Cholesterol is an essential structural lipid and metabolic precursor that supports membrane organization, steroid hormone biosynthesis, bile acid production, and embryonic development. Its cellular and systemic homeostasis is maintained through tightly coordinated biosynthesis, dietary absorption, intracellular trafficking, storage, and efflux. Disruption of this balance contributes not only to atherosclerotic cardiovascular disease, but also to cancer, neurodegeneration, and metabolic liver disease. Accordingly, cholesterol metabolism has become a central topic in both basic and translational research. Recent advances in lipidomics, metabolomics, proteomics, and multi-omics integration are further expanding our understanding of cholesterol biology at molecular and systems levels. This review summarizes the production, metabolism, physiological functions, and disease relevance of cholesterol, with particular emphasis on emerging omics approaches for cholesterol research.
1. WHAT IS CHOLESTEROL? STRUCTURE AND BIOSYNTHESIS
Cholesterol is a 27-carbon sterol with a rigid tetracyclic ring system, a hydroxyl group at C-3, and a branched aliphatic side chain. This amphipathic architecture explains why cholesterol is especially well suited to membranes: the hydroxyl group aligns near polar head groups, whereas the hydrophobic ring system and side chain pack within the lipid bilayer.
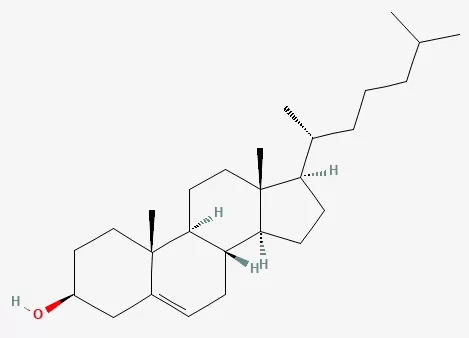
Figure 1. The molecular structure of cholesterol. Image reproduced from PubChem.
1.1 Biosynthesis: The Mevalonate Pathway
In mammalian cells, cholesterol is synthesized primarily in the liver and intestine through the mevalonate pathway. Acetyl-CoA is converted through a series of enzymatic steps to HMG-CoA and then to mevalonate by HMG-CoA reductase (HMGCR), the canonical rate-limiting enzyme of the pathway. Downstream reactions generate isoprenoids, squalene, lanosterol, and ultimately cholesterol (Goldstein & Brown, 2015; Luo et al., 2020; Duan et al., 2022).
The pathway is tightly regulated at multiple levels:
- Transcriptional control: Sterol regulatory element-binding proteins (SREBPs) activate expression of biosynthetic genes when cellular cholesterol is low.
- Post-translational control: HMG-CoA reductase undergoes sterol-sensitive degradation and other regulatory inputs that adjust pathway flux.
- Feedback regulation: Excess cellular cholesterol suppresses endogenous synthesis and modulates LDL receptor expression and the PCSK9 axis (Goldstein & Brown, 2015; Luo et al., 2020).
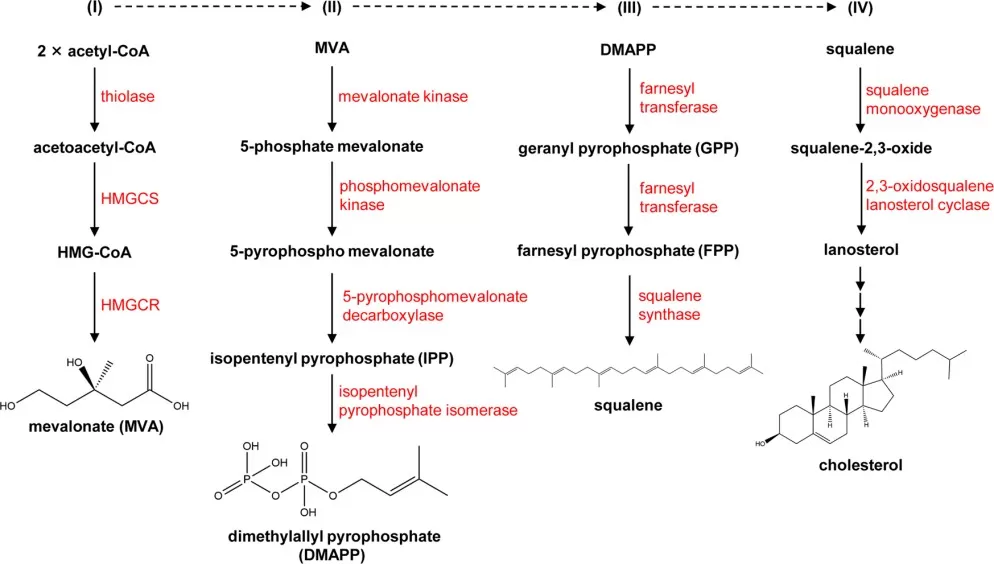
Figure 2. The pathway for cholesterol biosynthesis. Image reproduced from Duan et al., 2022, Signal Transduction and Targeted Therapy, 7(1), 265.
