has become a cornerstone of omics data analysis. The method's widespread adoption stems from its ability to handle the high-dimensional, multicollinear nature of modern biological datasets while explicitly incorporating class information.
2.1 Concept and Applications of PLS‑DA
Partial Least Squares Discriminant Analysis extends the principles of Partial Least Squares regression to classification problems. The method constructs latent variables—linear combinations of the original predictors—that maximize the covariance between the predictor matrix (X) and a binary response variable (Y) indicating class membership. This approach effectively combines dimensionality reduction with supervised classification, making it particularly suited for omics data where variables far outnumber samples.
PLS-DA has found widespread application across diverse omics disciplines. In metabolomics, the method enables discrimination between phenotypic states and identification of differentially abundant metabolites (Ren et al., 2026). Proteomic studies employ PLS-DA to distinguish disease subtypes based on protein expression patterns, while lipidomic investigations utilize the technique to characterize lipid signatures associated with specific physiological conditions. The method's capacity to handle multicollinear predictors—a common feature of spectroscopic and spectrometric data—has established it as a cornerstone of chemometric analysis in food authenticity assessment, clinical diagnostics, and environmental monitoring (Rivera-Pérez et al., 2022).
2.2 Strengths and Limitations of PLS‑DA
The principal strength of PLS-DA lies in its robust handling of ill-conditioned data matrices. Unlike classical discriminant methods that require matrix inversion, PLS-DA effectively manages situations where the number of predictors exceeds the number of observations. The algorithm's iterative structure provides computational efficiency and numerical stability even with highly collinear variables.
However, PLS-DA carries important limitations that warrant consideration. The method's supervised nature renders it susceptible to overfitting, particularly when model complexity is not rigorously controlled. Spurious class separations can emerge from random noise when cross-validation is inadequate or when the number of features vastly exceeds sample size (Bevilacqua and Bro, 2020). Furthermore, model interpretation can prove challenging because biological signal and systematic noise become intermingled within the same latent variables, obscuring the distinction between class-related variation and orthogonal technical artifacts.
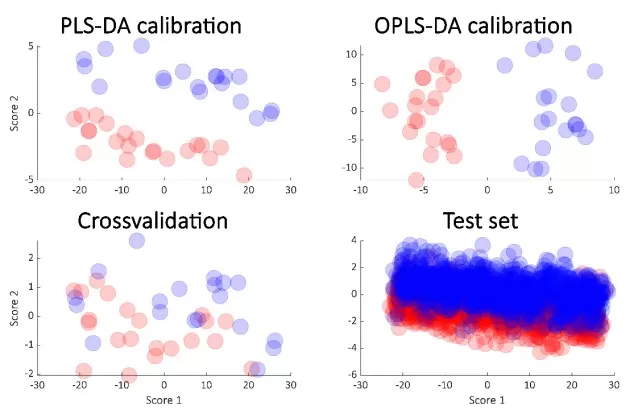
Figure 2. Calibrated score plot from PLS-DA (upper left) and the OPLS-DA (orthogonal PLS-DA) (upper right) from the simulated data. The cross-validated scores from PLS-DA (lower left) and the score plot obtained from the test samples (lower right). Image reproduced from Bevilacqua and Bro (2020), Metabolites, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
3. OPLS‑DA: Advancing PLS‑DA for Cleaner Interpretation
While PLS-DA effectively models covariance between predictors and response, its interpretive capacity is constrained by the mixing of predictive and non-predictive variation within the initial latent variables. Biological differences of interest become entangled with systematic variation unrelated to class distinction—such as batch effects, instrument drift, or physiological variation orthogonal to the phenotype under investigation. This mixing complicates model interpretation and biomarker selection. To address this limitation, Trygg and Wold introduced Orthogonal Partial Least Squares Discriminant Analysis (OPLS-DA) in 2002, incorporating an orthogonal signal correction filter that separates predictive variation from structured noise.
3.1 Methodological Improvements and Applications of OPLS‑DA
OPLS-DA extends PLS-DA by decomposing the predictor matrix (X) into three components: predictive variation correlated with the response (Y), orthogonal variation uncorrelated with Y, and residual noise. This orthogonal signal correction filters out systematic variation unrelated to class membership, concentrating all class-related information into a single predictive component (for two-class comparisons). Orthogonal components capture structured noise such as batch effects or physiological variation unrelated to the phenotype.
The method has found broad application across omics disciplines. In metabolomics, OPLS-DA enables visualization of disease-related metabolic perturbations while filtering confounding variation. Proteomic studies leverage the technique to distinguish cancer subtypes, with orthogonal components absorbing technical artifacts. Recent applications include distinguishing food geographical origins, identifying volatile markers differentiating processing conditions, and identifying disease-related metabolic signatures and biomarkers (Rivera-Pérez et al., 2022; Lyu et al., 2025).
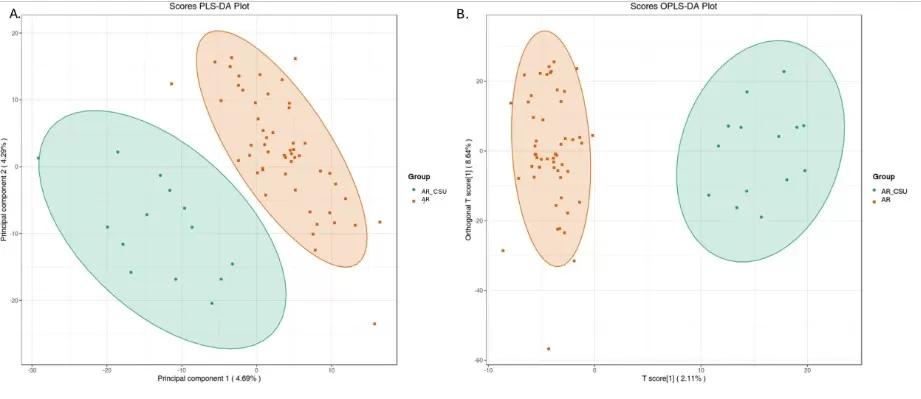
Figure 3. Multivariate statistical analysis of metabolic data from AR and AR_CSU patients. (A) The score plot generated by Partial Least Squares Discriminant Analysis (PLS-DA) and (B) the score plot produced by Orthogonal Partial Least Squares Discriminant Analysis (OPLS-DA) clearly illustrate distinct metabolomic profiles between AR and AR_CSU patients. Image reproduced from Lyu et al. (2025), Frontiers in Immunology, licensed under the Creative Commons Attribution 4.0 International License (CC BY 4.0).
3.2 Advantages and Limitations of OPLS‑DA
The principal advantage of OPLS-DA lies in enhanced interpretability. By isolating predictive from orthogonal variation, the method provides clearer visualization of class separation—score plots display discriminatory patterns without contamination from structured noise. The S-plot, unique to OPLS-DA, combines covariance with correlation to facilitate confident biomarker nomination. Variables in extreme S-plot quadrants exhibit both high discriminatory power and high consistency across samples.
However, OPLS-DA does not improve predictive performance over PLS-DA; with equivalent components, both methods yield identical prediction accuracy. The technique remains susceptible to overfitting in high-dimensional datasets, requiring rigorous validation through cross-validation and permutation testing. Additionally, orthogonal components may absorb biologically relevant variation uncorrelated with class distinction, warranting careful examination for unexpected biological structure.
4. PLS‑DA vs OPLS‑DA: Comparison and Decision Guide
While PLS-DA and OPLS-DA share fundamental algorithmic roots and produce equivalent predictive performance when equivalent components are considered, their structural differences carry important implications for data interpretation and research application. The table below summarizes key distinctions between these methods.
| Feature | PLS-DA | OPLS-DA |
|---|---|---|
| Latent Variable Decomposition | Predictive and orthogonal variation mixed within same components | Predictive and orthogonal variation separated into distinct components |
| Model Interpretability | Moderate; class separation may appear across multiple components | Enhanced; predictive information concentrated in first component |
| Handling of Orthogonal Variation | Orthogonal noise incorporated into predictive components | Orthogonal noise explicitly modeled and removed |
| Visualization Clarity | Score plots may obscure separation due to mixed noise | Score plots display clean separation in first predictive component |
| Risk of Overfitting | Substantial; requires rigorous cross-validation and permutation testing | Substantial; requires identical validation rigor as PLS-DA |
| Feature Selection Tools | VIP scores identify important variables | S-plots combine covariance and correlation for marker selection |
| Predictive Performance | Benchmark predictive capacity | Equivalent to PLS-DA with same number of components |
The choice between PLS-DA and OPLS-DA should be guided by the research objective. For purely predictive applications where accurate classification of unknown samples constitutes the primary goal, PLS-DA provides adequate performance with simpler implementation. However, when biological interpretation and reliable biomarker identification represent central aims, OPLS-DA's separation of predictive and orthogonal variation offers substantial advantages. The method's enhanced interpretability and specialized visualization tools support more confident identification of features genuinely associated with the phenotype of interest.