1.2 Dietary Intake and Absorption
Endogenous synthesis is only part of the story. Dietary cholesterol is absorbed in the small intestine through the Niemann-Pick C1-Like 1 (NPC1L1) transporter and then packaged into chylomicrons for transport to the liver and peripheral tissues. In a typical Western diet, approximately 300–500 mg of cholesterol enters the intestine from food each day, although the fraction ultimately absorbed varies substantially among individuals because cholesterol absorption is influenced by transporters, micellar solubilization, and host physiology. The identification of NPC1L1 also clarified the molecular basis for the cholesterol-lowering action of ezetimibe (Altmann et al., 2004; Wang, 2007).
2. PHYSIOLOGICAL FUNCTIONS OF CHOLESTEROL
Cholesterol is indispensable across multiple biological systems. Its functions extend well beyond passive structural support.
2.1 Membrane Structure and Lipid Rafts
Cholesterol is a critical modulator of plasma membrane biophysical properties. By intercalating between phospholipid acyl chains, it helps tune membrane order, permeability, curvature, and protein organization.
Cholesterol-rich membrane microdomains, often discussed as lipid rafts, can concentrate or segregate receptors, kinases, ion channels, and trafficking machinery, thereby shaping signal transduction and membrane sorting events. Depletion of membrane cholesterol disrupts raft integrity and perturbs downstream signaling in many experimental systems (Simons & Toomre, 2000).
2.2 Precursor for Steroid Hormones and Bile Acids
Cholesterol is the universal precursor for several classes of bioactive molecules (Luo et al., 2020; Peet et al., 1998):
- Steroid hormones: glucocorticoids, mineralocorticoids, and sex steroids that regulate immunity, metabolism, electrolyte balance, and reproduction.
- Bile acids: hepatically synthesized molecules that emulsify dietary lipids and also act as signaling ligands through receptors such as FXR and TGR5.
- Vitamin D: derived from 7-dehydrocholesterol in the skin and subsequently activated through hepatic and renal hydroxylation steps.
- Oxysterols: oxidized cholesterol derivatives that participate in cholesterol homeostasis and nuclear receptor signaling.
2.3 Developmental Signaling
During development, cholesterol contributes directly to Hedgehog signaling. A classic discovery showed that Hedgehog proteins undergo covalent cholesterol modification during autoprocessing, which influences ligand distribution and signaling range (Porter et al., 1996). This helps explain why inherited defects in cholesterol biosynthesis can have profound developmental consequences.
3. CHOLESTEROL TRANSPORT AND HOMEOSTASIS
Maintaining cholesterol homeostasis requires coordinated control of synthesis, uptake, trafficking, storage, efflux, and excretion across tissues.
3.1 Lipoprotein Transport System
Because cholesterol is highly hydrophobic, it cannot circulate freely in plasma and must be transported in lipoprotein particles (Luo et al., 2020). The major classes include:
- Chylomicrons: carry dietary lipids and cholesterol from the intestine to peripheral tissues and then to the liver.
- Very-low-density lipoprotein (VLDL), intermediate-density lipoprotein (IDL), and low-density lipoprotein (LDL): represent a hepatic secretion and remodeling pathway that ultimately produces LDL, the major apolipoprotein B-containing particle delivering cholesterol to peripheral cells.
- High-density lipoprotein (HDL): participates in cholesterol exchange and reverse cholesterol transport, helping remove excess cholesterol from peripheral tissues.
PCSK9 is a major post-transcriptional regulator of LDL receptor abundance because it promotes lysosomal degradation of LDLR. This pathway is clinically important: current cardiovascular guidelines still position statins as first-line lipid-lowering therapy, while ezetimibe, PCSK9 inhibitors, inclisiran, and bempedoic acid are layered in as adjunctive options in selected patients rather than replacing statins outright (Virani et al., 2023).
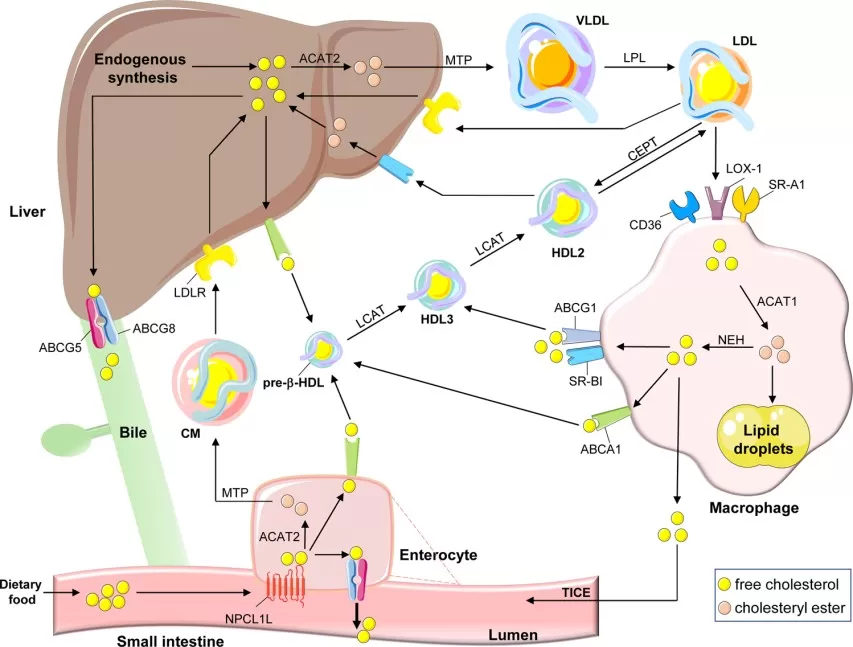
Figure 3. Regulation of cholesterol transport. Image reproduced from Duan et al., 2022, Signal Transduction and Targeted Therapy, 7(1), 265.
3.2 Intracellular Cholesterol Trafficking
After LDL uptake, cholesteryl esters are hydrolyzed in endolysosomal compartments, and free cholesterol is redistributed to the endoplasmic reticulum, plasma membrane, mitochondria, and other organelles. NPC1 and NPC2 are central to this intracellular transport pathway; loss of their function causes Niemann-Pick disease type C, a disorder characterized by lysosomal cholesterol accumulation and progressive neurovisceral pathology (Ikonen, 2006). When cellular free cholesterol rises above immediate needs, part of it is esterified by SOAT/ACAT enzymes and stored in lipid droplets (Luo et al., 2020).
3.3 Reverse Cholesterol Transport
Efflux pathways are equally important. ABCA1 plays a central role in transferring cholesterol and phospholipids to lipid-poor apolipoproteins and initiating nascent HDL formation, while related pathways support additional cholesterol export to HDL particles. In macrophages, impaired efflux favors foam-cell formation and lesion progression, whereas effective cholesterol export and centripetal transport toward the liver are key anti-atherogenic mechanisms (Tall et al., 2002).
4. CHOLESTEROL DYSREGULATION AND DISEASE
Disruption of cholesterol homeostasis contributes to a broad spectrum of diseases.