5. Implementing PLS‑DA and OPLS‑DA in Omics Studies
Successful application of supervised multivariate methods requires adherence to a systematic workflow that ensures model validity and biological relevance. The following steps outline a robust approach to implementing PLS-DA and OPLS-DA in omics data analysis.
Step 1: Data Quality Assessment with PCA
Before undertaking supervised analysis, examination of data structure through unsupervised Principal Component Analysis (PCA) provides essential quality control. PCA reveals natural clustering patterns, identifies potential outliers, and detects batch effects or other systematic technical variation. Samples exhibiting extreme distances within or orthogonal to the projection plane warrant investigation before proceeding to supervised modeling. This preliminary step ensures that subsequent supervised analyses rest upon sound data foundations.
Step 2: Initial Model Construction with PLS-DA
PLS-DA model construction begins with selection of an appropriate number of latent variables through cross-validation. The Q² metric—indicating predictive ability estimated through cross-validation—guides this selection. Rigorous validation includes permutation testing, wherein class labels are randomly reassigned and model performance recalculated across numerous iterations. This procedure assesses whether observed discrimination exceeds that expected by chance. A valid model demonstrates Q² values substantially higher than those obtained from permuted data.
Step 3: Model Optimization and Interpretation with OPLS-DA
When model validation confirms meaningful discrimination, OPLS-DA provides enhanced interpretive capabilities. The method's decomposition into predictive and orthogonal components enables clearer visualization of group separation and more refined feature selection. S-plots generated from OPLS-DA models combine covariance (magnitude of variable contribution) with correlation (reliability of that contribution). Variables appearing in the extreme upper-right and lower-left quadrants of the S-plot—exhibiting both high magnitude and high reliability—represent candidates for biomarker nomination. Variable Importance in Projection (VIP) scores, available for both PLS-DA and OPLS-DA, further support feature selection by quantifying each variable's contribution to the model.
Recent applications demonstrate the power of this approach. Rivera-Pérez and colleagues (2022) employed PLS-DA to discriminate thyme samples based on geographical origin, achieving perfect classification accuracy. Subsequent OPLS-DA enabled discrimination of sterilized versus non-sterilized samples and facilitated identification of 24 volatile markers distinguishing these processing conditions.
Step 4: Model Validation
Final model validation demands rigorous assessment of predictive performance on independent test data when available. Cross-validated scores provide more realistic visualization of model performance than calibrated scores, particularly when R² substantially exceeds Q². Researchers should remain cognizant that calibrated score plots can present misleadingly optimistic impressions of class separation, and cross-validated or test set projections offer more scientifically meaningful representations.
6. Avoiding Common Pitfalls in PLS‑DA and OPLS‑DA Analysis
Despite their utility, PLS-DA and OPLS-DA carry inherent risks that demand careful attention throughout the analytical process.
Avoiding Misapplication: OPLS-DA cannot salvage fundamentally non-discriminatory data. If two classes lack meaningful differences detectable by PLS-DA, OPLS-DA will not produce valid discrimination. The method separates predictive from orthogonal variation but cannot create predictive information where none exists. Initial PLS-DA assessment remains essential to establish the presence of genuine class-related signal.
Overfitting Risk in High-Dimensional Data: The "large p, small n" paradigm endemic to omics research creates substantial overfitting potential. Complex models can achieve perfect classification of calibration samples while generalizing poorly to new observations. Overfitting manifests as substantial discrepancy between R² (calibration performance) and Q² (cross-validated performance). Rigorous validation through cross-validation, permutation testing, and independent test sets provides essential protection against spurious findings.