4.1 Atherosclerotic Cardiovascular Disease
The causal relationship between elevated LDL cholesterol and atherosclerotic cardiovascular disease is among the most firmly established principles in translational lipid biology. Excess apoB-containing lipoproteins promote arterial retention of cholesterol, inflammation, foam-cell formation, plaque progression, and, eventually, myocardial infarction or stroke. Familial hypercholesterolaemia provides a clear human model of this biology because inherited defects in LDLR, APOB, or PCSK9 profoundly elevate LDL-C and accelerate coronary disease (Goldstein & Brown, 2015; Nordestgaard et al., 2013; Duan et al., 2022).
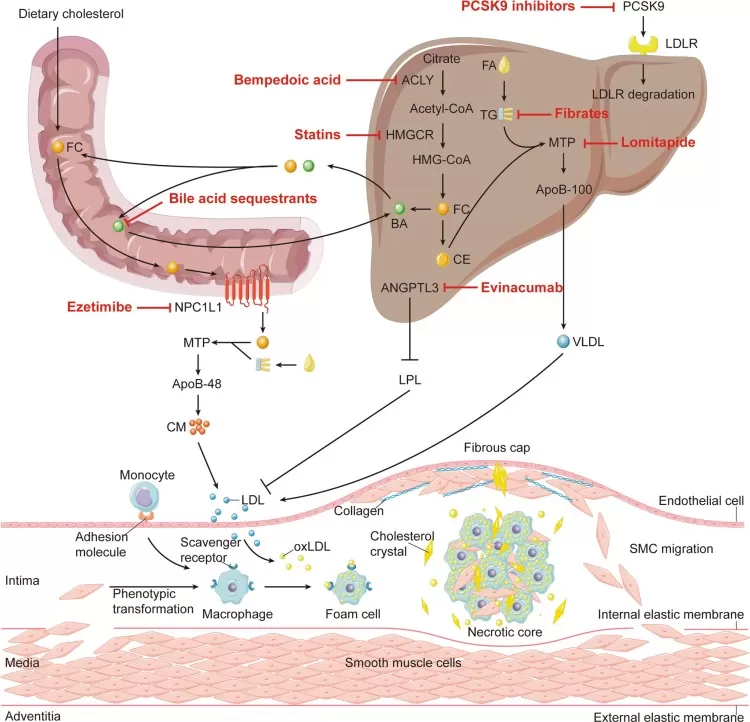
Figure 4. Inhibition of atherosclerosis by cholesterol-lowering interventions. Image reproduced from Duan et al., 2022, Signal Transduction and Targeted Therapy, 7(1), 265.
4.2 Cholesterol Metabolism in Cancer
Cancer cells frequently reprogram cholesterol metabolism. Increased synthesis, uptake, esterification, and compartmental redistribution can support membrane biogenesis, receptor signaling, adaptation to stress, and crosstalk with the tumor microenvironment. Cholesterol-derived metabolites can also reshape immune responses in tumors (Huang et al., 2020; Clendening & Penn, 2012).
Although preclinical evidence for targeting cholesterol pathways in cancer is substantial, clinical translation remains context dependent and tumor type specific. Statins and other interventions remain of strong mechanistic interest, but efficacy is unlikely to be uniform across tumor settings (Huang et al., 2020; Clendening & Penn, 2012).
4.3 Cholesterol and Neurodegeneration
The central nervous system is highly cholesterol enriched and relies largely on local synthesis rather than direct import of plasma lipoproteins. Accordingly, disturbances in cholesterol trafficking, esterification, and oxysterol metabolism can have outsized neurological consequences.
In Alzheimer's disease, APOE genotype, membrane cholesterol distribution, and amyloid processing are tightly intertwined, whereas NPC-related lysosomal cholesterol accumulation provides another example of how defective sterol trafficking can damage neuronal function (Puglielli et al., 2003; Ikonen, 2006).
4.4 Cholesterol in Metabolic Liver Disease
In metabolic liver disease and steatohepatitis, hepatic free cholesterol accumulation is increasingly recognized as a contributor to lipotoxicity, inflammation, and fibrogenesis. Importantly, the biologically harmful species is not simply bulk neutral lipid storage, but the dysregulated handling of free cholesterol and its downstream effects on hepatocytes, Kupffer cells, and stellate cells (Ioannou, 2016).