Data Preprocessing Effects: Scaling decisions profoundly influence model outcomes. Unit variance (UV) scaling assigns equal weight to all variables regardless of original magnitude, while Pareto scaling reduces but does not eliminate the influence of large-magnitude features. The choice among scaling approaches should reflect the underlying biology and measurement characteristics. Additionally, transformations addressing heteroscedasticity or non-normality may improve model performance and interpretability.
Interpretation of VIP Scores and S-plots: While VIP scores and S-plots support feature selection, statistical significance does not guarantee biological relevance. Selected features require independent validation and biological context for meaningful interpretation. Furthermore, VIP thresholds for feature selection remain somewhat arbitrary; researchers should consider multiple criteria and assess consistency across validation strategies.
7. Future Trends in Supervised Multivariate Omics Analysis
PLS-DA and OPLS-DA represent powerful and complementary tools for supervised analysis of omics data. PLS-DA provides robust classification capabilities with straightforward implementation, suitable for predictive applications where accurate sample categorization constitutes the primary objective. OPLS-DA, through its separation of predictive and orthogonal variation, offers enhanced interpretability and refined biomarker selection capabilities, making it particularly valuable for mechanistic investigation and feature discovery.
Both methods share fundamental requirements: rigorous validation, transparent reporting of performance metrics, and cautious interpretation grounded in biological context. The distinction between R² and Q², the results of permutation testing, and the use of cross-validated projections provide essential safeguards against overinterpretation. Researchers should select between these methods based on their specific objectives—prediction accuracy for PLS-DA, interpretive clarity for OPLS-DA—while maintaining consistent validation rigor regardless of choice.
Looking forward, the integration of PLS-DA and OPLS-DA with emerging analytical approaches promises expanded capabilities. Combination with machine learning algorithms may enhance predictive performance while preserving interpretability. Multi-omics integration, wherein complementary data types (genomics, transcriptomics, proteomics, metabolomics) are analyzed jointly, represents a frontier for supervised multivariate analysis. As these methods evolve, their fundamental strengths—handling high-dimensional correlated data, revealing class structure, and supporting biological interpretation—will ensure their continued centrality to omics research.
Explore MetwareBio’s Omics Analysis Services
MetwareBio provides integrated proteomics, metabolomics, and multi-omics analysis services to support biomarker discovery, pathway analysis, mechanism research, and molecular profiling. With advanced technology platforms and rigorous data analysis workflows, we help researchers generate reliable, high-quality insights from complex omics data.
Interested in omics analysis services for your next project? Contact MetwareBio to discuss your research goals and find the right solution for your study.
Contact UsReferences
- González-Domínguez, R., Sayago, A., & Fernández-Recamales, Á. (2022). An Overview on the Application of Chemometrics Tools in Food Authenticity and Traceability. Foods (Basel, Switzerland), 11(23), 3940. https://doi.org/10.3390/foods11233940
- Ren, Y., Zheng, T., Liu, R., Zhao, Y., Zhang, Z., Si, T., & Zhang, R. (2026). Development of a machine learning-based prediction model for bipolar disorder relapse via integration of 1H-NMR metabolomics and clinical features. Journal of Affective Disorders, 396, 120858. https://doi.org/10.1016/j.jad.2025.120858
- Rivera-Pérez, A., Romero-González, R., & Garrido Frenich, A. (2022). Fingerprinting based on gas chromatography-Orbitrap high-resolution mass spectrometry and chemometrics to reveal geographical origin, processing, and volatile markers for thyme authentication. Food Chemistry, 393, 133377. https://doi.org/10.1016/j.foodchem.2022.133377
- Bevilacqua, M., & Bro, R. (2020). Can We Trust Score Plots?. Metabolites, 10(7), 278. https://doi.org/10.3390/metabo10070278
- Lyu, X., Liu, Y., Zheng, H., Li, H., Wu, Z., Sun, Y., Wu, S., Jiang, X., Wu, S., Tang, R., Gao, Y., & Sun, J. (2025). Distinct metabolomic signatures in allergic rhinitis with concurrent chronic spontaneous urticaria: an untargeted metabolomics analysis reveals novel biomarkers and pathway alterations. Frontiers in Immunology, 16, 1555664. https://doi.org/10.3389/fimmu.2025.1555664