5. EMERGING FRONTIERS IN CHOLESTEROL RESEARCH
5.1 Cholesterol and Ferroptosis
Ferroptosis is an iron-dependent form of regulated cell death driven by lipid peroxidation. Its best-established biology centers on oxidizable membrane lipids, iron handling, and GPX4-related antioxidant defenses. Cholesterol metabolism intersects with this landscape through oxysterol generation, membrane organization, and broader lipid remodeling, but the precise mechanistic links remain an active area of investigation rather than a settled consensus (Stockwell et al., 2017; Huang et al., 2020).
5.2 Gut Microbiota, Bile Acids, and Cholesterol Metabolism
The gut microbiota exerts major effects on bile acid composition and signaling, which in turn feed back on cholesterol balance, intestinal absorption, hepatic metabolism, and inflammation (Collins et al., 2023):
- Conversion of primary bile acids to secondary bile acids, altering host receptor signaling and enterohepatic feedback loops.
- Production of microbial metabolites that influence host lipid and sterol metabolism.
- Modulation of gut-liver signaling networks that shape cholesterol homeostasis and inflammatory tone.
This gut-bile acid-cholesterol axis is now a major research frontier in metabolic disease, gastrointestinal disease, and host-microbe signaling (Collins et al., 2023).
5.3 Therapeutic Targeting of Cholesterol Metabolism
The therapeutic landscape continues to expand beyond statins. Existing and emerging strategies act at distinct points in cholesterol homeostasis (Virani et al., 2023):
- Intestinal cholesterol absorption inhibitors such as ezetimibe.
- PCSK9-targeted therapies, including monoclonal antibodies and small interfering RNA approaches.
- ATP-citrate lyase inhibition with bempedoic acid.
- Layered combination strategies that build on statins rather than displacing them as first-line therapy.
- Experimental modulation of LXR signaling, which remains of interest because of its roles in cholesterol efflux and bile acid metabolism (Peet et al., 1998).
6. OMICS APPROACHES FOR CHOLESTEROL RESEARCH
The complexity of cholesterol biology — spanning biosynthesis, transport, oxidative metabolism, signaling, and compartment-specific regulation — demands multidimensional analytical strategies. Modern omics platforms have transformed the depth and breadth of cholesterol research.
6.1 Lipidomics for Cholesterol and Sterol Profiling
Mass spectrometry-based lipidomics is central to modern cholesterol research because it can profile multiple classes of cholesterol-related analytes within the same experimental framework:
- Free cholesterol and cholesteryl esters, including ester species distinguished by acyl-chain composition.
- Oxysterols, such as 25-hydroxycholesterol and 27-hydroxycholesterol.
- Bile acids across primary and secondary pools.
- Sterol intermediates, such as lanosterol, desmosterol, and lathosterol, that report on pathway activity.
At the same time, cholesterol-related analytics require careful method design: accurate mass alone is often insufficient to distinguish isomeric sterols, so robust workflows typically rely on chromatographic separation, internal standards, derivatization strategies, and/or multistage fragmentation (Karu et al., 2007).
6.2 Metabolomics in Cholesterol Pathway Analysis
Targeted and untargeted metabolomics place cholesterol within broader metabolic context. Rather than functioning only as a biomarker platform, metabolomics increasingly supports mechanism-oriented studies that connect perturbations in sterol metabolism to coordinated changes in central carbon metabolism, redox state, bile acid biochemistry, and signaling metabolites (Johnson et al., 2016).
NMR-based metabolomics can also provide complementary information, particularly for lipoprotein subclass profiling and systems-level metabolic phenotyping.
 principles_1776132640_WNo_1007d446.webp)
Figure 5. Liquid chromatography–tandem mass spectrometry (LC–MS/MS) principles. Image reproduced from Dewi et al., 2023, Journal of Analytical Science and Technology, 14, 13.
6.3 Proteomics for Cholesterol Regulatory Networks
Proteomics adds complementary information about the enzymes, transporters, receptors, and signaling pathways that govern cholesterol homeostasis. Depending on the study question, quantitative proteomics can be applied to:
- Lipoprotein proteome characterization beyond canonical apolipoproteins.
- Mapping proteins associated with cholesterol-rich membrane environments or sterol-responsive signaling states.
- Quantifying pathway regulators such as LDLR- and PCSK9-related proteins with targeted or discovery workflows.
Phosphoproteomics can further reveal how cholesterol depletion or accumulation reshapes kinase signaling networks downstream of receptor systems enriched in cholesterol-sensitive membrane domains.
6.4 Transcriptomics and Multi-Omics Integration
Transcriptomics adds another layer by capturing how cholesterol-related gene networks — including HMGCR, LDLR, ABCA1, ABCG1, PCSK9, CYP7A1, and other metabolic regulators — respond across tissues, cell states, and disease contexts.
Combined multi-omics strategies are especially powerful when the scientific question is not simply whether cholesterol changes, but which molecular programs change with it, in which direction, and in which biological compartment. Integrative frameworks can therefore link sterol measurements with pathway regulation, cellular phenotypes, and disease mechanisms more effectively than any single omics layer alone (Picard et al., 2021).
MetwareBio: Omics Support for Cholesterol Research
As multi-omics approaches continue to advance cholesterol research, choosing the right analytical strategy is key. MetwareBio supports cholesterol-focused studies with integrated lipidomics, metabolomics, proteomics, and multi-omics solutions for mechanism exploration, biomarker discovery, and pathway analysis.
To learn how our omics workflows can support your cholesterol research, contact our team.
Contact UsReferences
- Altmann, S. W., Davis, H. R., Jr., Zhu, L. J., Yao, X., Hoos, L. M., Tetzloff, G., Iyer, S. P., Maguire, M., Golovko, A., Zeng, M., Wang, L., Murgolo, N., & Graziano, M. P. (2004). Niemann-Pick C1 Like 1 protein is critical for intestinal cholesterol absorption. Science, 303(5661), 1201–1204. https://doi.org/10.1126/science.1093131
- Clendening, J. W., & Penn, L. Z. (2012). Targeting tumor cell metabolism with statins. Oncogene, 31(48), 4967–4978. https://doi.org/10.1038/onc.2012.6
- Collins, S. L., Stine, J. G., Bisanz, J. E., Okafor, C. D., & Patterson, A. D. (2023). Bile acids and the gut microbiota: metabolic interactions and impacts on disease. Nature Reviews Microbiology, 21, 236–247. https://doi.org/10.1038/s41579-022-00805-x
- Dewi, K. R., Ismayati, M., Solihat, N. N., et al. (2023). Advances and key considerations of liquid chromatography–mass spectrometry for porcine authentication in halal analysis. Journal of Analytical Science and Technology, 14, 13. https://doi.org/10.1186/s40543-023-00376-3
- Duan, Y., Gong, K., Xu, S., Zhang, F., Meng, X., & Han, J. (2022). Regulation of cholesterol homeostasis in health and diseases: from mechanisms to targeted therapeutics. Signal Transduction and Targeted Therapy, 7(1), 265. https://doi.org/10.1038/s41392-022-01125-5
- Goldstein, J. L., & Brown, M. S. (2015). A century of cholesterol and coronaries: from plaques to genes to statins. Cell, 161(1), 161–172. https://doi.org/10.1016/j.cell.2015.01.036
- Huang, B., Song, B.-L., & Xu, C. (2020). Cholesterol metabolism in cancer: mechanisms and therapeutic opportunities. Nature Metabolism, 2, 132–141. https://doi.org/10.1038/s42255-020-0174-0
- Ikonen, E. (2006). Mechanisms for cellular cholesterol transport: defects and human disease. Physiological Reviews, 86(4), 1237–1261. https://doi.org/10.1152/physrev.00022.2005
- Ioannou, G. N. (2016). The role of cholesterol in the pathogenesis of NASH. Trends in Endocrinology and Metabolism, 27(2), 84–95. https://doi.org/10.1016/j.tem.2015.11.008
- Johnson, C. H., Ivanisevic, J., & Siuzdak, G. (2016). Metabolomics: beyond biomarkers and towards mechanisms. Nature Reviews Molecular Cell Biology, 17, 451–459. https://doi.org/10.1038/nrm.2016.25
- Karu, K., Hornshaw, M., Woffendin, G., Bodin, K., Hamberg, M., Alvelius, G., Sjovall, J., Turton, J., Wang, Y., & Griffiths, W. J. (2007). Liquid chromatography-mass spectrometry utilizing multi-stage fragmentation for the identification of oxysterols. Journal of Lipid Research, 48(4), 976–987. https://doi.org/10.1194/jlr.M600497-JLR200
- Luo, J., Yang, H., & Song, B.-L. (2020). Mechanisms and regulation of cholesterol homeostasis. Nature Reviews Molecular Cell Biology, 21, 225–245. https://doi.org/10.1038/s41580-019-0190-7
- Nordestgaard, B. G., Chapman, M. J., Humphries, S. E., Ginsberg, H. N., Masana, L., Descamps, O. S., Wiklund, O., Hegele, R. A., Raal, F. J., Defesche, J. C., Wiegman, A., Santos, R. D., Watts, G. F., Parhofer, K. G., Hovingh, G. K., Kovanen, P. T., Boileau, C., Averna, M., Boren, J., et al. (2013). Familial hypercholesterolaemia is underdiagnosed and undertreated in the general population: guidance for clinicians to prevent coronary heart disease. European Heart Journal, 34(45), 3478–3490. https://doi.org/10.1093/eurheartj/eht273
- Peet, D. J., Turley, S. D., Ma, W., Janowski, B. A., Lobaccaro, J.-M., Hammer, R. E., & Mangelsdorf, D. J. (1998). Cholesterol and bile acid metabolism are impaired in mice lacking the nuclear oxysterol receptor LXRalpha. Cell, 93(5), 693–704. https://doi.org/10.1016/S0092-8674(00)81432-4
- Picard, M., Scott-Boyer, M.-P., Bodein, A., Perin, O., & Droit, A. (2021). Integration strategies of multi-omics data for machine learning analysis. Computational and Structural Biotechnology Journal, 19, 3735–3746. https://doi.org/10.1016/j.csbj.2021.06.030
- Porter, J. A., Young, K. E., & Beachy, P. A. (1996). Cholesterol modification of Hedgehog signaling proteins in animal development. Science, 274(5285), 255–259. https://doi.org/10.1126/science.274.5285.255
- Puglielli, L., Tanzi, R. E., & Kovacs, D. M. (2003). Alzheimer's disease: the cholesterol connection. Nature Neuroscience, 6(4), 345–351. https://doi.org/10.1038/nn0403-345
- Simons, K., & Toomre, D. (2000). Lipid rafts and signal transduction. Nature Reviews Molecular Cell Biology, 1(1), 31–39. https://doi.org/10.1038/35036052
- Stockwell, B. R., Friedmann Angeli, J. P., Bayir, H., Bush, A. I., Conrad, M., Dixon, S. J., Fulda, S., Gascon, S., Hatzios, S. K., Kagan, V. E., Noel, K., Jiang, X., Linkermann, A., Murphy, M. E., Overholtzer, M., Oyagi, A., Pagnussat, G. C., Park, J., Ran, Q., et al. (2017). Ferroptosis: a regulated cell death nexus linking metabolism, redox biology, and disease. Cell, 171(2), 273–285. https://doi.org/10.1016/j.cell.2017.09.021
- Tall, A. R., Costet, P., & Wang, N. (2002). Regulation and mechanisms of macrophage cholesterol efflux. Journal of Clinical Investigation, 110(7), 899–904. https://doi.org/10.1172/JCI16391
- Virani, S. S., Newby, L. K., Arnold, S. V., Bittner, V., Brewer, L. C., Demeter, S. H., Dixon, D. L., Fleg, J. L., Hess, B., Hiatt, W. R., et al. (2023). 2023 AHA/ACC/ACCP/ASPC/NLA/PCNA guideline for the management of patients with chronic coronary disease. Circulation, 148(9), e9–e119. https://doi.org/10.1161/CIR.0000000000001168
- Wang, D. Q.-H. (2007). Regulation of intestinal cholesterol absorption. Annual Review of Physiology, 69, 221–248. https://doi.org/10.1146/annurev.physiol.69.031905.160725


